![]() Lorena Lopes Almeida1,2,3
Lorena Lopes Almeida1,2,3 ![]() ,
, ![]() Maurício Hostim-Silva1,2,4,5 and
Maurício Hostim-Silva1,2,4,5 and ![]() Ana Paula Cazerta Farro1,3,4
Ana Paula Cazerta Farro1,3,4
PDF: EN XML: EN | Cite this article
Abstract
O mero, Epinephelus itajara, está Criticamente Em Perigo no Brasil e depende dos habitats de manguezais como berçários. Ele é extremamente vulnerável à sobrepesca; no entanto, avaliações genéticas de longo prazo continuam escassas, deixando uma lacuna em nossa compreensão das dinâmicas populacionais. Avaliamos a variação genética de 126 juvenis coletados em 2013, 2015, 2018 e 2022, utilizando 18 loci microssatélites. Treze loci foram polimórficos, com o número de alelos variando de 3 a 17. Os valores médios de heterozigosidade observada (Ho) e esperada (He) foram 0,706 (variação: 0,556–0,897) e 0,710 (variação: 0,524–0,890), respectivamente. Não houve diferenças significativas em He, FIS e FST. O tamanho efetivo da população (Ne) foi estimado em 413,3 (IC 95%: 260,8–897,1), sugerindo riscos potenciais para a viabilidade a longo prazo da espécie. A AMOVA indicou que 99,64% da variação genética total correspondeu a diferenças dentro dos indivíduos, enquanto a análise de agrupamento Bayesiana indicou que todos os indivíduos apresentaram ancestrais mistos. A análise de parentesco identificou um número notável de relações de irmandade, sugerindo um estoque reprodutivo reduzido. Nossos resultados sugerem que, devido a longa expectativa de vida da espécie, mudanças na estrutura populacional e na diversidade genética provavelmente não serão refletidas prontamente neste período de amostragem. Isso destaca a importância do monitoramento contínuo e da manutenção da proibição da pesca para proteger a espécie.
Palavras-chave: Diversidade genética, Genética populacional de peixes, Loci de microssatélites, Parentesco.
Introduction
The Atlantic goliath grouper, Epinephelus itajara (Lichtenstein, 1822), is the largest grouper species in the Atlantic Ocean. They can grow up to 2.5 m in total length and weigh 320 kg (Heemstra, Randall, 1993), living up to 37 years (Bullock et al., 1992). The species inhabits tropical and subtropical waters of the western Atlantic, from Florida to southern Brazil, including the Gulf of Mexico and the Caribbean Sea (Sadovy, Eklund, 1999; Hostim-Silva et al., 2005) as well as the African coast in the Eastern Atlantic, from Senegal to Angola (Bertoncini et al., 2018).
This iconic species presents biological traits such as being protogynous hermaphrodites (Murie et al., 2023), slow growing, late maturing, and long-lived, which makes them extremely vulnerable to overfishing (Sadovy, Eklund, 1999). Additionally, the loss of mangrove habitats has been linked to declines in Atlantic goliath grouper populations, as these areas serve as critical nursery grounds for juveniles (Koenig et al., 2007; Moreno-Santos et al., 2022). Due to these features, the Atlantic goliath grouper is currently categorized as Vulnerable (VU) (IUCN; Bertoncini et al., 2018) by the IUCN Red List, and Critically Endangered (CR) in Brazil (ICMBio, 2018). The species is one of the few epinephelids that live in brackish water (Lara et al., 2009; Condini et al., 2023, 2024) and areas with extensive mangrove development appear to be not only essential but critical nursery areas for juveniles, potentially limiting the species distribution (Koenig et al., 2007). The Atlantic goliath grouper life cycle involves movement between marine and estuarine habitats. Spawning takes place in marine areas, and the larvae are then carried to estuarine regions where they will settle and grow. Upon reaching sexual maturity, they migrate to deeper coastal marine environments (Eklund, Schull, 2001; Hostim-Silva et al., 2018).
Marine fish have complex genetic structures and high genetic connectivity due to the environment they inhabit (Jørgensen et al., 2005). Populations with significant genetic diversity can better respond to selection because the favored genotypes are more likely to be present (Taboun et al., 2021). Several groups of reef fish, such as the Atlantic goliath grouper, demonstrate spawning aggregation behavior that is crucial for either promoting long-distance dispersal or limiting exchange among populations (Jackson et al., 2014). A considerable number of spawning aggregation sites have been identified in Florida (Sadovy, Eklund, 1999; Koenig, Coleman, 2009; Mann et al., 2009; Koenig et al., 2016, 2017; Malinowski et al., 2019), however, the locations of these events along the southwest Atlantic are still little known (Gerhardinger et al., 2006; Bueno et al., 2016; Giglio et al., 2016).
The dynamics of adult Atlantic goliath grouper spawning in oceanic environments have been investigated along its distribution (Gerhardinger et al., 2006; Mann et al., 2009; Bueno et al., 2016; Koenig et al., 2017; Malinowski et al., 2019), however, the relationships between their reproduction and the number of juveniles present at a nursery area, have not been fully clarified. The strong reproduction site fidelity of adult Atlantic goliath grouper (Sadovy, Eklund, 1999; Koenig et al., 2007) may contribute to population structure, nevertheless, its prolonged pelagic larval phase of 30 to 80 days, with an average of 60 days (Lara et al., 2009) may contribute to larval dispersion and gene flow (Cowen, Sponaugle, 2009). Therefore, understanding the behavior, biology, and genetic diversity of individuals will provide new insights into adult population dynamics.
