![]() Murilo S. Dias1
Murilo S. Dias1 ![]() ,
, ![]() Céline Jézéque2,
Céline Jézéque2, ![]() Bernard Hugueny2,
Bernard Hugueny2, ![]() Fabien Leprieur3,
Fabien Leprieur3, ![]() Sébastien Brosse2,
Sébastien Brosse2, ![]() James Albert4,
James Albert4, ![]() Fernanda A. S. Cassemiro5,
Fernanda A. S. Cassemiro5, ![]() Renata G. Frederico6,
Renata G. Frederico6, ![]() Max Hidalgo7,
Max Hidalgo7, ![]() Hernan Ortega7,
Hernan Ortega7, ![]() Pablo A. Tedesco2,
Pablo A. Tedesco2, ![]() Gislene Torrente-Vilara8,
Gislene Torrente-Vilara8, ![]() Jansen Zuanon9,10,
Jansen Zuanon9,10, ![]() Rémy Bigorne2,
Rémy Bigorne2, ![]() Koen Martens11,
Koen Martens11, ![]() Astrid Acosta12,
Astrid Acosta12, ![]() Edwin Agudelo12,
Edwin Agudelo12, ![]() Soraya Barrera Maure13,
Soraya Barrera Maure13, ![]() Douglas A. Bastos9,
Douglas A. Bastos9, ![]() Juan Bogotá-Gregory14,
Juan Bogotá-Gregory14, ![]() Fernando G. Cabeceira14,
Fernando G. Cabeceira14, ![]() André L. C. Canto15,
André L. C. Canto15, ![]() Fernando M. Carvajal-Vallejos16,
Fernando M. Carvajal-Vallejos16, ![]() Lucélia N. Carvalho17,
Lucélia N. Carvalho17, ![]() Ariana Cella-Ribeiro18,
Ariana Cella-Ribeiro18, ![]() Raphaël Covain19,
Raphaël Covain19, ![]() Carlos DoNascimiento20,
Carlos DoNascimiento20, ![]() Carolina R. C. Dória21,
Carolina R. C. Dória21, ![]() Cleber Duarte9,
Cleber Duarte9, ![]() Efrem J. G. Ferreira9,
Efrem J. G. Ferreira9, ![]() André V. Galuch9,22,
André V. Galuch9,22, ![]() Tommaso Giarrizzo23,
Tommaso Giarrizzo23, ![]() Rafael P. Leitão6,
Rafael P. Leitão6, ![]() John G. Lundberg24,
John G. Lundberg24, ![]() Mabel Maldonado16,
Mabel Maldonado16, ![]() Luciano F. A. Montag25,
Luciano F. A. Montag25, ![]() Willian Massaharu Ohara26,
Willian Massaharu Ohara26, ![]() Tiago H. S. Pires9,
Tiago H. S. Pires9, ![]() Marc Pouilly27,
Marc Pouilly27, ![]() Saúl Prada-Pedreros28,
Saúl Prada-Pedreros28, ![]() Luiz J. de Queiroz29,
Luiz J. de Queiroz29, ![]() Lucia Rapp Py-Daniel9,
Lucia Rapp Py-Daniel9, ![]() Frank R. V. Ribeiro15,
Frank R. V. Ribeiro15, ![]() Raúl Ríos Herrera30,
Raúl Ríos Herrera30, ![]() Jaime Sarmiento13,
Jaime Sarmiento13, ![]() Leandro M. Sousa31,
Leandro M. Sousa31, ![]() Lis F. Stegmann9,32,
Lis F. Stegmann9,32, ![]() Jonathan Valdiviezo-Rivera33,
Jonathan Valdiviezo-Rivera33, ![]() Francisco Villa34,
Francisco Villa34, ![]() Takayuki Yunoki35,
Takayuki Yunoki35, ![]() Aaike de Wever36 and
Aaike de Wever36 and ![]() Thierry Oberdorff2
Thierry Oberdorff2
PDF: Download Here | Supplementary: S1 S2 S3 S4 S5 S6 S7 S8 S9 S10 | Cite this article
Associate Editor: ![]() Caroline Arantes
Caroline Arantes
Section Editor: ![]() William Crampton
William Crampton
Editor-in-chief: ![]() José Birindelli
José Birindelli
Abstract
Identificar as principais dimensões taxonômica, filogenética e de atributos da diversidade beta, e avaliar seus possíveis fatores determinantes, aprimora nossa compreensão dos padrões e processos envolvidos na evolução das assembleias biológicas. Utilizando bancos de dados abrangentes sobre a distribuição, filogenia e atributos morfológicos dos peixes de água doce amazônicos, analisamos os padrões de substituição de espécies dessas três dimensões para avaliar seus potenciais determinantes históricos e contemporâneos usando regressão múltipla em matrizes de distâncias. Observamos que a diversidade beta taxonômica média foi cerca de duas vezes maior que a diversidade beta filogenética média e seis vezes maior que a diversidade beta baseada em atributos das espécies, com padrões espaciais coincidentes nas dimensões Taxoβsim e Phyloβsim, enquanto a Traitβsim apresentou-se mais difusa e heterogênea no espaço. Nós encontramos a influência proeminente das distâncias geográficas entre sub-bacias, da severidade do habitat e dos tipos de cor da água nas dimensões taxonômica e filogenética da diversidade beta, junto com efeitos individuais menores da temperatura atual, tipos de habitat, conexões históricas entre sub-bacias, incursões marinhas e esforço amostral. Em contraste, a Traitβsim foi pouco explicada, sendo influenciada apenas pelo esforço amostral e pelas condições hidromorfológicas atuais das sub-bacias. Esses resultados apontam para efeitos predominantes da limitação de dispersão, filtragem ambiental e contingências históricas na explicação dos padrões de diversidade beta taxonômica e filogenética dos agrupamentos de peixes amazônicos, mas não na substituição de atributos funcionais.
Palavras-chave: Assembleias de peixes, Bacia Amazônica, Eventos históricos, Filtragem ambiental, Métricas de diversidade beta.
Introduction
Determining the drivers of variation in species composition between assemblages (i.e., taxonomic beta diversity, Taxoβ) is key to understanding the distribution of biodiversity across space and time. Taxonomic beta diversity is linked to important ecological mechanisms, such as environmental filtering (Leibold, Chase, 2018), species dispersal limitation (Hubbell, 2001; Wu et al., 2017; Leibold, Chase, 2018; Guilherme et al., 2022), biotic interactions (Graham, Fine, 2008) and historical contingency (Leprieur et al., 2011; Fitzpatrick et al., 2013; Dias et al., 2014; Fluck et al., 2020; Wu et al., 2017). In addition to taxonomic beta diversity, phylogenetic and functional traits beta diversity (Phyloβ and Traitβ, respectively) have been adopted as complementary biodiversity dimensions (Graham, Fine, 2008; Swenson et al., 2011; Leprieur et al., 2012; Villéger et al., 2013; Brum et al., 2017). While Taxoβ measures species composition dissimilarity among assemblages (Whittaker, 1960), the two other dimensions measure the changes in lineages and species functional traits, respectively, providing insights into the evolutionary history and environmental conditions that shape assemblages (Graham, Fine, 2008; Weinstein et al., 2014; Penone et al., 2016). Although these three beta diversity measures are correlated due to phylogenetic constraints (Cadotte et al., 2009; Swenson et al., 2011; Fritz, Rahbek, 2012; Weinstein et al., 2014; Fluck et al., 2020), each metric may add distinct information on the causal mechanisms involved in shaping assemblage formation (Mcgill et al., 2006; Graham, Fine, 2008; Weinstein et al., 2014; Penone et al., 2016; Brum et al., 2017; Mazel et al., 2017; Villéger et al., 2017, 2013; Lu et al., 2019; McLean et al., 2021). Thus, analyzing these three beta diversity dimensions together, and highlighting the drivers affecting each of them, may advance our understanding of patterns and processes involved in the evolution of biological assemblages.
The Amazon Basin is a major biodiversity hotspot (Antonelli et al., 2018), containing, among others, the highest freshwater biodiversity on earth (Tisseuil et al., 2013). This is notably true for Amazonian freshwater fishes which represent ~15% of all freshwater fish species described worldwide (Jézéquel et al., 2020a; Tedesco et al., 2017a). Much of this diversity is thought to be associated with major geological landscape events that took place between 30–7 Ma, as suggested by a continental-scale study using the entire South American freshwater fish fauna (Cassemiro et al., 2023). Compared to other large riverine ecosystems worldwide, the Amazon Basin is still relatively well preserved (Reis et al., 2016) but is facing rising anthropogenic threats, including damming, deforestation and climate change, that may potentially endanger its exceptional fish fauna in the near future (Reis et al., 2016; Pelicice et al., 2017; Albert et al., 2020, 2023; Herrera-R et al., 2020; Frederico et al., 2021; Couto et al., 2024; Correa et al., 2025). Basin-wide approaches to understand the mechanisms underlying the origin and maintenance of Amazonian fish diversity patterns are thus urgently needed to organize evidence-based policy developmentand conservation planning (Socolar et al., 2016; Jézéquel et al., 2020b; Dagosta et al., 2021; Flecker et al., 2022; Caldas et al., 2023).
Despite recent advances describing Amazonian fish diversity patterns and diversification (Dagosta, Pinna, 2017, 2019; Boschman et al., 2023; Cassemiro et al., 2023), only two studies covering the entire Amazon drainage basin have attempted so far to quantitatively analyze its basin-wide fish diversity drivers. The first one showed prominent influences of current and past climatic conditions, habitat size, and natural fragmentation on Amazonian sub-basins species richness, but also highlighted an independent West-East gradient of decreasing species richness, suggesting a center of fish diversity located westward and a colonization eastward from this center not yet achieved (Oberdorff et al., 2019). Based on the same framework, the second study showed, in particular, that Western Amazon sub-basins were more phylogenetically clustered but less phylogenetically diverse compared to Eastern Amazon ones, suggesting deeper evolutionary divergences among taxa located to the East, and more recent radiations in the Western sub-drainages (Salgueiro et al., 2022). Based on these findings and integrating the West-East gradient of decreasing richness (Oberdorff et al., 2019), Salgueiro et al. (2022) concluded that Western Amazon was an evolutionary “cradle” of biodiversity (i.e., location with unusually high rates of recent speciation), in line with the accelerating net diversification rates during the last 20 Ma (mainly in Western Amazon) found by Cassemiro et al. (2023) and Boschman et al. (2023). Taken together, results from Salgueiro et al. (2022), Cassemiro et al. (2023), and Boschman et al. (2023) suggest that Western Amazon is the most important source of fish lineages for the Amazon Basin.
Here, we analyze how historical and contemporary ecological factors interact to generate modern patterns of taxonomic, phylogenetic and functional traits beta diversity across the Amazon Basin. Given the large spatial grain (i.e., sub-basins) and geographical extent (i.e., the whole Amazon Basin) of our study and the limited dispersal capacity of species, we expect spatial distance (i.e., dispersal limitation), and to a lesser extent environmental filtering, to be the most important drivers of taxonomic beta diversity (Barton et al., 2013). On the other hand, we expect phylogenetic changes in lineages across sub-basins (i.e., turnover) to be highly dictated by the geological and historical events that occurred in the basin over the last 20 Ma (Boschman et al., 2023; Cassemiro et al., 2023). Finally, we anticipate functional (i.e., species morphological traits in our case, but see Olden et al., 2025) beta diversity to be notably dictated by current drivers such as differences in hydrological habitat types between sub-basins.
Material and methods
Biological data. Fish data were retrieved from the AmazonFish project (www.amazon-fish.com), which offers the most complete fish distribution database available for the entire Amazon drainage basin (Jézéquel et al., 2020a). Briefly, occurrence fish records were compiled (and are regularly updated) by mobilizing and integrating all information available in published articles, books, gray literature, online databases, worldwide museums and Universities, and expeditions conducted during the project. All occurrence records were validated by freshwater fish specialists and taxonomists (for details on data construction, see Jézéquel et al., 2020a,b). The database currently contains 300,000 georeferenced records for 2,500 valid native freshwater fish species from 514 genera and 56 families. As we were interested in riverine organisms, we excluded species from the genus Orestias because they are mostly restricted to lakes in the Andean highlands (Scott et al., 2020).
Here, our defined fish assemblages consist of 2,392 species presence/absence in each of our 97 sub-basins. The corresponding point data containing the geographic coordinates of all georeferenced records and the information sources can be freely downloaded from the AmazonFish website at https://figshare.com/articles/dataset/A_database_of_freshwater_fish_species_of_the_Amazon_Basin/9923762. The choice of this large spatial grain was motivated by the heterogeneous distribution of sampling effort within the basin, limiting the possibilities to work at smaller spatial grains (Oberdorff et al., 2019). The full method for sub-basins delineation and dataset construction are detailed in (Oberdorff et al., 2019; Jézéquel et al., 2020a,b), respectively. Briefly, we classified our sub-drainage basins based on the HydroBASIN framework (Lehner, Grill, 2013) and combined different HydroBASIN levels to retain only sub-basins >20,000 km2 to optimize sampling effort (Oberdorff et al., 2019). According to (Jézéquel et al., 2020b) all but nine of these 97 sub-basins have >60% of completeness estimated based on Chao 2 richness estimators (i.e., Demini, Paru_Este, Jari, Jamanxim, Grande, Tapaua, Apurimac1, Ucayali2, Curaray; see fig. S1 from Oberdorff et al., 2019). A sub-basin with completeness ratio >0.6 is considered well surveyed (Troia, McManamay, 2016). The nine sub-basins having < 60% of completeness may thus generate potential biases in turnover values linked to insufficient sub-basin sampling effort. However, including or excluding these nine under-sampled sub-basins from analyses (see below) did not change the overall results so that we decided to discuss results obtained for the whole dataset (i.e., 97 sub-basins).
We obtained phylogenetic information on Amazonian fishes from the supertree established by (Rabosky et al., 2018). The backbone of this ultrametric supertree consists of 11,638 species with genetic data available, a calibration process including 130 fossil points, and the insertion of 19,888 species based on a stochastic polytomy resolution algorithm (Rabosky, 2020). In the Amazon Basin, we found 635 species for which genetic data were available and 1451 for which inclusion was based on the polytomy algorithm, resulting in 2086 fish species in the final pruned tree (~87% of Amazonian species).As the supertree consists of 100 samples from the posterior distribution (Rabosky et al., 2018; Rabosky, 2020), we relied our analyses on each one of the 100 samples and estimated their mean dissimilarity values over all phylogenetic pruned trees. To confirm results obtained from this phylogenetic information, we further calculated the phylogenetic beta diversity metrics with a newly compiled, comprehensive phylogeny of Neotropical freshwater fish taxa based exclusively on DNA sequences (Cassemiro et al., 2023) and containing 1,111 species from our AmazonFish dataset. As phylogenetic beta diversity index calculated with both phylogenies were highly correlated (Mantel r= 0.90; p < 0.001; Fig. S1) and our results were similar whatever the phylogenetic information used, we discuss below only those stemming from Rabosky’s phylogeny (i.e., 2086 species), but also provide those obtained using the 1,111 species available in the phylogeny from Cassemiro et al. (2023).
Fish species functional traits were gathered from the database developed by Brosse et al. (2021) and concerned exclusively morphological attributes related to species body size and body shape. The body shape of species was described using nine unitless measures (i.e., body elongation, eye vertical position, relative eye position, oral gape position, relative maxillary length, body lateral shape, pectoral fin vertical position, pectoral fin size, and caudal peduncle throttling) linked to species’ locomotion and food acquisition (see fig. 1 from Brosse et al., 2021). The data set used here included 1,661 fish species for which trait information was consistently available (>69% of our Amazonian fish species pool). To summarize, the calculation of the three beta diversity metrics was performed on distinct pool of species for each facet (i.e., taxonomic: 2,392 species; phylogenetic: 2,086 species; functional: 1,661 species).
Current, past and spatial predictors. Large scale biodiversity patterns can be explained by a range of ecological and historical drivers (Ricklefs, 2004; Brown, 2014), and most of these drivers also apply for freshwater fishes at large spatial scales (Hugueny et al., 2010). These drivers can be summarized under climate/productivity, area/environmental heterogeneity, historical/evolutionary, and spatial hypotheses (Ricklefs, 2004; Hugueny et al., 2010; Brown, 2014). Data sources and definitions of the drivers used in this study are presented in detail in (Oberdorff et al., 2019), and we only provide here a brief overview of each of them. All predictors described below were extracted for each of the 97 sub-drainage basins, providing a single mean value for each variable, and subsequently transformed into dissimilarity matrices (see below).
Current environmental predictors. To estimate the effects of climate and productivity, we used the annual mean and seasonality (Coefficient of Variation of intra-year monthly values calculated as the standard deviation divided by the mean) of temperature (Temp), precipitation (Prec), actual evapotranspiration (AET), potential evapotranspiration (PET), net primary productivity (NPP), solar radiation (SolRad), run-off (RO), and the lowest (or highest) value of minimum (or maximum) temperature of the coldest (or warmest) month from WorldClim (version 1). These variables measure the mean current climatic condition, the seasonal climatic variability and the energy availability within each sub-drainage basin.
Habitat size and diversity were estimated using the surface area of the sub-drainage basin (km2; Area), the network density (i.e., length of the riverine network divided by the surface area of the sub-basin, a measure of habitat availability for fishes; NetwD), the land cover heterogeneity (i.e., the proportion of native land cover classes within each sub-drainage basin; VegCover), and the soil heterogeneity (i.e., the proportion of each soil type within each sub-drainage basin; SoilCover). Finally, we used the hydromorphological heterogeneity (i.e., the proportion of river hydromorphological types within each sub-basin; HydromorphoTypes). This last variable is derived from a recently developed classification scheme defining homogenous geomorphological river types throughout the Amazon Basin (Jézéquel et al., 2022). Each river type, based on similar processes and river form characteristics (e.g., channel planform, islands, bars, river bed dominant substrates size), is supposed to partly reflect hydrological constrains for biological assemblages (e.g., Chessman et al., 2006).
Amazonian waters were divided into three distinct biogeochemical water types or “colors”, differentiated by sediment composition, geochemistry and optical characteristics: whitewaters have an Andean origin (e.g., the Madeira River and the Amazon mainstem), characterized by their usually low transparency due to large amounts of sediment particles and a neutral pH (pH ~6–8.5); nutrient poor blackwaters mostly draining the Precambrian Guiana shield (e.g., the Negro River) characterized by their brownish color and acidic waters (pH < 5); and nutrient-poor, low-sediment, high-transparency and slightly acidic clearwaters (pH ~ 5.3–7.2) mostly draining the Brazilian and Guianas shields (e.g., the Tapajós and Xingu Rivers) (Ríos-Villamizar et al., 2020). Fish assemblages respond to water types (e.g., Bogotá-Gregory et al., 2020), hence we classified sub-basins according to their main water type as detailed in (Venticinque et al., 2016) (see also fig. S2 from Oberdorff et al., 2019). The three water color types were coded as categorical variables (coded as black, white and clear water (see Predictors of dissimilarity matrices).
We further estimated the fragmentation and the isolation of sub-drainage basins, both key drivers of freshwater fish diversity at large spatial scale (Dias et al., 2013; Tedesco et al., 2017b), by using the number of waterfalls (>30 m) within each sub-basin using data from Lehner, Grill (2013), the sub-basin elevation (mean, and range; ElevMean and ElevRange), the proportion of the sub-basin surface with terrain slope above 15% (Csup15) and the proportion of the sub-basin surface above 1,000 m in altitude (ElevProp1000) (Oberdorff et al., 2019).
The number of sampling sites divided by the surface area of each sub-basin was also included in our models to further control for a potential sampling effort effect (SamplingEffort).
Historical factors. We included variables related to the Amazon Basin geological history from distinct time periods. We identified the sub-basins potentially belonging or not to the Pebas system at ∼23 Mya (Hoorn et al., 2010), the surface area of each sub-basin under sea water considering a sea-level rise of 25m during recent Pleistocene marine incursions (<1 Mya, Miller et al., 2005), and the Quaternary climate stability within the sub-basin (from ∼21 kyr to present; the strongest climatic shift in recent history, Sandel et al., 2011). We used Quaternary climate reconstructions of mean, max and min annual temperatures and precipitations at the Last Glacial Maximum (LGM; 21 kyr) from three General Circulation Models (CCSM, MIROC, and MPI; three independent initiatives from distinct research communities to estimate past, present and future Earth climate metrics) and calculated the difference between current and LGM mean values to describe climate stability (Diff_CurrentLGM).
We used the distance of each of the sub-drainages to the current Amazon river mouth (km) and the longitudinal position (downriver-upriver) of the principal tributary hosting a sub-basin to represent the position of each of the sub-drainages within the Amazon River network (see Oberdorff et al., 2019 for a detailed explanation).
Spatial predictors. Watercourse distance is known to better represent the connections between sites in a riverine network than does the Euclidean spatial distance (Grant et al., 2007; Tonkin et al., 2018; Larsen et al., 2021). Therefore, we calculated the watercourse distance (km) between each of our sub-drainages to represent the path to current fish dispersal. In this way, the watercourse distance matrix represents the distance (km) between each pair of sub-drainages following the dendritic configuration of the Amazon Basin.
Statistical analyses
Biological dissimilarity matrices. To quantify the taxonomic, phylogenetic and functional beta diversity, we used the pure turnover component of the Sørensen dissimilarity index for all three facets (i.e., the Simpson dissimilarity, βsim). The Sørensen dissimilarity index (βSør) can be partitioned into two components: pure turnover (i.e., βsim) and nestedness (βsne) (Baselga, 2010, 2012,). As, in our case, βsim represented a larger fraction of the βSør than βsne (see results), we chose here to discuss only results obtained from the former. However, results obtained using βsne are given for information.
The βsim for taxonomic data (i.e., Taxoβsim) quantifies β-diversity due to pure turnover (replacement) of species between sub-basins, independently of their respective differences in species richness (Baselga, 2010, 2012). It is calculated as Taxoβsim = min (b,c) / (a+min (b,c)), where a is the number of species shared by the two concerned sub-basins, b is the number of species unique to one sub-basin and c is the number of species unique to the other sub-basin.
Taxonomic dissimilarity has been extended to phylogenetic and functional (i.e., species traits) data (Leprieur et al., 2012; Villéger et al., 2013). To quantify the pure turnover component of phylogenetic dissimilarity of sub-basins assemblage, we used the Phyloβsim metric which is analogous to Taxoβsim with shared and unique species being replaced by shared and unique branch lengths on the phylogenetic tree (Leprieur et al., 2012).
Finally, we adopted a similar framework to calculate the functional dissimilarity between assemblages (Traitβsim) (Villéger et al., 2013). We used species body size and the nine continuous morphological body ratios (centered and reduced so that they all have zero mean and unity standard deviation) available in Brosse et al. (2021) to compute four Principal Components axes in order to construct the species traits multidimensional space. We used four axes summarizing 71% of variability of the total morphological matrix and giving a good representation of functional spaces (low mean residual variation; 0.61; see Tabs. S2, S3; Figs. S4, S5) without excessively increasing the computation time requested to calculate trait volumes for higher dimensions (Maire et al., 2015). The morphological dissimilarity metric (i.e., Traitβsim) between two assemblages was calculated as the proportion of unshared convex hull volume between a pair of sub-basins after controlling for their size differences in the morphological space (Villéger et al., 2013). All metrics were calculated using the vegan (Oksanen et al., 2022), betapart package (Baselga, Orme, 2012), and picante (Kembel et al., 2010) packages. Taxoβsim, Phyloβsim and Traitβsim metrics vary from 0 (total similarity in assemblage taxonomic or phylogenetic or species traits composition between two sub-basins) to 1 (total dissimilarity in assemblage taxonomic or phylogenetic or species traits composition between two sub-basins).
Matrices of predictors. Predictors were log-transformed, when necessary, to minimize potential effects of extreme values (see Oberdorff et al., 2019). We further standardized all metrics so that they all had mean zero and 0.5 standard deviation (Gelman, 2008); this procedure is key to homogenize their scales and ranges, and assures each variable to have the same weight when computing the dissimilarity predictor matrices. Then, we applied the Euclidean distance, a common approach to deal with continuous environmental variables, to calculate each of the predictor dissimilarity matrices. As the water type variable was categorical with three levels (Tab. 1), we constructed a pairwise binary matrix coding pairs of sub-basins of the same water type as 1, and 0 when different. We calculated the similarity matrix of Pebas past connection/disconnection by coding 1 for pairs of sub-basins both connected to Pebas, and 0 for all other configurations. As spatial watercourse distance was a pairwise matrix (watercourse distance between two sub-basins), no pairwise calculation was further needed. We finally calculated fourteen matrices representing major ecological, historical and evolutionary hypotheses in order to estimate their independent effects on fish assemblage dissimilarities (Tab. 1).
TABLE 1 | All predictors used to construct each dissimilarity matrix and their corresponding explanation group. Some predictors have been transformed (log[x+1] or arcsin of the square root) to improve distribution of values along their gradients (predictors are named following the adopted transformation). Standardized predictor means predictors have been centered (mean = 0) and reduced (sd = 0.5) to give them equal weight prior distance matrix computation. Variance Inflation Factor (VIF) values are shown for all resulting pairwise matrices.