Microsatellites, also known as single sequence repeats (SSRs), are highly variable sequences that have been extensively used in population genetics studies to assess genetic diversity (Antoro et al., 2005; Taboun et al., 2021; Wenne, 2023), gene flow (Berry et al., 2012; Liu et al., 2016; Snead et al., 2023), temporal population structure (Diaz-Suarez et al., 2022; Burimski et al., 2024), reproductive success dynamics (Tringali, 2023), and relatedness (López et al., 2015; Sánchez-Velásquez et al., 2022; Jeannot et al., 2024). Temporal genetic analysis has become widely used to assess changes in the diversity and population structure of wild fish populations (Østergaard et al., 2003; López et al., 2015; Taboun et al., 2021; Burimski et al., 2024). These studies are vital for exploited and endangered fish species since they can track genetic diversity changes over time (Burimski et al., 2024), investigate the impacts of fishing pressure on genetic diversity (Taboun et al., 2021), and inform fisheries management strategies (Petrou et al., 2021). Measuring relatedness in wild populations is challenging due to partial or lacking pedigree information (Jeannot et al., 2024), especially for wild populations of polygamous species (Nance et al., 2011), such as the Atlantic goliath grouper, which forms relatively small spawning aggregations (10 to 100 individuals) (Sadovy, Eklund, 1999) with individuals migrating up to 500 km (Ellis et al., 2014). In this scenario, half-sibling relationships are prevalent and although they are less accurate than full-sibling or parent-offspring relationships (Melero et al., 2017), they are particularly informative for populations with large effective population size (Ne) and polygamous reproductive behavior, where full siblings occur at a lower frequency (Wang, Santure, 2009).
The present study aims to enhance our understanding of the genetic diversity and population structure of the Atlantic goliath grouper, particularly focusing on temporal variations. Given the absence of previous data, our results provide the first baseline for understanding the temporal genetic diversity and population dynamics of the Atlantic goliath grouper in the Southwestern Atlantic. Here, we sought to determine whether more than one genetically distinct population contributes to the juvenile presence in the study area. To achieve this, we monitored the temporal genetic composition of juvenile Atlantic goliath grouperusing microsatellite loci. We also evaluated the genetic diversity of this species and tested the hypothesis of temporal genetic structuring on the central Brazilian coast over the course of 10 years. The results herein are crucial, as they provide valuable insights for the species into a region where such information is currently unavailable.
Material and methods
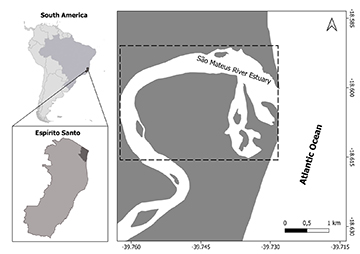
Sample collection. This study was conducted in the São Mateus River estuary, within the Environmental Protection Area of Conceicão da Barra, on the southeastern Brazilian coast (Fig. 1). The study area is located south of the Abrolhos Bank continental shelf, habitat with the largest and richest coral reef system in the South Atlantic Ocean (Leão et al., 2003; Francini-Filho et al., 2019), being an important habitat for Atlantic goliath grouper (Freitas et al., 2015; Zapelini et al., 2017). The region features distinctive environments, including well-preserved sandbanks and a vast mangrove forest. The dominant vegetation species is the white mangrove, Laguncularia racemosa, followed by the red mangrove, Rhizophora mangle (Tognella et al., 2020). These, along with distinct physical, morphological, and structural features, such as muddy substrates, mangrove roots, and varying depths, make the area a suitable nursery habitat for juvenile Atlantic goliath grouper (Moreno-Santos et al., 2022; Condini et al., 2024; Damasceno et al., 2025).
FIGURE 1| Map showing the study area in the estuary of the São Mateus River in Conceição da Barra, Espírito Santo State, Brazil, where juvenile Atlantic goliath grouper individuals (Epinephelus itajara) were captured (dotted rectangle) in 2013, 2015, 2018, and 2022.
A total of 136 juvenile Atlantic goliath grouper were sampled from 2013 to 2022 (2013, n = 21; 2015, n = 36; 2018, n = 39; 2022, n = 40). Juvenile Atlantic goliath grouper were collected using baited blue crab traps, and we also received donations from artisanal fishers’ incidental captures using a variety of gears, such as cast nets, gill nets, surrounding nets, setlines, and line and hook. To make sure we did not sample the same fish over time, we marked each fish with an identification microchip. Samples were collected employing a nonlethal method using caudal fin clips, causing minimal effects on fish, which were immediately released after sampling procedures. Samples were conserved in 96% ethanol and stored at -20 oC.
Microsatellite DNA amplification and genotyping. The genomic DNA was isolated from caudal fin samples using the DNeasy® Blood & Tissue Kit according to the manufacturer’s protocols (https://www.qiagen.com). Eighteen highly polymorphic microsatellite loci originally developed for the Atlantic goliath grouper(Seyoum et al., 2013) were used to assess nuclear genetic variation among samples (Tab. 1). The forward primer from each locus was 5′-fluorescently labeled with one of the three dyes: 6-FAM, HEX, and NED (Applied Biosystems). Polymerase chain reaction (PCR) was performed in 25 μl reactions containing 10 to 20 ng/μl of template DNA, 21 μl of Platinum PCR Supermix (Applied Biosystems), and 0.6 μM forward and reverse primers. For all loci, samples were subjected to an initial hot-start step at 94° C for 2 min; followed by 35 cycles of: denaturation at 94° C for 30 s, annealing temperature at 57° C for 30 s, extension at 72° C for 30 s; and a final extension at 72° C for 7 min. PCR amplification products were verified on 1.5% agarose gel, and fragment analysis of eight multiplex sets was performed on an Applied Biosystems genetic analyzer ABI 3730xl by Macrogen Inc. (South Korea).
TABLE 1 | Microsatellite loci used to assess the genetic diversity of juvenile Atlantic goliath grouper (Epinephelus itajara). Loci names (Loci); primers sequences (Sequence 5’-3’), forward (F) and reverse (R) sequences; allele size range (base pairs, bp); fluorescent primer labeling (Dye); and Multiplex PCR reaction (Multiplex). Original loci developed by Seyoum et al. (2013).
Loci | Sequence (5′ – 3′) | bp | Dye | Multiplex |
Eita01 | F:GTGTGTAAGGCTGATGTGATTTATTC | 259-281 | HEX | E |
R:AGCCTTAATTAGAGAGTCGTGTTCTT | ||||
Eita03 | F:ATCAGATTTGGAATTAGATCCATTG | 274-318 | FAM | D |
R:GTAGGCCAACAGTCTTCTCTATCTG | ||||
Eita06 | F:GAATGTACCTTTGCTACAGGAGTGT | 239-257 | FAM | F |
R:ATACATTTTTGGTCATGTCTGTGTG | ||||
Eita07 | F:TGCTTGTATATGTGTGTGAGAAAGA | 197-231 | FAM | C |
R:TAAGGTCATATTGATCTCCCTGTGT | ||||
Eita12 | F:ATTGTGTAATGTGTTCCTGTTTGTG | 274-286 | NED | G |
R:AGATGGTCTTACGTGTGATCTTGTT | ||||
Eita19 | F:AAATAACGGGAGTAGCTACCAAGAC | 192-218 | FAM | H |
R:CCCCTCACCCGTTTATATTTTATTA | ||||
Eita20 | F:CGTATTCAAGGTGTGTGTTTAAATG | 196-220 | HEX | G |
R:ACAGTGGGGTAATGAATTATGAAAG | ||||
Eita25 | F:GTGTGTGCTTTTGTATGTTGAACTG | 167-197 | HEX | F |
R:ATATTATGCCACAGCGTCAGAGT | ||||
Eita27 | F:AAGCAGTTCCGATTAGATAACGTC | 173-179 | FAM | B |
R:GCTGTGCAGTAAATATGCTTTGAC | ||||
Eita28 | F:CTCTGTCTTGAAAACACATAAAGCA | 142-158 | FAM | E |
R:GGTAGATTGCTCTGACATCTCATCT | ||||
Eita29 | F:ACCTGTATTTCTTAATGCTGATGTCTT | 155-175 | FAM | D |
R:GGGAAGTATTCCTTTAACATACCGTAA | ||||
Eita30 | F:AGAAATCTGTCTAGCGACTACATGC | 114-160 | FAM | C |
R:CTGAGTAACCTCTGACCAAATAAGC | ||||
Eita31 | F:CTTGGTGCTCAAGTATCTTTTCTCATA | 141-159 | NED | H |
R:GATCACTACCTCTTTTTCTTCATCAAA | ||||
Eita33 | F:TATAGACCAATCAGGTTGCATGAAT | 154-182 | HEX | A |
R:ATGAGGGATCATGAAGCAAAATC | ||||
Eita34 | F:TAAATCAGCACATTTATGTAACAAGGT | 129-175 | FAM | F |
R:GTTCTCTTTCTCCATGTAGGTGAGTTA | ||||
Eita36 | F:GAGGAAGAAAGACGGAAGATAGACT | 139-163 | FAM | G |
R:TTCATGGTTAGTTGATCACTGGAAT | ||||
Eita38 | F:CTACACCAGTTTATTCTGTGACCAG | 89-113 | HEX | B |
R:GAGTCGACACAGCTCTCATTAACTT | ||||
Eita41 | F:CAAGGGCAGAGCGAAGAC | 161-173 | FAM | A |
R:TGTACACAACATCCTCTGGGTACT |
Data analysis. Allele peaks were visualized in Geneious v. 6.1.4 (Kearse et al., 2012) (www.geneious.com). Ten percent of the homozygous individuals were re-genotyped, to ensure their homozygosity and minimize genotyping errors. Cervus v. 3.0.3 (Kalinowski et al., 2007) was used to assess the average non-exclusion probability for individual identity, and potential duplicate samples were searched by comparing their genotypes. All loci were checked for the presence of null alleles and other genotyping errors (allele dropout and stutter peaks) using MICRO-CHECKER 2.2.3 (Van Oosterhout et al., 2004) with Bonferroni correction. Linkage disequilibrium and Hardy-Weinberg equilibrium were tested using GENEPOP on the Web (Raymond, Rousset, 1995; Rousset, 2008).