Predictors | Combined variables | Matrix calculation; dissimilarity/similarity | Group | VIF |
D_Energy | AET (min, max, annual, cv), PET (min, max, annual, cv), NPP (mean, sd, cv) | Euclidean distance of standardized predictors; dissimilarity | Current environment | 8.4 |
D_Water | Precipitation (min, max, annual, cv), Runoff (mean) | Euclidean distance of standardized predictors; dissimilarity | Current environment | 2.4 |
D_Temp | Temperature (min, max, mean, cv), Solar Radiation (mean, cv) | Euclidean distance of standardized predictors; dissimilarity | Current environment | 6.2 |
D_HabitatSize | NetworkDensity, SurfaceArea_log | Euclidean distance of standardized predictors; dissimilarity | Current environment | 1.1 |
D_HabitatType | VegType (relative proportion), SoilType (relative proportion) | Euclidean distance of binded VegCover and SoilCover categories; dissimilarity | Current environment | 1.3 |
D_HabitatHarsh | Csup15, ElevMean_log, ElevRange_log, ElevPropBasin-Above1000m_asin_sqrt | Euclidean distance of standardized predictors; dissimilarity | Current/Historical | 2.2 |
D_Fragmentation | NumberWaterfall_log | Euclidean distance of standardized predictor; dissimilarity | Current/Historical | 1.1 |
D_Position | DistanceRiverMouth, PositionCodeAlongWestEast | Euclidean distance of standardized predictors; dissimilarity | Current/Historical | 2.6 |
D_WaterColor | WaterColor (categories black, clear, white) | Same water color = 1, Distinct water color = 0; similarity | Current environment | 1.2 |
D_SampEff | Density of sampling points_log (log[NumbSamplingPoints/Subdrainage surface area]) | Euclidean distance between standardized predictor; dissimilarity | Current environment | 1.2 |
D_CurrentLGM | Difference between Current and Past temp (min, max, mean), precip (min, max, annual) | Euclidean distance of standardized predictors; dissimilarity | Historical | 1.5 |
D_MarineInc | Proportion (%) of Surface area below 25m_log | Euclidean distance of standardized predictor; dissimilarity | Historical | 1.8 |
D_PastConnect | Connected/not Connected to Pebas system | Two sub-basin connected to Pebas = 1, only one connected or no connections to Pebas = 0; similarity | Historical | 1.5 |
D_HydromorphoType | Relative proportion of riverine hydrological habitats (%) | Euclidean distance between relative proportion of hydrogeomorphology riverine categories; dissimilarity | Current environment | 1.4 |
D_Spatial | Spatial distance | WaterCourse pairwise distance among sub-drainages (km); dissimilarity | Space | 2.8 |
Prior to modeling, we assessed multicollinearity among all predictor distance matrices using the Variance Inflation Factor (VIF). Although there are no canonical rules of thumb, VIF values <10 are usually considered as acceptable and indicate low collinearity among predictors (Dormann et al., 2012). As none of our predictor distance matrices presented VIF values >10, all predictors were included simultaneously in our modeling framework.
Concordance among metrics and mapping. We first performed Pearson’ correlation among vectorized Taxoβsim, Phyloβsim, and Traitβsim matrices to estimate correlations between all assemblage beta diversity metrics. As we were dealing with pairwise distance matrices, we mapped spatial assemblage dissimilarity by calculating the mean dissimilarity values of a focal sub-drainage over all pairwise combinations with the other sub-drainages (i.e., calculating mean values over rows of the pairwise matrices), allowing therefore to visualize the spatial patterns of the three beta diversity facets. In order to improve visualization, we also performed Non-Metric Multidimensional Scaling analysis (NMDS) on each individual dissimilarity matrix and used the scores of the two resulting axes as Red Green Blue (RGB) color parameters for each map. As a result, sub-basins with similar species composition are represented with similar colors on the maps (Fig. S6).
Drivers of fish assemblage dissimilarities. As usually done, we modeled the pairwise dissimilarity (i.e., Taxoβsim, Phyloβsim, Traitβsim) against all the dissimilarity matrices related to current climate and environmental factors, historical and geological events, and spatial distance between sub-drainage basins (Tab. 1) using Multiple Regression on Distance Matrices (i.e., ordinary MRM based on Gaussian error distribution; i.e., MRM_lm) (Lichstein, 2007), which are equivalent to partial Mantel analysis. However, due to the properties of dissimilarity values (i.e., bounded between 0 and 1 and their relationship with predictors often involving non-linear trends), we also adopted the MRM approach based on Generalized Linear Models with Binomial error distribution (i.e., MRM_glm, performed with ecodist package and MRM function), as MRM_glm models appear more suitable for our response variables. As results obtained using the two approaches were very similar, we chose to provide both of them for transparency. These analyses allow us to estimate the independent effects of predictors current, historical and spatial effects while controlling for those peculiarities. The statistical significance was assessed with permutation of the response matrix, using the frequency at which 3999 null simulated coefficients were higher and/or lower than the observed coefficients. All analyses and graphics were performed under R environment (R Development Core Team, 2022), modeling routines were implemented in the ecodist package (Goslee et al., 2007) and all codes are available as Appendix files and at https://github.com/msversutdias/AmazonFISHBetaDiversity.git.
Results
Over all pairwise combinations, the turnover component is higher than the nestedness for taxonomic (mean Taxoβsor = 0.66, range = 0.22–1.00, sd = 0.16; mean Taxoβsim = 0.51, 0.03–1.00, sd = 0.18; mean Taxoβsne = 0.15, 0.00–0.77, sd = 0.13) and phylogenetic metrics (mean Phyloβsor = 0.42, range = 0.13–0.95, sd = 0.15; mean Phyloβsim = 0.24, 0.01–0.69, sd = 0.11; mean Phyloβsne = 0.17, 0.00–0.89, sd = 0.16). The nestedness component was higher than the turnover component for trait beta diversity (mean Traitβsor = 0.38, range = 0.02–1.00, sd = 0.23; mean Traitβsim = 0.08, 0.00–0.70, sd = 0.09; mean Traitβsne = 0.30, 0.00–1.00, sd = 0.26).
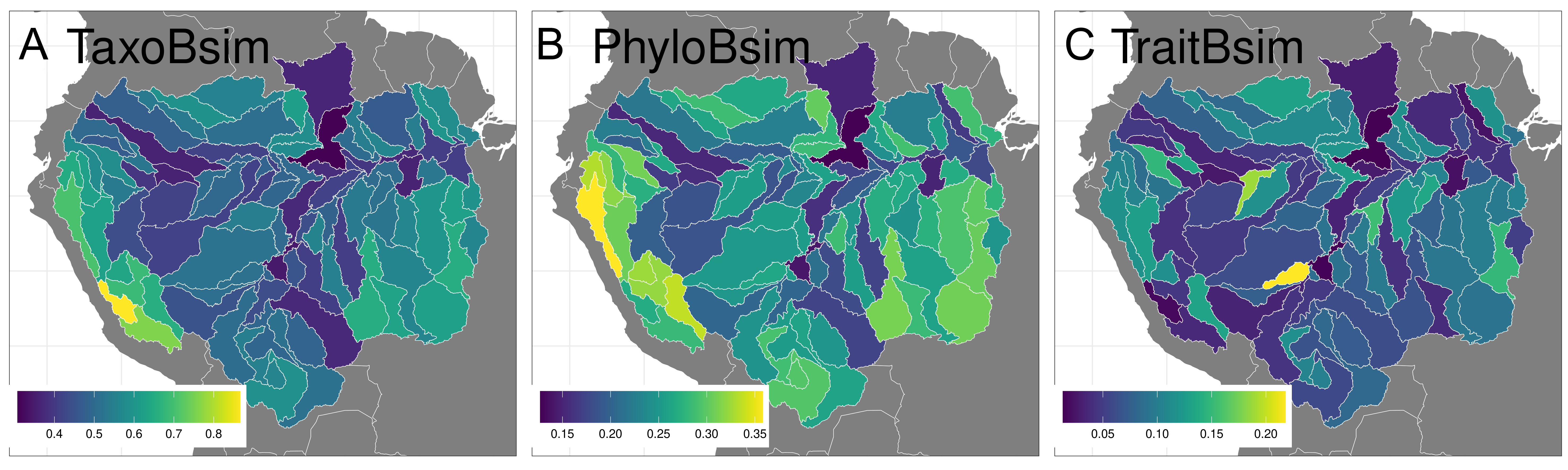
Geographical patterns in the three beta diversity facets. Mean taxonomic beta diversity among sub-basins (Taxoβsim= 0.51) is around two times higher than mean phylogenetic beta diversity (Phyloβsim=0.24) and six times higher than species traits beta diversity (Traitβsim=0.08). Using all pairwise distance values, all three beta diversity dimensions are positively and significantly related to each other, but correlations vary a lot among metrics (Tab. 2). Using mean pairwise dissimilarity values for each focal sub-basin (i.e., mean dissimilarity values over rows in matrices), Taxoβsim and Phyloβsim dimensions show coincident spatial patterns of low dissimilarity levels at the center (and along the river mainstem) of the Amazon Basin and higher dissimilarity at its periphery, Andean sub-basins displaying the highest dissimilarity values together with sub-basins located in the southeastern part of the basin (Figs. 1A, B). The Traitβsim pattern is more diffuse and heterogeneous across space, with highest assemblage dissimilarity values found, for instance, in sub-basins located in the northern part of the basin (e.g., upstream parts of the Negro River) (Fig. 1C; see also Fig. S6).
TABLE 2 | Pearson correlation among all pairwise indices of fish beta diversity in the Amazon 97 sub-drainages. All correlation values are significant at p<0.001.
| Taxoβsim | Phyloβsim | Traitβsim |
Taxoβsim | – | – | – |
Phyloβsim | 0.89 | – | – |
Traitβsim | 0.36 | 0.56 | – |
FIGURE 1| Spatial mapping of mean (a) taxonomic (Taxoβsim), (b) phylogenetic (Phyloβsim), and c) functional (Traitβsim) beta diversity dimensions. The values within sub-drainages are mean dissimilarity values calculated over all pairwise combinations targeting each of the 97 sub-basins analyzed. As they are all dissimilarities, all values are bounded between 0 and 1.
Factors explaining patterns in the three beta diversity facets. MRM_glm and ordinary MRM models show Pseudo-R2 and R2 ranging from 8 to 54% of explained variance. Our multiple MRM_glm models reveal a significant and prominent positive influence of geographic distance between sub-basins on the taxonomic and phylogenetic dimensions of beta diversity (Tab. 3; Fig. 2). In addition to geographic distance, Taxoβsim increases with dissimilarity in habitat harshness and habitat types (i.e., soil and vegetation cover), and decreases in Amazonian sub-basins having the same chemical water type. The model also suggests negative effects of the historical connection to the Pebas system, of marine incursions, and a positive effect of mean current temperature on Taxoβsim, but these last three effects are only marginally significant. Besides geographic distance, Phyloβsim is negatively related to dissimilarity in sampling effort, water type, and positively related to dissimilarity in habitat harshness. These effects suggest high phylogenetically dissimilar assemblages in sub-basins having distinct degree of harshness; whereas phylogenetically similar assemblages are found in sub-basins of the same water type and having high differences in sampling effort. Sampling effort and hydromorphological types significantly affect Traitβsim patterns. This suggests trait assemblage dissimilarity decreases with increasing difference in sampling effort, whereas trait dissimilarity is high when differences in rivers’ hydromorphological type increase between sub-basins.
TABLE 3 | Observed coefficient estimates from Multiple Regression on distance Matrices (MRM) models using Binomial (MRM_glm) and Gaussian (MRM_lm) error distribution on taxonomic, phylogenetic and trait beta diversity from the Amazon River basin. Pseudo-R2 is defined as (Null Deviance – Residual Deviance) / Null deviance. **p< 0.01; *p< 0.05; ¶ 0.05<p<0.10.