For the genetic diversity analyses, only polymorphic loci were used. We calculated for each locus the total number of alleles (Na), mean number of alleles (Nam), observed (Ho) and expected (He) heterozygosities, the polymorphic information content (PIC), and the inbreeding coefficient (FIS) for each sample using the software Fstat 2.9.3 (Goudet, 2001). The program NeEstimator 2.0 (Do et al., 2014) was used to estimate contemporary effective population size (Ne) for all sampling units grouped as one population (2013, 2015, 2018, 2022) using a method based on linkage disequilibrium (Waples et al., 1989, 2010). A Pcrit value of 0.02 and 95% confidence intervals were chosen to reduce the potential bias for low-frequency alleles.
To test the hypothesis of recent events of population size reduction, we calculated the standardized Garza and Williamson M index, which quantifies the reduction in the number of alleles relative to the allelic size range of a population to detect recent bottleneck events, applying the threshold of 0.68. In the following, we used the software Bottleneck v. 1.2.02 (Cornuet, Luikart, 1996), and calculations were based on heterozygosity excess. The analysis was performed under both the Stepwise Mutational Model (SMM) and the Two-Phase Model (TPM; 30 variations, mutational model 70 gradual, 10,000 interactions; DiRenzo et al., 1994). The probability of significant heterozygosity excess was assessed using the one-tailed Wilcoxon signed-rank test (α = 0.05) based on 10,000 replications, along with the Sign testand Standardized Differences Test (SDT) to further assess the evidence for recent bottleneck events. Populations showing significant heterozygosity excess were considered as having likely undergone a recent genetic bottleneck.
The software Arlequin v. 3.5.2.2 (Excoffier, Lischer, 2010) was used to investigate the temporal population structuring of juvenile Atlantic goliath groupers sampled in 2013, 2015, 2018, and 2022. Pairwise FST was estimated based on the number of differences, with 10,000 random generations. A Bayesian clustering analysis was performed using Structure v. 2.3.2 (Pritchard et al., 2000) to verify the existence of different genetic clusters (K) in the samples, using an admixture model without prior population information and a correlated allele frequencies model. The burn-in was set at 150,000 length steps, followed by 750,000 iterations of the Markov Chain Monte Carlo (MCMC) simulations. Twenty independent iterations were performed for each value of K ranging from one to 10 and the consistency of clustering assignments across runs was evaluated to ensure the stability of results and to assess convergence. The Structure Harvester (Earl, VonHoldt, 2012) (available at http://taylor0.biology.ucla.edu/structureHarvester/), a web-based program was used to assess and determine the optimal values of k according to Evanno’s ∆K method (Evanno et al., 2005). The postprocessing of the outputs was performed using the CLUMPAK program (Kopelman et al., 2015), which assured convergence by summarizing and visualizing clustering patterns across independent runs. In addition, a principal coordinate analysis (PCoA) was performed using the R package PopGenReport v. 3.0.4 (Adamack, Gruber, 2014) to visualize genetic diversity among sampled individuals.
COLONY (Jones, Wang, 2010) was used to infer full- and half-sibship relationships among all sampling units grouped as a single population (126 juvenile Atlantic goliath grouper individuals) and within each sampling unit (2013, 2015, 2018, 2022). The program, which relies on multi-locus genotype data, is suitable for analyzing sibling relationships even in the absence of known parental genotypes (Wang, 2012). Sibling pair assignments were accepted if the probability was greater than 0.75, as used for other coral reef fishes (Herrera et al., 2016; Robitzch et al., 2020). The following parameters were used: polygamous diploid dioecious species, no inbreeding, unknown population allele frequencies, no information on the maternal or paternal mean sibship size, and using the full likelihood (FL) method, as it is the most accurate one as verified by simulated and empirical data analyses (Wang, 2012). One random seed number was used and the results were verified by running the analyses twice. Later, the Pedigree Viewer program (Kinghorn, Kinghorn, 2009) was used to visualize and analyze the results.
Results
Genetic structure and diversity. Of the 136 caudal fin samples, 126 (2013, n = 19; 2015, n = 35; 2018, n = 37; 2022, n = 35) were used in further analyses and provided genotypes for 12-13 loci. Two pairs of samples (PMB246 with PMB247 from 2013, and Ep22 with Ep30 from 2015) had matching genotypes probably due to labeling errors, therefore PMB246 and Ep30 were excluded from further analyses. Eight samples were also excluded due to either missing genotypes or low-quality peaks. Five loci were not used in the analysis: Eita31 was rejected because it had too many peaks, null alleles were found in Eita27 and Eita30, and linkage disequilibrium was shown in Eita19 and Eita29. The thirteen other microsatellite loci were moderately variable, e.g., k = 3–17 (Tab. 2). None of the 13 loci showed a significant departure from Hardy-Weinberg equilibrium (Tab. 2) and the probability of identity was 3.051 X 10-13.
TABLE 2 | Genetic diversity of juvenile Atlantic goliath grouper (Epinephelus itajara) from Conceição da Barra, Espírito Santo state, Brazil, for 13 microsatellite loci. N: sample size for each locus; Na: total number of alleles; Ho: observed heterozygosity; He: expected heterozygosity; PIC: Polymorphic Information Content; Ra: allelic richness; Dg: genetic diversity; FIS: inbreeding coefficient; F (Null): estimation of the frequency of null alleles; Hardy-Weinberg Equilibrium P (HWE). No statistically significant values for HWE P < 0.05.
Loci | N | Na | Ho | He | PIC | Ra | Dg | FIS | F(Null) | P(HWE) |
Eita01 | 126 | 5 | 0.587 | 0.563 | 0.500 | 4.984 | 0.562 | -0.044 | -0.018 | 0.322 |
Eita03 | 126 | 15 | 0.794 | 0.807 | 0.780 | 14.984 | 0.807 | 0.017 | 0.008 | 0.768 |
Eita06 | 125 | 9 | 0.784 | 0.803 | 0.779 | 9.000 | 0.803 | 0.024 | 0.015 | 0.747 |
Eita07 | 126 | 8 | 0.690 | 0.740 | 0.691 | 7.992 | 0.740 | 0.067 | 0.033 | 0.253 |
Eita12 | 126 | 3 | 0.563 | 0.524 | 0.451 | 3.000 | 0.524 | -0.076 | -0.042 | 0.731 |
Eita20 | 125 | 10 | 0.696 | 0.652 | 0.623 | 10.000 | 0.652 | -0.068 | -0.047 | 0.657 |
Eita25 | 125 | 14 | 0.824 | 0.828 | 0.803 | 14.000 | 0.828 | 0.005 | 0.001 | 0.393 |
Eita28 | 126 | 7 | 0.635 | 0.692 | 0.640 | 7.000 | 0.692 | 0.082 | 0.046 | 0.147 |
Eita33 | 126 | 8 | 0.817 | 0.810 | 0.780 | 8.000 | 0.810 | -0.010 | -0.008 | 0.952 |
Eita34 | 126 | 17 | 0.897 | 0.890 | 0.876 | 16.976 | 0.890 | -0.008 | -0.007 | 0.388 |
Eita36 | 126 | 7 | 0.556 | 0.583 | 0.524 | 6.984 | 0.583 | 0.046 | 0.029 | 0.203 |
Eita38 | 126 | 7 | 0.659 | 0.664 | 0.624 | 6.992 | 0.664 | 0.008 | 0.010 | 0.111 |
Eita41 | 126 | 7 | 0.675 | 0.674 | 0.624 | 6.984 | 0.674 | -0.002 | -0.001 | 0.832 |
Mean | 125.77 | 9 | 0.706 | 0.710 | 0.669 | 8.992 | 0.710 |
|
|
|
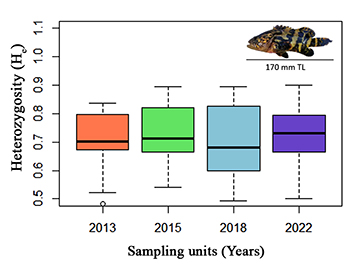
The average number of microsatellite alleles per locus was nine (range: 3–17 alleles), Eita34 was the most variable locus (17 alleles) and Eita12 was the least variable locus (three alleles) (Tab. 2). There were no He significant differences between the sampling units (Fig. 2). The average Ho was 0.706 (range: 0.556–0.897), and the average He was 0.710 (range: 0.524–0.890, Tab 2). The PIC at each microsatellite locus was always higher than 0.5, except for Eita12 (range: 0.451–0.876) (Tab. 2). The contemporary effective population size of the juvenile Atlantic goliath grouper population, estimated according to the linkage disequilibrium method, was 413.3 (95% CI 260.8–897.1; harmonic mean sample size: 126) for all sampled units grouped into one population.
FIGURE 2| Expected heterozygosity (He) across sampling units (years; 2013, 2015, 2018, and 2022) for 13 microsatellite loci of juvenile Atlantic goliath grouper (Epinephelus itajara, top right – TL stands for total length) sampled in Conceição da Barra, Espírito Santo State, Brazil. For each sampling unit, the thick black lines show the median values and the box depicts the quartiles of the loci values. Whiskers show 1.5× the interquartile range or the maximum value, with circles representing outliers.
The standardized Garza-Williamson M index ranged from 0.23 to 0.58, which is below the commonly used threshold of 0.68, suggesting a recent or past population bottleneck. However, the results of bottleneck detection tests varied among methods (Tab. 3). The Wilcoxon one-tailed test showed no evidence of a bottleneck under either the TPM (P = 0.31775) or SMM (P = 0.99957) models. In contrast, both the Sign Test and the SDT detected significant heterozygosity excess under the SMM model (P < 0.05), suggesting a potential reduction in genetic diversity.
TABLE 3 | Results of the Bottleneck Test for juvenile Atlantic goliath grouper (Epinephelus itajara) sampled in 2013, 2015, 2018, and 2022 in Conceição da Barra, Espírito Santo State, Brazil. SDT: Standardized different test; TPM: Two-phase model; SMM: Stepwise mutational model. Bold values (P < 0.05) indicate evidence of a recent bottleneck. The normal mode-shift indicates the approximately L-shaped allele frequency distribution.