| MRM_glm | MRM_lm | ||||
Dissimilarity predictors | Taxoβsim | Phyloβsim | Traitβsim | Taxoβsim | Phyloβsim | Traitβsim |
(Intercept) | 0.07** | -1.16** | -2.51** | 0.51** | 0.24** | 0.08** |
D_Energy | -0.16 | -0.21 | -0.39 | -0.04 | -0.04¶ | -0.03 |
D_Water | -0.02 | 0.02 | 0.11 | 0.00 | 0.00 | 0.01 |
D_Temp | 0.37¶ | 0.15 | -0.14 | 0.08* | 0.03 | -0.01 |
D_HabitatSize | -0.06 | 0.05 | 0.06 | -0.01 | 0.01 | 0.00 |
D_HabitatType | 0.15* | 0.09 | 0.02 | 0.04** | 0.02¶ | 0.00 |
D_HabitatHarsh | 0.40** | 0.25** | 0.15 | 0.09** | 0.05** | 0.01 |
D_Position | 0.14 | 0.03 | 0.04 | 0.03 | 0.01 | 0.01 |
D_HabitatFragment | 0.02 | -0.02 | -0.13 | 0.00 | 0.00 | -0.01 |
D_HabitatWaterCol | -0.20** | -0.12* | -0.07 | -0.05** | -0.02** | -0.01 |
D_SamplEff | -0.07 | -0.25** | -0.58** | -0.02 | -0.04** | -0.04** |
D_MarineInc | -0.24¶ | -0.13 | -0.29 | -0.06* | -0.02 | -0.02 |
D_CurrentLGM | 0.17 | 0.13 | 0.22 | 0.04* | 0.02¶ | 0.01 |
D_PastConnect | -0.22¶ | -0.13 | -0.09 | -0.05* | -0.02¶ | -0.01 |
D_HydromorphoType | -0.03 | 0.03 | 0.16* | -0.01 | 0.01 | 0.01* |
D_Spatial | 0.54** | 0.40** | 0.20 | 0.13** | 0.07** | 0.01 |
Null deviance | 675.58 | 289.24 | 452.78 | – | – | – |
Residual Deviance | 324.39 | 186.92 | 409.81 | – | – | – |
Pseudo-R2 (%) | 52.00 | 35.37 | 9.49 | – | – | – |
F statistic | – | – | – | 364.50** | 174.88** | 26.23** |
R2 (%) | – | – | – | 54.00** | 36.00** | 8.00** |
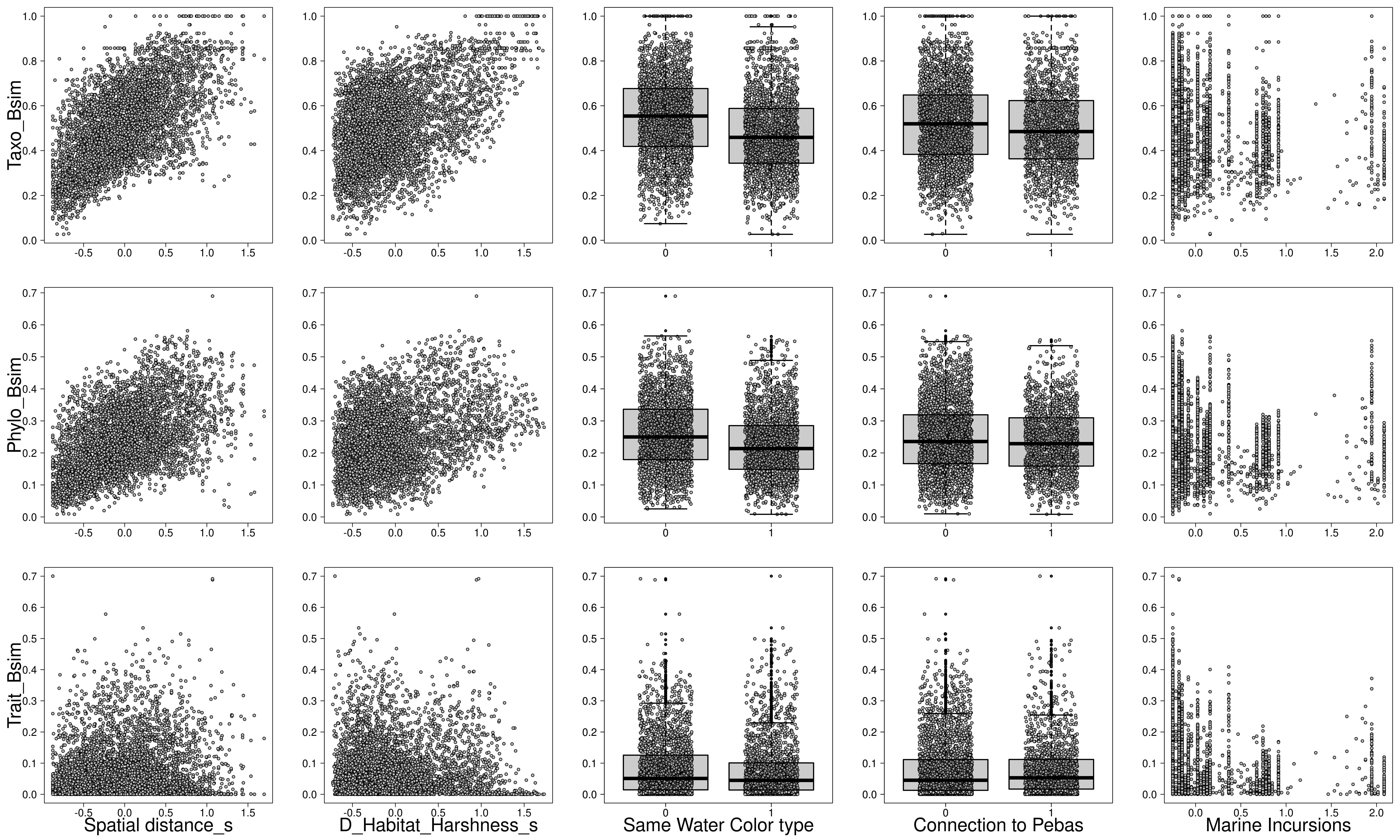
FIGURE 2| Relationships between fish dissimilarity facets from the Amazon sub-basins pairs and the most important predictors identified in Tab. 2.
Ordinary MRMs (MRM_lm) for the three beta diversity dimensions retain the same variables as MRM_glm but also highlight supplementary significant effects of historical connection to the Pebas system, of marine incursions (Fig. 2), Quaternary (D_CurrentLGM) and differences in current temperature (D_Temp) on sub-basins taxonomic dissimilarity (Taxoβsim). Assemblages in two sub-basins connected to the Pebas system are overall less taxonomically dissimilar (more similar) than their counterparts, while sub-basins having been subject to increasing levels of past marine incursions are overall less dissimilar to other sub-basins. Phyloβsim is related to the same predictors detected with MRM_glm (D_HabitatHarsh, D_HabitatWaterCol, D_SamplEff), but D_CurrentLGM and D_PastConnect also show marginally significant effects similar to Taxoβsim (Tab. 3). Sampling effort and riverine hydromorphology were detected again as the only significant predictors for Traitβsim (Tab. 3). Results obtained are similar when using a recently published Neotropical freshwater fish phylogeny (Cassemiro et al., 2023) instead of the one proposed by Rabosky et al. (2018) (Tabs. S7, S8) or when excluding pairwise dissimilarity from the 9 under-sampled sub-basins (Tab. S9). See also results for βsne component (Tab. S10).
Discussion
By analyzing, for the first time, three fish beta diversity dimensions among 97 sub-basins covering the whole Amazon Basin, our study unravels the role of several drivers and associated processes that act together to shape diversity patterns in this hyperdiverse basin.
A leading effect of dispersal limitation by geographic distance on taxonomic and phylogenetic beta diversity patterns. Regardless of the models retained (i.e., MRM_glm or MRM_lm), taxonomic (Taxoβsim) and phylogenetic beta diversity (Phyloβsim) dimensions are most-strongly related to geographic distance. This result is controlled for and independent of the other potential drivers used in the modeling, suggesting that species dispersal limitation by space strongly influences patterns of assemblage turnover among sub-basins as already detected for fish in other regions (Albert et al., 2011; Craig et al., 2020) and for several other taxonomic groups (Dambros et al., 2016, 2020; Fluck et al., 2020). Despite the fact that riverine fishes, with the exception of migratory species, are considered poor dispersers (Rodríguez, 2002; Comte, Olden, 2018), the Amazonian fish fauna is in addition largely dominated by small-bodied species evenly distributed among the richest families (Dagosta, Pinna, 2019; Albert et al., 2020). Given the large spatial grain used in our study (i.e., sub-basins), the huge geographical area of the Amazon Basin (~6 million km2, excluding the Tocantins Basin), and considering that small-sized fishes display lower dispersal abilities than their larger counterparts (Jenkins et al., 2007; Radinger, Wolter, 2013; Giam, Olden, 2018), it is not surprising that dispersal limitation by geographic distance strongly drives the variation in these two beta diversity facets (Rodrigues‐Filho et al., 2018). Moreover, as the metric Phyloβsim is sensitive to turnover among recently diverged lineages (Leprieur et al., 2012) and two times lower than Taxoβsim, the increase in assemblage phylogenetic dissimilarity with spatial distance also suggests recent allopatric diversification processes, rather than adaptive radiation in distant sub-basins and maintenance of these neo-endemic species in these sub-basins due to their usually smaller body size and consequent limited dispersal capacities (Tedesco et al., 2012).
According to metacommunity theory (Leibold, Chase, 2018), the spatial distribution of assemblages in a region is determined by the rate of dispersal of organisms and the environmental heterogeneity of habitat patches (Ricklefs, 2007). Beta diversity is often thought to be driven by environmental heterogeneity (Soininen et al., 2018) because different conditions among patches lead to distinct species using different niche opportunities due to their distinct habitat requirements (i.e., environmental filtering, Chase, Leibold, 2003). However, under the neutral model (Hubbell, 2001), dispersal limitation restricts species occurrence in different patches, resulting in spatially structured beta diversity patterns independent of environmental distance (Leibold, Chase, 2018). In practice, the relative importance of environmental conditions and dispersal as factors affecting species geographic ranges shifts from stronger environmental filtering to stronger dispersal limitation while changing from smaller to larger regional scales (Antonelli et al., 2018; Benone et al., 2020; Yunoki, 2023). Considering the large spatial grain involved (sub-basins >20,000 km2), that most Amazonian fishes are presumably poor dispersers (see above) and the high taxonomic beta diversity among sub-basins, we argue that at this spatial grain, dispersal limitation contributes strongly to the formation of Amazonian fish species assemblages (Albert et al., 2020), leading assemblages dissimilarity to increase with increasing geographic distance, and this independently of any environmental distance. In contrast, we found only a slightly positive influence of current temperature and habitat types on the Taxoβsim dimension, which may come from the fact that the analyzed sub-basins, given their size, broadly include most climatic conditions and types of habitat found in the Amazon Basin (Jézéquel et al., 2022).
Environmental filtering through habitat types and habitat harshness also drives taxonomic, phylogenetic and to a lesser extent functional beta diversity patterns. Besides geographic distance, our models depict a significant effect of habitat harshness (i.e., high elevation streams/rivers with steep gradients, cold climatic conditions and high hydrological variability) on assemblage taxonomic and phylogenetic turnover among sub-basins. This environmental harshness strongly constrains dispersal (Oberdorff et al., 2019) and drives deterministic selection over evolutionary times, leading to a reduced number of species filtered from the regional species pool, and possessing biological traits that allow them to cope with these specific environmental conditions (Datry et al., 2016). Consequently, in sub-basins harboring these habitats (mainly in the Andes and cratonic shields), fish assemblages display more distinct species composition and distinct lineages compared to other sub-basins (Fig. S6), leading to higher taxonomic and phylogenetic turnover between pairs of sub-basins. Assemblage composition changes produced by environmental harshness have previously been noticed locally in the Amazon Basin (e.g., Datry et al., 2016; Herrera-R et al., 2020). High taxonomic beta diversity is also found among sub-basins draining different types of soil and habitat cover, suggesting that these habitat types exercise a significant filtering process on the compositional structure of fish assemblages (Lo et al., 2020), usually affecting differentially species depending on their biological traits (e.g., trophic traits; Arantes et al., 2017).
We also find high taxonomic and phylogenetic beta diversity among sub-basins draining distinct water chemistry types, independently of the other considered drivers. According to our dataset, 83 genera and 258 species are found exclusively in whitewaters; 59 genera and 342 species are found exclusively in clearwaters;and 39 genera and 150 species are found exclusively in blackwaters. This suggests that differences in biogeochemical water types between sub-basins of the Amazon River act as filters to species dispersal and may also provide effective barriers to gene flow between populations leading to speciation processes and finally accumulation of distinctive fish assemblages between sub-basins. Both processes, i.e., filter to dispersal (e.g., Saint-Paul et al., 2000; Winemiller et al., 2008; Yunoki, Velasco, 2016; Bogotá-Gregory et al., 2020; Borges et al., 2023) and regulation of genetic exchanges between populations (e.g., Cooke et al., 2012; Beheregaray et al., 2015; Pires et al., 2018), have already been highlighted for fish assemblages inhabiting different water color types of the Amazon Basin.
Finally, one of our models also highlights a weak, though significant, positive effect of hydro-geomorphological river types on assemblages functional (species morphological traits) beta diversity. This pattern suggests that rivers’ hydro-geomorphology do act as an environmental filter, influencing the morphological trait distribution (i.e., the structure of the morphological space) of sub-basins fish assemblages through variations in habitat structural complexity (e.g., Lamouroux et al., 2002; Bower et al., 2021), and independently of any phylogenetic signal (i.e., morphological traits divergence among assemblages occupying habitats of distinct hydromorphological conditions). However, as previously said, the effect noticed is rather weak, despite a morphologically hyperdiverse Neotropical freshwater fish fauna (especially the Amazonian fauna) (Toussaint et al., 2016). The fact that our functional beta diversity metric (exclusively based on morphological traits) showed very low mean values and very low spatial variability (Fig. 1) may be partly due to the large grain size used in our study but is consistent with earlier findings obtained for tropical reef fishes (Maxwell et al., 2022). On the other hand, the Amazonian ichthyofauna has ancient origins, and many clades of Amazonian fishes had achieved modern phenotypes by the early Neogene [~23 million years (Ma) ago], when the aquatic system was mostly fluvio-lacustrine (Hoorn et al., 2010; Albert et al., 2020). It is thus likely that most of the diversification of species forms has been preferentially driven in this enduring, stable environment. The establishment of the transcontinental Amazon River during the last 10 Ma due to the uplift of the Andes, did not change drastically hydrological constraints for fishes as most of the basin (except the Andean Foreland) is represented by lowland areas with a flat topography (Jézéquel et al., 2022). In that, our results may plead for an overall historical and current non-binding physical environment for species morphological adaptation in the basin and suggest that the majority of morphological traits are shared among sub-basin assemblages. Furthermore, within the whole Amazon Basin, the trait composition of species-poor sub-basin assemblages is found to be nested within the trait composition of species-rich ones, as βsne was much more influential than βsim in explaining overall beta diversity patterns of functional (morphological) traits (see results).