Sign test | SDT | Wilcoxon sign-rank test | ||||
TPM | SMM | TPM | SMM | TPM | SMM |
Mode-shift |
0.53874 | 0.00172 | 0.41514 | 0.00001 | 0.31775 | 0.99957 | |
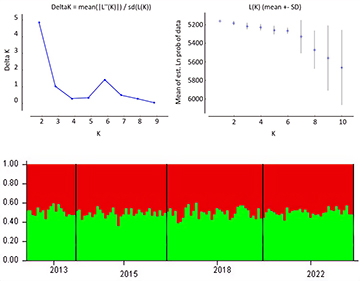
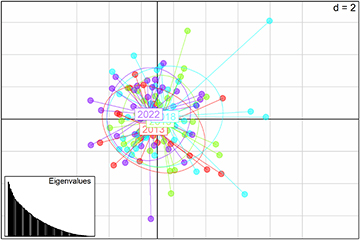
There were no statistically significant values for FIS (Tab. 2) and FST (Tab. 4) for the temporal genetic structuring test between the individuals sampled in the different years. AMOVA results indicated that 99.64% of total genetic variation corresponded to differences within individuals, 0.35% corresponded to differences among individuals within sampling units, and there were no differences among sampling units. There was no detectable substructure by sampling period or overall, according to the Bayesian clustering analysis. ∆K revealed that the greatest likelihood was K = 1, and, K = 2 showed all individuals to have similarly mixed ancestries and were mostly assigned to a single cluster (Fig. 3). Similarly, the PCoA plot showed an overlap of individuals from all sampling units with no evident clustering (Fig. 4), as well as no differences, were observed for the pairwise FST and the AMOVA results (Tab. 4).
TABLE 4 | Pairwise FST (lower diagonal) and P values (upper diagonal) of Atlantic goliath grouper (Epinephelus itajara) sampled in Conceição da Barra, Espírito Santo State, Brazil, for thirteen microsatellite loci among the sampling units (2013, 2015, 2018, and 2022). No statistically significant values for FST P < 0.01.
Sampling units | 2013 | 2015 | 2018 | 2022 |
2013 | ******* | 0.39333 | 0.77022 | 0.20354 |
2015 | 0.00083 | ******* | 0.80923 | 0.48787 |
2018 | -0.00319 | -0.00292 | ******* | 0.23107 |
2022 | 0.00429 | -0.00021 | 0.00213 | ******* |
FIGURE 3| Bayesian clustering analysis inferred with the program structure 2.3.4 for 13 microsatellite loci of juvenile Atlantic goliath grouper (Epinephelus itajara) (n = 126) sampled in Conceição da Barra, Espírito Santo State, Brazil. (a) mean log probability (Lk) is given over 20 iterations for each K value, (b) ∆K calculated from Evanno’s method for successive K values, and (c) barplot with assignment probabilities of individuals to putative population clusters at k = 2 according to sampling units (2013, 2015, 2018, and 2022).
FIGURE 4| Principal Coordinate Analysis (PCoA) based on the genetic distance matrix of FST values for 13 microsatellite loci of juvenile Atlantic goliath grouper (Epinephelus itajara) sampled in Conceição da Barra, Espírito Santo State, Brazil, for the sampling units of 2013 (red), 2015 (green), 2018 (blue), and 2022 (purple).
Parentage analysis. The sibship analysis with COLONY for the sampling units grouped as a single population indicated 48 and 54 possible fathers and mothers, respectively, with a maximum of seven paternity and six maternity families. The contribution of each mother varied from one to six offspring individuals, while each father’s contribution varied from one to seven offspring individuals. One possible mother contributed to six offspring individuals distributed into the four sampling units (2013, 2015, 2018, 2022), this pattern was also observed for another possible mother who contributed to five offspring individuals. One father contributed to seven offspring individuals distributed into three of the four sampling units (2013, 2018, 2022) and one father contributed to six offspring individuals distributed into three of the four sampling units (2015, 2018, 2022). No fathers were observed to contribute to all four sampling units.
Out of 7875 possible sibling pairs, three (0.03%) full-sibling pairs (Epita22-2015 and Eita49-2018, Eita6-2018 and Eita62-2018, Eita38-2018, and Eita44-2018) and 89 (1.1%) half-sibling pairs were above the threshold probability of 0.75 and were accepted as true full-siblings and half-siblings, respectively. Three possible mothers and three possible fathers were responsible for the three full-sibling pairs. The full-sibling pair Epita22-2015 and Eita49-2018 had one half-sibling on their mother’s side (Eita206-2022), the full-sibling pair Eita6-2018 and Eita62-2018 had one half-sibling on their father’s side (Epita33-2015), and the full-sibling pair Eita38-2018 and Eita44-2018 had one half-sibling on their mother’s side (Eita12-2018) and three half-siblings on their father’s side (Eita17-2018, Eita213-2022, and Eita216-2022).
The temporal sibship analysis for each sampling unit identified four half-sibling pairs and no full-sibling pairs for 2013, with a probability of nine fathers and nine mothers; 20 half-sibling pairs and one full-sibling pair for 2015, with a probability of 17 fathers and 17 mothers; 21 half-sibling pairs and three full-sibling pairs for 2018, with a probability of 15 fathers and 17 mothers; and 33 half-sibling pairs and two full-sibling pairs for 2022, with a probability of 15 fathers and 15 mothers.