Historical contingencies also matter for explaining taxonomic and phylogenetic beta diversity patterns. Independently of the effects of other considered drivers, assemblages in sub-basins historically connected to the Pebas system tend to be overall less taxonomically and phylogenetically dissimilar than their counterparts. This result is in phase with the well accepted historical hypothesis proposing the presence of a megawetland known as the Pebas system occupying western Amazonian during the Miocene and separated from a much smaller fluvial eastern Amazonia system by the Purus Arch (Hoorn et al., 2010; Shephard et al., 2010). Isolation of these two systems during millions of years before the Amazon River assumed its modern course toward the Atlantic (Campbell et al., 2006; Hoorn et al., 2010; Latrubesse et al., 2010; Oberdorff et al., 2019) may have generated two centers of diversification of the current Amazonian fish fauna by favoring independent diversification processes resulting in a pattern of high phylogenetic and high taxonomic beta dissimilarities between sub-basins located on both sides of this historical barrier.
A marginally significant (MRM_glm) or significant (MRM_lm) negative effect of Pleistocene marine incursion (<1 Ma ago, magnitude ~25 m) (Miller et al., 2005) on sub-basins taxonomic beta diversity patterns was also revealed by our models; fish assemblages in sub-basins having experienced increasing marine incursion (measured here as the percentage of area of sub-basins having suffered from marine incursions) are overall less dissimilar to other sub-basins. A parsimonious hypothesis is that this recent marine incursion has eliminated freshwater habitats in the low-lying areas of the lower and central Amazon (see also fig. S4 from Oberdorff et al., 2019), leading to populations extirpation of lowland freshwater fish species that homogenized fish assemblages in these concerned sub-basins. This last hypothesis is reinforced by the significant negative relationship we found between the nestedness component of taxonomic beta diversity and our Pleistocene marine incursion variable (see Tab. S10).
At last, one of our two models (MRM lm) highlights a supplementary positive effect of Quaternary climate variability (LGM) on sub-basins taxonomic and phylogenetic beta diversity patterns (also only marginally significant for the latter), suggesting that assemblages in sub-basins having undergone strong climatic variations since the LGM are more taxonomically and phylogenetically distant than their counterparts. This different history of climate evolution between sub-basins may have increased extinction rates in sub-basins where climatic variations were high (Cheng et al., 2013) and, on the contrary, increased speciation rates in sub-basins having experienced stable climatic conditions through time (Tedesco et al., 2012).
A significant effect of sampling effort on the functional and phylogenetic facets of beta diversity. After controlling for all other factors, our models highlight that both functional and phylogenetic assemblage dissimilarities decrease with increasing difference in sampling effort between sub-basins. A previous study using the same fish species occurrence database already noticed a positive and significant effect of sampling effort on our sub-basins species richness (Oberdorff et al., 2019). Together, these results support the view that assemblages in under-sampled sub-basins most often represent a subset of species found in better sampled sub-basins (i.e., species found in under-sampled areas are usually the most widespread ones, decreasing functional and phylogenetic sub-basins assemblage dissimilarities). Sampling gaps are pervasive in the Amazon, irrespective of the studied biological group, and are overall explained by difficulties to access some regions and/or remoteness of these regions from research infrastructure (Carvalho et al., 2023; Herrera-R et al., 2023).
To conclude, our large-scale analysis of spatial variation in the fish species assemblages inhabiting the Amazon sub-drainages provides insight into the processes structuring present-day fish assemblages. By analyzing three beta diversity facets (i.e., taxonomic, phylogenetic and functional), our results indicate that, at the spatial grain (97 sub-basins) and geographical extent (whole Amazon Basin) of this study, dispersal limitation by geographic distance is the major process shaping taxonomic and/or phylogenetic differences in fish assemblages composition between sub-drainages. However, we further noticed that taxonomic and phylogenetic beta diversity patterns also arise due to contemporary environmental constraints (i.e., environmental filtering) and to historical events, whereas functional turnover is only weakly linked to sampling effort and current sub-basins hydro-geomorphology. Finally, our results show that the Amazon Basin still suffers from incomplete knowledge on species spatial distribution, biological traits and phylogenetic information even though we used the most complete and up-to-date information currently available for these 3 biodiversity facets (Rabosky et al., 2018; Jézéquel et al., 2020a; Brosse et al., 2021; Cassemiro et al., 2023). There is thus an urgent need to address these gaps by increasing field surveys in currently under-sampled areas (Jézéquel et al., 2020b; Carvalho et al., 2023).
Acknowledgments
We thank Leandro Duarte for his advice on phylogenetic metrics. This paper is dedicated to J. Maldonado-Ocampo, who passed away during a fishing trip in the Río Vaupés (Colombia). We thank the Ecosystem Lab/UnB for cluster computation facilities.
References
Albert JS, Carnaval AC, Flantua SGA, Lohmann LG, Ribas CC, Riff D et al. Human impacts outpace natural processes in the Amazon. Science. 2023; 379(6630):eabo5003. https://doi.org/10.1126/science.abo5003
Albert JS, Carvalho TP, Petry P, Holder MA, Maxime EL, Espino J et al. Aquatic biodiversity in the Amazon: habitat specialization and geographic isolation promote species richness. Animals. 2011; 1(2):205–41. https://doi.org/10.3390/ani1020205
Albert JS, Tagliacollo VA, Dagosta F. Diversification of Neotropical freshwater fishes. Annu Rev Ecol Evol Syst. 2020; 51(1):27–53. https://doi.org/10.1146/annurev-ecolsys-011620-031032
Antonelli A, Ariza M, Albert J, Andermann T, Azevedo J, Bacon C et al. Conceptual and empirical advances in Neotropical biodiversity research. PeerJ. 2018 ; 6:e5644. https://doi.org/10.7717/peerj.5644
Arantes CC, Winemiller KO, Petrere M, Castello L, Hess LL, Freitas CE. Relationships between forest cover and fish diversity in the Amazon River floodplain. J Appl Ecol. 2017; 55(1):386–95. https://doi.org/10.1111/1365-2664.12967
Barton PS, Cunningham SA, Manning AD, Gibb H, Lindenmayer DB, Didham RK. The spatial scaling of beta diversity. Glob Ecol Biogeogr. 2013; 22(1):1–11. https://doi.org/10.1111/geb.12031
Baselga A. The relationship between species replacement, dissimilarity derived from nestedness, and nestedness. Glob Ecol Biogeogr. 2012; 21(12):1223–32. https://doi.org/10.1111/j.1466-8238.2011.00756.x
Baselga A. Partitioning the turnover and nestedness components of beta diversity. Glob Ecol Biogeogr. 2010; 19(1):134–43. https://doi.org/10.1111/j.1466-8238.2009.00490.x
Baselga A, Orme CDL. betapart: an R package for the study of beta diversity. Methods Ecol Evol. 2012; 3(5):808–12. https://doi.org/10.1111/j.2041-210X.2012.00224.x
Beheregaray LB, Cooke GM, Chao NL, Landguth EL. Ecological speciation in the tropics: insights from comparative genetic studies in Amazonia. Front Genet. 2015; 5:477. https://doi.org/10.3389/fgene.2014.00477
Benone NL, Leal CG, Santos LL, Mendes TP, Heino J, Montag LFA. Unravelling patterns of taxonomic and functional diversity of Amazon stream fish. Aquat Sci. 2020; 82(4):75. https://doi.org/10.1007/s00027-020-00749-5
Bogotá-Gregory JD, Lima FCT, Correa SB, Silva-Oliveira C, Jenkins DG, Ribeiro FR et al. Biogeochemical water type influences community composition, species richness, and biomass in megadiverse Amazonian fish assemblages. Sci Rep. 2020; 10(1):15349. https://doi.org/10.1038/s41598-020-72349-0
Borges SH, D’Aquino DD, Cruz MV, Souza RF. Turnover in fish species composition is related to water colour of Amazonian rivers. J Trop Ecol. 2023; 39:e7. https://doi.org/10.1017/S0266467422000451
Boschman LM, Carraro L, Cassemiro FAS, Vries J, Altermatt F, Hagen O et al. Freshwater fish diversity in the western Amazon basin shaped by Andean uplift since the Late Cretaceous. Nat Ecol Evol. 2023; 7(12):2037–44. https://doi.org/10.1038/s41559-023-02220-8
Bower LM, Saenz DE, Winemiller KO. Widespread convergence in stream fishes. Biol J Linn Soc. 2021; 133(3):863–79. https://doi.org/10.1093/biolinnean/blab043
Brosse S, Charpin N, Su G, Toussaint A, Herrera-R GA, Tedesco PA et al. FISHMORPH: A global database on morphological traits of freshwater fishes. Glob Ecol Biogeogr. 2021; 30(12):2330–36. https://doi.org/10.1111/geb.13395
Brown JH. Why are there so many species in the tropics? J Biogeogr. 2014; 41(1):8–22. https://doi.org/10.1111/jbi.12228
Brum FT, Graham CH, Costa GC, Hedges SB, Penone C, Radeloff VC et al. Global priorities for conservation across multiple dimensions of mammalian diversity. PNAS. 2017; 114(29):7641–46. https://doi.org/10.1073/pnas.1706461114
Cadotte MW, Hamilton MA, Murray BR. Phylogenetic relatedness and plant invader success across two spatial scales. Divers Distrib. 2009; 15(3):481–88. https://doi.org/10.1111/j.1472-4642.2009.00560.x
Caldas B, Thieme ML, Shahbol N, Coelho ME, Grill G, Van Damme PA et al. Identifying the current and future status of freshwater connectivity corridors in the Amazon Basin. Conserv Sci Pract. 2023; 5(1):e12853. https://doi.org/10.1111/csp2.12853
Campbell KE, Frailey CD, Romero-Pittman L. The Pan-Amazonian Ucayali Peneplain, late Neogene sedimentation in Amazonia, and the birth of the modern Amazon River system. Palaeogeogr Palaeoclimatol Palaeoecol. 2006; 239(1–2):166–219. https://doi.org/10.1016/j.palaeo.2006.01.020
Carvalho RL, Resende AF, Barlow J, França FM, Moura MR, Maciel R et al. Pervasive gaps in Amazonian ecological research. Curr Biol. 2023; 33(13):R1–R3. https://doi.org/10.1016/j.cub.2023.06.077
Cassemiro FAS, Albert JS, Antonelli A, Menegotto A, Wüest RO, Cerezer F et al. Landscape dynamics and diversification of the megadiverse South American freshwater fish fauna. PNAS. 2023; 120(2):e2211974120. https://doi.org/10.1073/pnas.2211974120
Chase JM, Leibold MA. Ecological niches: linking classical and contemporary approaches. Chicago: University of Chicago Press; 2003.
Cheng H, Sinha A, Cruz FW, Wang X, Edwards RL, d’Horta FM et al. Climate change patterns in Amazonia and biodiversity. Nat Commun. 2013; 4:1411. https://doi.org/10.1038/ncomms2415
Chessman BC, Fryirs KA, Brierley GJ. Linking geomorphic character, behaviour and condition to fluvial biodiversity: implications for river management. Aquat Conserv. 2006; 16(3):267–88. https://doi.org/10.1002/aqc.724
Comte L, Olden JD. Evidence for dispersal syndromes in freshwater fishes. Proc Biol Sci. 2018; 285(1871):20172214. https://doi.org/10.1098/rspb.2017.2214
Cooke GM, Chao NL, Beheregaray LB. Marine incursions, cryptic species and ecological diversification in Amazonia: the biogeographic history of the croaker genus Plagioscion (Sciaenidae). J Biogeogr. 2012; 39(4):724–38. https://doi.org/10.1111/j.1365-2699.2011.02635.x
Correa SB, Coronado-Franco KV, Jézéquel C, Cantarute Rodrigues A, Evans KO, Granger JJ et al. Floodplain forests drive fruit-eating fish diversity at the Amazon Basin-scale. PNAS. 2025; 122(3):e2414416122. https://doi.org/10.1073/pnas.2414416122
Couto TBA, Jenkins CN, Beveridge CF, Heilpern SA, Herrera-R GA, Piland NC et al. Translating science into actions to conserve Amazonian freshwaters. Conserv Sci Pract. 2024; 6(11):e13241. https://doi.org/10.1111/csp2.13241
Craig JM, Carvalho TP, Chakrabarty P, Derouen V, Ortega H, Petry P et al. Using community phylogenetics to assess phylogenetic structure in the Fitzcarrald region of Western Amazonia. Neotrop Ichthyol. 2020;18(2):e200004. https://doi.org/10.1590/1982-0224-2020-0004
Dagosta FCP, de Pinna M. Biogeography of Amazonian fishes: deconstructing river basins as biogeographic units. Neotrop Ichthyol. 2017; 15(3):e170034. https://doi.org/10.1590/1982-0224-20170034
Dagosta FCP, de Pinna M. The fishes of the Amazon: distribution and biogeographical patterns, with a comprehensive list of species. Am Mus Novit. 2019; 431:1–163. https://doi.org/10.1206/0003-0090.431.1.1
Dagosta FCP, de Pinna M, Peres CA, Tagliacollo VA. Existing protected areas provide a poor safety-net for threatened Amazonian fish species. Aquat Conserv. 2021; 31(5):1167–89. https://doi.org/10.1002/aqc.3461
Dambros C, Zuquim G, Moulatlet GM, Costa FRC, Tuomisto H, Ribas CC et al. The role of environmental filtering, geographic distance and dispersal barriers in shaping the turnover of plant and animal species in Amazonia. Biodivers Conserv. 2020; 29(13):3609–34. https://doi.org/10.1007/s10531-020-02040-3
Dambros CS, Morais JW, Azevedo RA, Gotelli NJ. Isolation by distance, not rivers, control the distribution of termite species in the Amazonian rain forest. Ecography. 2016; 40(10):1242–50. https://doi.org/10.1111/ecog.02663
Datry T, Melo AS, Moya N, Zubieta J, Barra ED, Oberdorff T. Metacommunity patterns across three Neotropical catchments with varying environmental harshness. Freshw Biol. 2016; 61(3):277–92. https://doi.org/10.1111/fwb.12702
Dias MS, Cornu J-F, Oberdorff T, Lasso CA, Tedesco PA. Natural fragmentation in river networks as a driver of speciation for freshwater fishes. Ecography. 2013; 36(6):683–89. https://doi.org/10.1111/j.1600-0587.2012.07724.x
Dias MS, Oberdorff T, Hugueny B, Leprieur F, Jézéquel C, Cornu J-F et al. Global imprint of historical connectivity on freshwater fish biodiversity. Ecol Lett. 2014; 17(9):1130–40. https://doi.org/10.1111/ele.12319
Dormann CF, Elith J, Bacher S, Buchmann C, Carl G, Carré G et al. Collinearity: a review of methods to deal with it and a simulation study evaluating their performance. Ecography. 2012; 36(1):27–46. https://doi.org/10.1111/j.1600-0587.2012.07348.x
Fitzpatrick MC, Sanders NJ, Normand S, Svenning J-C, Ferrier S, Gove AD et al. Environmental and historical imprints on beta diversity: insights from variation in rates of species turnover along gradients. Proc Biol Sci. 2013; 280(1768):20131201. https://doi.org/10.1098/rspb.2013.1201
Flecker AS, Shi Q, Almeida RM, Angarita H, Gomes-Selman JM, García-Villacorta R et al. Reducing adverse impacts of Amazon hydropower expansion. Science. 2022; 375(6582):753–60. https://doi.org/10.1126/science.abj4017
Fluck IE, Cáceres N, Hendges CD, Brum MN, Dambros CS. Climate and geographic distance are more influential than rivers on the beta diversity of passerine birds in Amazonia. Ecography. 2020;43(6):860–68. https://doi.org/10.1111/ecog.04753
Frederico RG, Dias MS, Jézéquel C, Tedesco PA, Hugueny B, Zuanon J et al. The representativeness of protected areas for Amazonian fish diversity under climate change. Aquat Conserv. 2021; 31(5):1158–66. https://doi.org/10.1002/aqc.3528
Fritz SA, Rahbek C. Global patterns of amphibian phylogenetic diversity. J Biogeogr. 2012; 39(8):1373–82. https://doi.org/10.1111/j.1365-2699.2012.02757.x
Gelman A. Scaling regression inputs by dividing by two standard deviations. Stat Med. 2008; 27(15):2865–73. https://doi.org/10.1002/sim.3107
Giam X, Olden JD. Drivers and interrelationships among multiple dimensions of rarity for freshwater fishes. Ecography. 2018; 41(2):331–44. https://doi.org/10.1111/ecog.02946
Goslee SC, Urban DL. The ecodist package for dissimilarity-based analysis of ecological data. J Stat Softw. 2007; 22(7):1–19.
Graham CH, Fine PVA. Phylogenetic beta diversity: linking ecological and evolutionary processes across space in time. Ecol Lett. 2008; 11(12):1265–77. https://doi.org/10.1111/j.1461-0248.2008.01256.x
Grant EHC, Lowe WH, Fagan WF. Living in the branches: population dynamics and ecological processes in dendritic networks. Ecol Lett. 2007; 10(2):165–75. https://doi.org/10.1111/j.1461-0248.2006.01007.x
Guilherme DR, Pequeno PACL, Baccaro FB, Franklin E, Santos Neto CR, Souza JLP. Direct and indirect effects of geographic and environmental factors on ant beta diversity across Amazon basin. Oecologia. 2022; 198(1):193–203. https://doi.org/10.1007/s00442-021-05083-7
Herrera-R GA, Oberdorff T, Anderson EP, Brosse S, Carvajal-Vallejos FM, Frederico RG et al. The combined effects of climate change and river fragmentation on the distribution of Andean Amazon fishes. Glob Chang Biol. 2020; 26(10):5509–23. https://doi.org/10.1111/gcb.15285
Herrera-R GA, Tedesco PA, DoNascimiento C, Jézéquel C, Giam X. Accessibility and appeal jointly bias the inventory of Neotropical freshwater fish fauna. Biol Conserv. 2023; 284:110186. https://doi.org/10.1016/j.biocon.2023.110186
Hoorn C, Wesselingh FP, Steege HT, Bermudez MA, Mora A, Sevink J et al. Amazonia through time: Andean uplift, climate change, landscape evolution, and biodiversity. Science. 2010; 330(6006):927–31. https://doi.org/10.1126/science.1194585
Hubbell SP. The unified neutral theory of biodiversity and biogeography. Princeton: Princeton University Press; 2001.
Hugueny B, Oberdorff T, Tedesco PA. Community ecology of river fishes: a large-scale perspective. In: Gido KB, Jackson DA, editors. Community ecology of stream fishes: concepts, approaches, and techniques. Bethesda: American Fisheries Society; 2010. p.29–62.
Jenkins DG, Brescacin CR, Duxbury CV, Elliott JA, Evans JA, Grablow KR et al. Does size matter for dispersal distance? Glob Ecol Biogeogr. 2007; 16(4):415–25. https://doi.org/10.1111/j.1466-8238.2007.00312.x
Jézéquel C, Oberdorff T, Tedesco PA, Schmitt L. Geomorphological diversity of rivers in the Amazon Basin. Geomorphology. 2022; 400:108078. https://doi.org/10.1016/j.geomorph.2021.108078
Jézéquel C, Tedesco PA, Bigorne R, Maldonado-Ocampo JA, Ortega H, Hidalgo M et al. A database of freshwater fish species of the Amazon Basin. Sci Data. 2020a; 7(1):96. https://doi.org/10.1038/s41597-020-0436-4
Jézéquel C, Tedesco PA, Darwall W, Dias MS, Frederico RG, Hidalgo M et al. Freshwater fish diversity hotspots for conservation priorities in the Amazon Basin. Conserv Biol. 2020b; 34(4):956–65. https://doi.org/10.1111/cobi.13466
Kembel SW, Cowan PD, Helmus MR, Cornwell WK, Morlon H, Ackerly DD et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics. 2010; 26(11):1463–64. https://doi.org/10.1093/bioinformatics/btq166
Lamouroux N, Poff NL, Angermeier PL. Intercontinental convergence of stream fish community traits along geomorphic and hydraulic gradients. Ecology. 2002; 83(7):1792–807. https://doi.org/10.1890/0012-9658(2002)083[1792:ICOSFC]2.0.CO;2
Larsen S, Comte L, Filipe AF, Fortin M-J, Jacquet C, Ryser R et al. The geography of metapopulation synchrony in dendritic river networks. Ecol Lett. 2021; 24(4):791–801. https://doi.org/10.1111/ele.13699
Latrubesse EM, Cozzuol M, Silva-Caminha SAF da, Rigsby CA, Absy ML, Jaramillo C. The Late Miocene paleogeography of the Amazon Basin and the evolution of the Amazon River system. Earth Sci Rev. 2010; 99(3–4):99–124. https://doi.org/10.1016/j.earscirev.2010.02.005
Lehner B, Grill G. Global river hydrography and network routing: baseline data and new approaches to study the world’s large river systems. Hydrol Process. 2013; 27(15):2171–86. https://doi.org/10.1002/hyp.9740
Leibold MA, Chase JM. Metacommunity ecology. Princeton: Princeton University Press; 2018.
Leprieur F, Albouy C, Bortoli JD, Cowman PF, Bellwood DR, Mouillot D. Quantifying phylogenetic beta diversity: distinguishing between ‘true’ turnover of lineages and phylogenetic diversity gradients. PLoS ONE. 2012; 7(8):e42760. https://doi.org/10.1371/journal.pone.0042760
Leprieur F, Tedesco PA, Hugueny B, Beauchard O, Dürr HH, Brosse S et al. Partitioning global patterns of freshwater fish beta diversity reveals contrasting signatures of past climate changes. Ecol Lett. 2011; 14(4):325–34. https://doi.org/10.1111/j.1461-0248.2011.01589.x
Lichstein JW. Multiple regression on distance matrices: a multivariate spatial analysis tool. Plant Ecol. 2007; 188(2):117–31. https://doi.org/10.1007/s11258-006-9126-3
Lo M, Reed J, Castello L, Steel EA, Frimpong EA, Ickowitz A. The influence of forests on freshwater fish in the tropics: a systematic review. BioScience. 2020; 70(5):404–14. https://doi.org/10.1093/biosci/biaa021
Lu M, Vasseur D, Jetz W. Beta diversity patterns derived from island biogeography theory. Am Nat. 2019; 194(3):E52–E65. https://doi.org/10.1086/704181
Maire E, Grenouillet G, Brosse S, Villéger S. How many dimensions are needed to accurately assess functional diversity? Glob Ecol Biogeogr. 2015; 24(6):728–40. https://doi.org/10.1111/geb.12299
Maxwell MF, Leprieur F, Quimbayo JP, Floeter SR, Bender MG. Global patterns and drivers of beta diversity facets of reef fish faunas. J Biogeogr. 2022; 49(5):954–67. https://doi.org/10.1111/jbi.14349
Mazel F, Wüest RO, Lessard J-P, Renaud J, Ficetola GF, Lavergne S. Global patterns of β-diversity along the phylogenetic time-scale: the role of climate and plate tectonics. Glob Ecol Biogeogr. 2017; 26(10):1211–21. https://doi.org/10.1111/geb.12632
McGill B, Enquist B, Weiher E, Westoby M. Rebuilding community ecology from functional traits. Trends Ecol Evol. 2006; 21(4):178–85. https://doi.org/10.1016/j.tree.2006.02.002
McLean M, Stuart-Smith RD, Villéger S, Auber A, Edgar GJ, MacNeil MA. Trait similarity in reef fish faunas across the world’s oceans. PNAS. 2021; 118(12):e2012318118. https://doi.org/10.1073/pnas.2012318118
Miller KG, Kominz MA, Browning JV, Wright JD, Mountain GS, Katz ME. The Phanerozoic record of global sea-level change. Science. 2005; 310(5752):1293–98. https://doi.org/10.1126/science.1116412
Oberdorff T, Dias MS, Jézéquel C, Albert JS, Arantes CC, Bigorne R et al. Unexpected fish diversity gradients in the Amazon basin. Sci Adv. 2019; 5(9):eaav8681. https://doi.org/10.1126/sciadv.aav8681
Oksanen J, Simpson GL, Blanchet FG, Kindt R, Legendre P, Minchin PR. vegan: community ecology package. R package version; 2022.