Discussion
Genetic structure and diversity. We found moderate nuclear genetic diversity that has been maintained over the years and no evidence of temporal genetic structure for the juvenile Atlantic goliath grouper population analyzed in this study for the last 10 years, indicating some degree of gene flow among the sampling units. However, the low Ne suggests potential risks to the population’s long-term genetic viability, despite the apparent stability in genetic diversity. In the following, we discuss these findings and their implications for the management and conservation of Atlantic goliath grouper in the estuary of the São Mateus River, an important area for the conservation of the species, due to the high frequency of juvenile individuals found in the estuarine mangrove area throughout the year (Damasceno et al., 2015; Almeida et al., 2022; Moreno-Santos et al., 2022; Condini et al., 2023, 2024).
Seyoum et al. (2013) was the first study to isolate and characterize microsatellite markers for the Atlantic goliath grouper, and to the best of our knowledge, our study is the first to apply 18 of those markers to investigate the genetic diversity of the Atlantic goliath grouper on the Southwestern Atlantic. Our results suggested that the mean number of alleles (Na = 9), observed and expected heterozygosity (Ho= 0.706, He= 0.710), were higher than those found by Seyoum et al. (2013) using the same set of markers (Na= 7, Ho= 0.610, and He= 0.653), but these findings could be a result of the number of samples analyzed in the present study (n = 126) compared to those analyzed by Seyoum et al. (2013) (n = 52). When compared to other Epinephelus Bloch, 1793 species, the average number of alleles (Na= 9) remained higher, while the average expected heterozygosity (He = 0.71) was lower than those found for E. tauvina (Fabricius, 1775) (Na = 8.09; He= 0.83), E. coioides (Hamilton, 1822) (Na = 8.36; He = 0.85), E. bleekeri (Vaillant, 1878) (Na = 7.36; He= 0.79), E. malabaricus (Bloch & Schneider, 1801) (Na = 8; He= 0.80), and E. areolatus (Forsskål, 1775) (Na = 7.9; He= 0.79) from the Arabian Gulf (Hassanien, Al-Rashada et al., 2021). The 13 microsatellites used in this study exhibited a moderate to high polymorphism (0.669 for mean PIC), indicating their effectiveness in detecting genetic variation within the population and our results are consistent with those found for the Giant grouper E. lanceolatus (Bloch, 1790) (Weng et al., 2021).
The Atlantic goliath grouper presents a series of biological and ecological traits that heighten its susceptibility to Ne reduction, including its long lifespan, late maturity, and reproductive aggregations. Ne is a key parameter in conservation genetics, as it reflects a population’s historical exposure to genetic drift and inbreeding while also providing insight into its long-term viability (Wang et al., 2016). Our estimate of the contemporary Ne for the juvenile Atlantic goliath grouper population was 413.3 (95% CI: 260.8–897.1), a value below the threshold of 500 recommended for long-term genetic stability and population viability (Frankham et al., 2014). Similarly, Tringali (2023) reported a Ne of approximately 659 (95% CI: 588–745) for adult Atlantic goliath grouper in Florida waters, also near this critical threshold. These findings highlight potential concerns regarding the species’ evolutionary potential and resilience to environmental changes and fishing pressures, reinforcing the need for continued genetic monitoring and conservation measures.
The effect of coastal current dynamics along the South American coast on Atlantic goliath grouper populations is still unknown, however, studies with E. marginatus suggested that ocean currents shape the genetic structure of the species by promoting specific local larval retention (Schunter et al., 2011). In general, larval dispersal is the main mechanism of connectivity in reef fish. For the Atlantic goliath grouper, the duration of the larval period generally varies from 40 to 60 days after spawning, sometimes extending from 30 to 80 days (Lara et al., 2009), allowing plenty of time for long-distance larval dispersal by winds and currents before settlement. The genetic diversity of the Atlantic goliath grouper varies considerably along the Brazilian coast with a significant degree of genetic structuring and a high degree of genetic differentiation between locations using the control region of the mitochondrial DNA, indicating that gene flow on the Brazilian coast is discontinuous, not homogeneous, and lacking evidence of any systematic relationship with geographic patterns (Damasceno et al., 2015). Despite being genome regions with high rates of differentiation, relatively low genetic variability has been described in the Atlantic goliath grouper using the control region (mtDNA) (Silva-Oliveira et al., 2008) and microsatellite loci (Seyoum et al., 2013), confirming the species’ vulnerability to ongoing anthropogenic impacts, such as fishing pressure and habitat destruction.
The detection of a bottleneck through the excess of heterozygosity in our study suggests that the population has gone through a substantial decline at some point in its history, most likely resulting in a loss of genetic diversity. This raises concerns about the species’ long-term viability, as reduced genetic variation may impact adaptability and resilience to environmental changes. A study conducted in the Florida coastal waters also reported evidence of reduced genetic diversity in the Atlantic goliath grouper, with few individuals contributing to offspring for the following generation and a high variance in reproductive performance among individuals (Tringali, 2023). However, interpreting bottleneck events in this species remains challenging due to the lack of studies specifically investigating historical population dynamics. It is important to note that heterozygosity excess tests only detect bottleneck events within a specific time frame, as a population bottleneck with an Ne as low as 50 remains detectable for 25–250 generations following a decline (Cornuet, Luikart, 1996). While our results suggest a bottleneck event, further research is needed to determine its timing, severity, and potential long-term effects on the population.