Olden J, Keppler FW, Jardine TD, Boulêtreau S, Harrod C, Jackson MC et al. Looks can be deceiving: morphological traits are weakly associated with the isotopic niches of freshwater fishes. Glob Ecol Biogeogr. 2025; 34(9):e70124. https://doi.org/10.1111/geb.70124
Pelicice FM, Azevedo-Santos VM, Vitule JRS, Orsi ML, Lima Junior DP, Magalhães ALB. Neotropical freshwater fishes imperilled by unsustainable policies. Fish Fish. 2017; 18(6):1119–33. https://doi.org/10.1111/faf.12228
Penone C, Weinstein BG, Graham CH, Brooks TM, Rondinini C, Hedges SB. Global mammal beta diversity shows parallel assemblage structure in similar but isolated environments. Proc Biol Sci. 2016; 283(1837):20161028. https://doi.org/10.1098/rspb.2016.1028
Pires THS, Borghezan EA, Machado VN, Powell DL, Röpke CP, Oliveira C. Testing Wallace’s intuition: water type, reproductive isolation and divergence in an Amazonian fish. J Evol Biol. 2018; 31(6):882–92. https://doi.org/10.1111/jeb.13272
R Development Core Team. R: a language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2022. Available from: https://www.r-project.org/
Rabosky DL. Speciation rate and the diversity of fishes in freshwaters and the oceans. J Biogeogr. 2020; 47(6):1207–17. https://doi.org/10.1111/jbi.13839
Rabosky DL, Chang J, Title PO, Cowman PF, Sallan L, Friedman M. An inverse latitudinal gradient in speciation rate for marine fishes. Nature. 2018; 559(7714):392–95. https://doi.org/10.1038/s41586-018-0273-1
Radinger J, Wolter C. Patterns and predictors of fish dispersal in rivers. Fish Fish. 2013; 15(1):456–473. https://doi.org/10.1111/faf.12028
Reis RE, Albert JS, Dario FD, Mincarone MM, Petry P, Rocha LA. Fish biodiversity conservation in South America. J Fish Biol. 2016; 89(1):12–47. https://doi.org/10.1111/jfb.13016
Ricklefs RE. History and diversity: explorations at the intersection of ecology and evolution. Am Nat. 2007; 170(S2):S56–S70. https://doi.org/10.1086/519402
Ricklefs RE. A comprehensive framework for global patterns in biodiversity. Ecol Lett. 2004; 7(1):1–15. https://doi.org/10.1046/j.1461-0248.2003.00554.x
Ríos-Villamizar EA, Adeney JM, Piedade MTF, Junk WJ. New insights on the classification of major Amazonian river water types. Sustain Water Resour Manag. 2020; 6(5):1–16. https://doi.org/10.1007/s40899-020-00440-5
Rodrigues-Filho CAS, Leitão RP, Zuanon J, Sánchez-Botero JI, Baccaro FB. Historical stability promoted higher functional specialization and originality in Neotropical stream fish assemblages. J Biogeogr. 2018; 45(6):1345–54. https://doi.org/10.1111/jbi.13205
Rodríguez MA. Restricted movement in stream fish: the paradigm is incomplete, not lost. Ecology. 2002; 83(1):1–13. https://doi.org/10.1890/0012-9658(2002)083[0001:RMISFT]2.0.CO;2
Saint-Paul U, Zuanon J, Correa MAV, García M, Fabré NN, Berger U. Fish communities in central Amazonian white- and blackwater floodplains. Environ Biol Fishes. 2000; 57(3):235–50. https://doi.org/10.1023/A:1007699130333
Salgueiro L, Cassemiro FAS, Albert JS, Frederico RG, Hidalgo M, Hugueny B et al. Drivers of phylogenetic structure in Amazon freshwater fish assemblages. J Biogeogr. 2022; 49(2):310–23. https://doi.org/10.1111/jbi.14300
Sandel B, Arge L, Dalsgaard B, Davies RG, Gaston KJ, Sutherland WJ. The influence of late Quaternary climate-change velocity on species endemism. Science. 2011; 334(6056):660–64. https://doi.org/10.1126/science.1210173
Scott S, Rojas P, Vila I. Meristic and morphological differentiation of Orestias species (Teleostei; Cyprinodontiformes) from the southern Altiplano. Environ Biol Fishes. 2020; 103(8):939–51. https://doi.org/10.1007/s10641-020-00995-4
Shephard GE, Müller RD, Liu L, Gurnis M. Miocene drainage reversal of the Amazon River driven by plate-mantle interaction. Nat Geosci. 2010; 3:870–75. https://doi.org/10.1038/ngeo1017
Socolar JB, Gilroy JJ, Kunin WE, Edwards DP. How should beta-diversity inform biodiversity conservation? Trends Ecol Evol. 2016; 31:67–80. https://doi.org/10.1016/j.tree.2015.11.005
Soininen J, Heino J, Wang J. A meta-analysis of nestedness and turnover components of beta diversity across organisms and ecosystems. Glob Ecol Biogeogr. 2018; 27:96–109. https://doi.org/10.1111/geb.12660
Swenson NG, Anglada-Cordero P, Barone JA. Deterministic tropical tree community turnover: evidence from patterns of functional beta diversity along an elevational gradient. Proc Biol Sci. 2011; 278:877–84. https://doi.org/10.1098/rspb.2010.1369
Tedesco PA, Beauchard O, Bigorne R, Blanchet S, Buisson L, Conti L et al. A global database on freshwater fish species occurrence in drainage basins. Sci Data. 2017a; 4:17141. https://doi.org/10.1038/sdata.2017.141
Tedesco PA, Leprieur F, Hugueny B, Brosse S, Dürr HH, Beauchard O et al. Patterns and processes of global riverine fish endemism. Glob Ecol Biogeogr. 2012; 21:977–87. https://doi.org/10.1111/j.1466-8238.2011.00749.x
Tedesco PA, Paradis E, Lévêque C, Hugueny B. Explaining global-scale diversification patterns in actinopterygian fishes. J Biogeogr. 2017b; 44:773–83. https://doi.org/10.1111/jbi.12905
Tisseuil C, Cornu J-F, Beauchard O, Brosse S, Darwall W, Holland R et al. Global diversity patterns and cross-taxa convergence in freshwater systems. J Anim Ecol. 2013; 82:365–76. https://doi.org/10.1111/1365-2656.12018
Tonkin JD, Altermatt F, Finn DS, Heino J, Olden JD, Pauls SU et al. The role of dispersal in river network metacommunities: patterns, processes, and pathways. Freshw Biol. 2018; 63:141–63. https://doi.org/10.1111/fwb.13037
Toussaint A, Charpin N, Brosse S, Villéger S. Global functional diversity of freshwater fish is concentrated in the Neotropics while functional vulnerability is widespread. Sci Rep. 2016; 6:22125. https://doi.org/10.1038/srep22125
Troia MJ, McManamay RA. Filling in the GAPS: evaluating completeness and coverage of open-access biodiversity databases in the United States. Ecol Evol. 2016; 6:4654–69. https://doi.org/10.1002/ece3.2225
Venticinque E, Forsberg B, Barthem R, Petry P, Hess L, Mercado A et al. An explicit GIS-based river basin framework for aquatic ecosystem conservation in the Amazon. Earth Syst Sci Data. 2016; 8:651–61. https://doi.org/10.5194/essd-8-651-2016
Villéger S, Brosse S, Mouchet M, Mouillot D, Vanni MJ. Functional ecology of fish: current approaches and future challenges. Aquat Sci. 2017; 79:783–801. https://doi.org/10.1007/s00027-017-0546-z
Villéger S, Grenouillet G, Brosse S, Baselga A. Decomposing functional β-diversity reveals that low functional β-diversity is driven by low functional turnover in European fish assemblages. Glob Ecol Biogeogr. 2013; 22:671–81. https://doi.org/10.1111/geb.12021
Weinstein BG, Tinoco B, Parra JL, Brown LM, McGuire JA, Stiles FG et al. Taxonomic, phylogenetic, and trait beta diversity in South American hummingbirds. Am Nat. 2014; 184:211–224. https://doi.org/10.1086/676991
Whittaker RH. Vegetation of the Siskiyou Mountains, Oregon and California. Ecol Monogr. 1960; 30:279–338. https://doi.org/10.2307/1943563
Winemiller KO, López-Fernández H, Taphorn DC, Nico LG, Duque AB. Fish assemblages of the Casiquiare River, a corridor and zoogeographical filter for dispersal between the Orinoco and Amazon basins. J Biogeogr. 2008; 35:1551–63. https://doi.org/10.1111/j.1365-2699.2008.01917.x
Wu L, Si X, Didham RK, Ge D, Ding P. Dispersal modality determines the relative partitioning of beta diversity in spider assemblages on subtropical land-bridge islands. J Biogeogr. 2017; 44:2121–31. https://doi.org/10.1111/jbi.13007
Yunoki T. In a niche-neutral continuum, a set of theoretical models in a metacommunity operates simultaneously in patchy habitats. Ecol Evol. 2023; 13:e9754. https://doi.org/10.1002/ece3.9754
Yunoki T, Velasco LT. Fish metacommunity dynamics in the patchy heterogeneous habitats of várzea lakes, turbid river channels and transparent clear and black water bodies in the Amazonian Lowlands of Bolivia. Environ Biol Fishes. 2016; 99:391–408. https://doi.org/10.1007/s10641-016-0481-1
Authors
![]() Murilo S. Dias1
Murilo S. Dias1 ![]() ,
, ![]() Céline Jézéque2,
Céline Jézéque2, ![]() Bernard Hugueny2,
Bernard Hugueny2, ![]() Fabien Leprieur3,
Fabien Leprieur3, ![]() Sébastien Brosse2,
Sébastien Brosse2, ![]() James Albert4,
James Albert4, ![]() Fernanda A. S. Cassemiro5,
Fernanda A. S. Cassemiro5, ![]() Renata G. Frederico6,
Renata G. Frederico6, ![]() Max Hidalgo7,
Max Hidalgo7, ![]() Hernan Ortega7,
Hernan Ortega7, ![]() Pablo A. Tedesco2,
Pablo A. Tedesco2, ![]() Gislene Torrente-Vilara8,
Gislene Torrente-Vilara8, ![]() Jansen Zuanon9,10,
Jansen Zuanon9,10, ![]() Rémy Bigorne2,
Rémy Bigorne2, ![]() Koen Martens11,
Koen Martens11, ![]() Astrid Acosta12,
Astrid Acosta12, ![]() Edwin Agudelo12,
Edwin Agudelo12, ![]() Soraya Barrera Maure13,
Soraya Barrera Maure13, ![]() Douglas A. Bastos9,
Douglas A. Bastos9, ![]() Juan Bogotá-Gregory14,
Juan Bogotá-Gregory14, ![]() Fernando G. Cabeceira14,
Fernando G. Cabeceira14, ![]() André L. C. Canto15,
André L. C. Canto15, ![]() Fernando M. Carvajal-Vallejos16,
Fernando M. Carvajal-Vallejos16, ![]() Lucélia N. Carvalho17,
Lucélia N. Carvalho17, ![]() Ariana Cella-Ribeiro18,
Ariana Cella-Ribeiro18, ![]() Raphaël Covain19,
Raphaël Covain19, ![]() Carlos DoNascimiento20,
Carlos DoNascimiento20, ![]() Carolina R. C. Dória21,
Carolina R. C. Dória21, ![]() Cleber Duarte9,
Cleber Duarte9, ![]() Efrem J. G. Ferreira9,
Efrem J. G. Ferreira9, ![]() André V. Galuch9,22,
André V. Galuch9,22, ![]() Tommaso Giarrizzo23,
Tommaso Giarrizzo23, ![]() Rafael P. Leitão6,
Rafael P. Leitão6, ![]() John G. Lundberg24,
John G. Lundberg24, ![]() Mabel Maldonado16,
Mabel Maldonado16, ![]() Luciano F. A. Montag25,
Luciano F. A. Montag25, ![]() Willian Massaharu Ohara26,
Willian Massaharu Ohara26, ![]() Tiago H. S. Pires9,
Tiago H. S. Pires9, ![]() Marc Pouilly27,
Marc Pouilly27, ![]() Saúl Prada-Pedreros28,
Saúl Prada-Pedreros28, ![]() Luiz J. de Queiroz29,
Luiz J. de Queiroz29, ![]() Lucia Rapp Py-Daniel9,
Lucia Rapp Py-Daniel9, ![]() Frank R. V. Ribeiro15,
Frank R. V. Ribeiro15, ![]() Raúl Ríos Herrera30,
Raúl Ríos Herrera30, ![]() Jaime Sarmiento13,
Jaime Sarmiento13, ![]() Leandro M. Sousa31,
Leandro M. Sousa31, ![]() Lis F. Stegmann9,32,
Lis F. Stegmann9,32, ![]() Jonathan Valdiviezo-Rivera33,
Jonathan Valdiviezo-Rivera33, ![]() Francisco Villa34,
Francisco Villa34, ![]() Takayuki Yunoki35,
Takayuki Yunoki35, ![]() Aaike de Wever36 and
Aaike de Wever36 and ![]() Thierry Oberdorff2
Thierry Oberdorff2
[1] Universidade de Brasília, Departamento de Ecologia, Programa de Pós-Graduação em Ecologia, 70910-900, Brasília, DF, Brazil. (MSD) msdias@unb.br (corresponding author).
[2] Centre de Recherche sur la Biodiversité et l’Environnement (CRBE), UMR 5300 CNRS, IRD276, INP, UT , Toulouse, France. (CJ) celine.jezequel@ird.fr, (BH) bernard.hugueny@ird.fr, (SB) sebastien.brosse@univ-tlse3.fr, (PAT) pablo.tedesco@ird.fr, (TO) thierry.oberdorff@ird.fr.
[3] MARBEC, Univ Montpellier, CNRS, Ifremer, IRD, 34095 Montpellier, France. (FL) fabien.leprieur@umontpellier.fr.
[4] University of Louisiana, Department of Biology, Lafayette, LA 70504-4245, USA. (JA) jalbert@louisiana.edu.
[5] Universidade Federal de Goiás, Departamento de Ecologia, Programa de Pós-Graduação em Ecologia e Evolução, 74690 900, Goiânia, GO, Brazil. (FASC) fernandacassemiro@gmail.com.
[6] Universidade Federal de Minas Gerais, Departamento de Genética, Ecologia e Evolução, 31270-901, Belo Horizonte, MG, Brazil. (RGF) renatafrederico@gmail.com, (RPL) ecorafa@gmail.com.
[7] Universidad Nacional Mayor de San Marcos, Departamento de Ictiología, Museo de Historia Natural, Lima 15081, Peru. (MH) mhidalgod@unmsm.edu.p, (HO) hortega.musm@gmail.com.
[8] Universidade Federal de São Paulo, Instituto do Mar, Campus Baixada Santista, 11070-100, Santos, SP, Brazil. (GTV) gtvilara@gmail.com.
[9] Instituto Nacional de Pesquisas da Amazônia, Coordenação de Biodiversidade, 69067-375, Manaus, AM Brazil. (JZ) jzuanon3@gmail.com, (DAB) avizdoug@gmail.com, (THSP) thspires@gmail.com, (LRP-D) luciarpd60@gmail.com, (EJGF) ferreira.efrem@gmail.com
[10] Universidade Santa Cecília (UNISANTA), Santos, SP, Brazil (Senior Visiting Researcher, PRAPG-CAPES).
[11] Royal Belgian Institute for Natural Sciences, Freshw Biol, B-1000 Brussels, Belgium. (KM) kmartens@naturalsciences.be.
[12] Instituto Amazónico de Investigaciones Científicas Sinchi, Colección Ictiológica de la Amazonia Colombiana, Leticia, Amazonas, 910001, Colombia. (JBG) juandbogota@gmail.com, (AA) astridacostasantos@gmail.com, (EA) eagudelo@sinchi.org.co.
[13] Museo Nacional de Historia Natural, Colección Boliviana de Fauna, La Paz, Bolivia. (SBM) sorayabarrera@gmail.com, (JS) jsarmientotavel@gmail.com
[14] Universidade Federal de Mato Grosso, Instituto de Biociências, 78060-900, Cuiabá, MT, Brazil. (FGC) fernando.cabeceira@gmail.com.
[15] Universidade Federal do Oeste do Pará, Instituto de Ciências e Tecnologia das Águas, 68040-050, Santarém, PA, Brazil. (ALCC) cantoandre@gmail.com, (FRVR) fraynner@yahoo.com.br.
[16] Universidad Mayor de San Simón, Unidad de Limnología y Recursos Acuáticos, Cochabamba, Bolivia. (FMCV) fmcvalle@yahoo.com, (MM) mabelmaldonado.m@fcyt.umss.edu.bo.
[17] Universidade Federal de Mato Grosso, Instituto de Ciências Naturais, Humanas e Sociais, 78550-728, Sinop, MT, Brazil. (LNC) carvalholn@yahoo.com.br.
[18] Instituto Chico Mendes de Conservação da Biodiversidade – ICMBio, 76887-000, Campo Novo de Rondônia, RO, Brazil (ACR) ariana.ribeiro@icmbio.gov.br.
[19] Muséum d’histoire naturelle de Genève, 1 route de Malagnou CH1208, Genève, Switzerland. (RC) raphael.covain@geneve.ch.
[20] Universidad de Antioquia, Medellín, Colombia. (CD) c.donascimiento@udea.edu.co.
[21] Universidade Federal de Rondônia, Departamento de Biologia, 76801-059, Porto Velho, RO, Brazil. (CRCD) carolinarcdoria@unir.br.
[22] Instituto Federal do Amazonas, Campus Manaus Centro, 69020-120, Manaus, Amazonas, Brazil. (AVG) andregaluch@ifam.edu.br.
[23] Instituto de Ciências do Mar (LABOMAR), Universidade Federal do Ceará (UFC), Fortaleza, Brazil. (TG) tgiarrizzo@gmail.com.
[24] Department of Ichthyology, The Academy of Natural Sciences of Drexel University, 1900 Benjamin Franklin Parkway, 19103-1195, Philadelphia, PA, USA. (JGL) jgl43@drexel.edu.
[25] Universidade Federal do Pará, Programa de Pós-Graduação em Ecologia, 66075-110, Belém, PA, Brazil. (LFAM) lfamontag@gmail.com.
[26] Universidade Federal do Amazonas, Departamento de Biologia, 69077-000, Manaus, AM, Brazil. (WMO) ohara@ufam.edu.br.
[27] Muséum National d’Histoire Naturelle, Laboratoire de Biologie des Organismes et Ecosystèmes Aquatiques, F-75005, Paris, France. (MP) marc.pouilly@ird.fr.
[28] Pontificia Universidad Javeriana, Laboratorio de Ictiología, Unidad de Ecología y Sistemática (UNESIS), Bogotá, Colombia. (SPP) saul.prada@javeriana.edu.co.
[29] Naturalis Biodiversity Center, 2333 CR Leiden, Netherlands. (LJQ) luiz.jardimdequeiroz@naturalis.nl.
[30] Instituto para la Investigación y la Preservación del Patrimonio Cultural y Natural, Cali, Colombia. (RRH) raulriosh@gmail.com.
[31] Universidade Federal do Pará, Laboratório de Ictiologia de Altamira, 68372-040, Altamira, PA, Brazil. (LMS) leandro.m.sousa@gmail.com.
[32] Museu Paraense Emílio Goeldi, Coordenação de Comunicação e Extensão, 66040-170, Belém-PA Brazil. (LFS) lisstegmann@museu-goeldi.br.
[33] Instituto Nacional de Biodiversidad, Quito, 170150, Ecuador. (JVR) jonathan.valdiviezo@biodiversidad.gob.ec
[34] Universidad del Tolima, Grupo de Investigación en Zoología, Ibagué, 730006299, Colombia. (FV) favilla@ut.edu.co.
[35] Universidad Autónoma del Beni, Centro de Investigación de Recursos Acuáticos, Trinidad, Bolivia. (TY) takayukiyunoki@yahoo.com.
[36] Research Institute for Nature and Forest, B-1000 Brussels, Belgium. (AW) aaike.dewever@inbo.be.
Authors’ Contribution 

Murilo S. Dias: Conceptualization, Formal analysis, Investigation, Methodology, Project administration, Software, Visualization, Writing-original draft, Writing-review and editing.
Céline Jézéquel: Data curation, Methodology, Project administration, Resources, Software, Validation, Writing-review and editing.
Bernard Hugueny: Conceptualization, Investigation, Supervision, Validation, Writing-review and editing.
Fabien Leprieur: Conceptualization, Formal analysis, Investigation, Methodology, Software, Supervision, Writing-original draft, Writing-review and editing.
Sébastien Brosse: Data curation, Funding acquisition, Methodology, Project administration, Resources, Supervision, Validation, Writing-review and editing.
James Albert: Investigation, Supervision, Validation, Writing-review and editing.
Fernanda A. S. Cassemiro: Investigation, Validation, Writing-original draft, Writing-review and editing.
Renata G. Frederico: Investigation, Validation, Writing-review and editing.
Max Hidalgo: Data curation, Investigation, Validation, Writing-review and editing.
Hernan Ortega: Data curation, Investigation, Validation, Writing-review and editing.
Pablo A. Tedesco: Conceptualization, Data curation, Funding acquisition, Investigation, Project administration, Resources, Supervision, Validation, Writing-original draft, Writing-review and editing.
Gislene Torrente-Vilara: Data curation, Funding acquisition, Investigation, Resources, Validation, Writing-review and editing.
Jansen Zuanon, Rémy Bigorne, Koen Martens, Astrid Acosta, Edwin Agudelo, Soraya Barrera Maure, Douglas A. Bastos, Juan Bogotá-Gregory, Fernando G. Cabeceira, André L. C. Canto, Fernando M. Carvajal-Vallejos, Lucélia N. Carvalho, Ariana Cella-Ribeiro, Raphaël Covain, Carlos DoNascimiento, Carolina R. C. Dória, Cleber Duarte, Efrem J. G. Ferreira, André V. Galuch, Tommaso Giarrizzo, Rafael P. Leitão, John G. Lundberg, Mabel Maldonado, Luciano F. A. Montag, William M. Ohara, Tiago H. S. Pires, Marc Pouilly, Saúl Prada-Pedreros, Luiz J. de Queiroz, Lucia Rapp Py-Daniel, Frank R. V. Ribeiro, Raúl Ríos Herrera, Jaime Sarmiento, Leandro M. Sousa, Lis F. Stegmann, Jonathan Valdiviezo-Rivera, Francisco Villa, Takayuki Yunoki, Aaike de Wever: Data curation, Investigation, Resources, Validation, Writing-review and editing.
Thierry Oberdorff: Conceptualization, Data curation, Funding acquisition, Investigation, Project administration, Resources, Supervision, Validation, Visualization, Writing-original draft, Writing-review and editing.
Ethical Statement
Not applicable.
Competing Interests
The author declares no competing interests.
Data availability statement
The corresponding point data containing the geographic coordinates of all georeferenced records and the information sources can be freely downloaded from the AmazonFish website at https://figshare.com/articles/dataset/A_database_of_freshwater_fish_species_of_the_Amazon_Basin/9923762. All codes are available as Appendix files and at https://github.com/msversutdias/AmazonFISHBetaDiversity.git.
AI statement
ChatGPT and DeepSeek has been used for preparing Bibliography list following NI standards.
Funding
This research benefited support from the ERANet-LAC (www.eranet-lac.eu/) “AmazonFish” (ELAC2014/DCC–0210) project. We thank the Field Museum of Natural History in Chicago and the John D. and Catherine T. MacArthur Foundation (Grant G–1607–151047) for having mobilized data for AmazonFish in Colombia, Ecuador, and Peru. French Laboratories of Excellence “CEBA” (ANR–10–LABX–25–01) and “TULIP” (ANR–10–LABX–41 and ANR–11–IDEX–0002–02) are also acknowledged. JZ (#313183/2014–7), RPL (#314464/2023–9) and CRCD (#304501/2023–9) acknowledge Brazil’s CNPq for productivity grants. MSD thanks CNPq for a post-doctoral fellowship (#150784/2015–5) and productivity (#310524/2022–9) grant, the INCT-Peixes (MCTIC/CNPq, #405706/2022–7), FAPDF (#00193.00001819/2018–75), and UnB/DPI/DPG (#04/2024) for funding. TO, BH and PAT thank the CEBA project “EMERGENCE”. GT-V. received grants from Foundation of Support to Research in the Amazon (PAREV/FAPEAM #019/2010), CAPES (Pro-Amazon Program: Biodiversity and Sustainability, process #6632/14–9), and FAPESP (São Paulo Research Foundation #2016/07910–0). All data was collected through the AmazonFish project (www.amazon-fish.com; https://freshwaterfishdata-dev.ird.fr).
Supplementary Material
Supplementary material S1
Supplementary material S2
Supplementary material S3
Supplementary material S4
Supplementary material S5
Supplementary material S6
Supplementary material S7
Supplementary material S8
Supplementary material S9
Supplementary material S10
How to cite this article
Dias MS, Jézéquel C, Hugueny B, Leprieur F, Brosse S, Albert J, Cassemiro FAS, Frederico RG, Hidalgo M, Ortega H, Tedesco PA, Torrente-Vilara G, Zuanon J, Bigorne R, Martens K, Acosta A, Agudelo E, Maure SB, Bastos DA, Bogotá-Gregory J, Cabeceira FG, Canto ALC, Carvajal-Vallejos FM, Carvalho LN, Cella-Ribeiro A, Covain R, DoNascimiento C, Dória CRC, Duarte C, Ferreira EJG, Galuch AV, Giarrizzo T, Leitão RP, Lundberg JG, Maldonado M, Montag LFA, Ohara WM, Pires THS, Pouilly M, Prada-Pedreros S, Queiroz LJ, Py-Daniel LR, Ribeiro FRV, Herrera RR, Sarmiento J, Sousa LM, Stegmann LF, Valdiviezo-Rivera J, Villa F, Yunoki T, Wever A, Oberdorff T. Dispersal limitation, environmental filtering, and historical contingencies explain the patterns of taxonomic and phylogenetic turnover in Amazonian freshwater fish faunas, but only poorly their functional traits turnover. Neotrop Ichthyol. 2026; 24(1):e250084. https://doi.org/10.1590/1982-0224-2025-0084
Copyright
This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Distributed under
Creative Commons CC-BY 4.0

© 2025 The Authors.
Diversity and Distributions Published by SBI
![]() Accepted December 2, 2025
Accepted December 2, 2025
![]() Submitted May 15, 2025
Submitted May 15, 2025
![]() Epub April 27, 2026
Epub April 27, 2026