A possible explanation for the lack of significant results in FST, AMOVA, and FIS, lies in the complex relationships between the variables influencing the Atlantic goliath grouper population. Even though it is well documented that the species experienced a sharp decline in the 1990s (Sadovy, 1994; Gerhardinger et al., 2006; Giglio et al., 2017; Zapelini et al., 2017; Malinowski et al., 2020; Locatelli et al., 2023) the effects of this demographic decrease may not be readily reflected during the sampling period of this study. With an average generation length of 21.5 years (Bertoncini et al., 2018) and a life span of up to 37 years (Sadovy, Eklund, 1999), the species late maturing and longevity suggest that shifts in population dynamics may take longer to be identified than our sampling period allows. Additionally, the AMOVA results indicated that most of the genetic variation was observed within individuals, rather than between individuals or sampling units, further supporting the possibility that temporal changes may not be easily detectable within the limited sampling time of this study.
Although no significant genetic structuring were detected in our study, it is still critical to recognize the challenges facing the Atlantic goliath grouper population. While the species is protected in Brazilian jurisdictional waters (Ordinance N° 121/2002 – IBAMA, 2002), the continued exploitation by fishers and illegal commercialization (Giglio et al., 2014; Almeida et al., 2024), leads to the depletion of local populations over time. These pressures may not immediately result in detectable genetic changes, but they continue to threaten the species long-term survival. Therefore, our findings highlight the critical need for increased monitoring efforts and the implementation of stronger conservation measures to ensure the genetic diversity and long-term survival of Atlantic goliath grouper populations. Consequently, our results suggest that the fishing ban protecting the species should be maintained and these population units need to be monitored for a longer period to enable the detection of changes that could pose threats to this species, since the reductions of Ne may result in elevated rates of genetic diversity loss (Larsson et al., 2010).
Parentage analysis. There are few studies concerning the spawning aggregations of Atlantic goliath grouper along the Brazilian coast (Félix-Hackradt, Hackradt, 2008; Bueno et al., 2016; Giglio et al., 2016), and yet, none of them include the coast of the state of Espírito Santo. This leaves a gap in understanding how many aggregation spawning sites are contributing to the juvenile Atlantic goliath grouper population investigated in this study. The closest known aggregation site to the study area is in the state of Bahia (Giglio et al., 2016), between the Coroa Vermelha and the Abrolhos Archipelago, approximately 100 km north of the study site, however, Damasceno et al. (2015) found genetic differentiation between Atlantic goliath grouper population units from these two geographically close areas (Municipality of Conceição da Barra in the state of Espírito Santo and the Municipality of Caravelas in the state of Bahia), but not between geographically distant areas from our study site, such as Municipality of Parnaíba (2,300 km north) in the state of Piauí and the Municipality of Bragança (2,600 km north) in the state of Pará. This suggests that the Abrolhos Bank may play an important role in larval retention and that these processes need a deeper investigation. Nevertheless, these results were obtained using mitochondrial markers, which are useful for studying maternal lineages and ancient population events, while microsatellites are useful for studying recent population events and both maternal and paternal lineages.
The accuracy obtained from COLONY is low for panels with few microsatellite loci, but it reaches a maximum of 100% for panels over eight loci (Weng et al., 2021). Our 13-locus microsatellite panel revealed a relatively low prevalence of closely related individuals among the collected juveniles in each sampling unit, as well as across all the sampling units as one population. While this initially suggests that the population in the São Mateus River estuary is large enough to support a diverse pool of breeding individuals, the observed parentage results raise some concerns. These results are notable and concerning, particularly given the relatively small dataset collected over a 10-year period, where each year’s sample represents a random draw from the total population. The pattern of sibling relationships observed points toward a reduced spawning stock rather than a diverse breeding pool. As the frequency of siblings increased throughout the study, it suggests that inbreeding or limited breeding opportunities may be occurring, contrary to what would be expected in a healthy population of broadcast-spawning marine species, where full and half-siblings are typically rare.
Furthermore, some fathers contributed as many as seven offspring, while some mothers produced as many as six. Two potential mothers participated in aggregations across all four sampling units, indicating that these females likely returned to the spawning grounds each year (Coleman et al., 2000). Interestingly, no fathers were observed to contribute to all four sampling units, which could indicate a well-mixed population, with dominant males disproportionately contributing to offspring production. However, this absence of fathers across multiple sampling units could also suggest that male individuals are being harvested, further supporting the possibility of reduced spawning stock. These findings suggest ongoing pressures on the population, highlighting the need for closer monitoring to assess the long-term vialbility of the breeding stock.
Implications for management and conservation. In Brazilian waters, the Atlantic goliath grouper has been protected for 22 years, through a series of fishing bans specifically targeting the species. The ban was initially established in 2002 (Ordinance N° 121/2002 – IBAMA, 2002) and was renewed in 2007 (Ordinance N° 42/2007 – IBAMA, 2007), 2012 (Ordinance N° 13/2012 – MMA/MPA, 2012), and 2015 (Ordinance N° 13/2015 – MMA/MPA, 2015), protecting the species until 2023. Since 2023, the species no longer benefits from a specific fishing ban; however, it remains listed as Critically Endangered (CR) under Ordinance N° 445/2014 (MMA, 2014). Despite these conservation efforts, illegal captures are still taking place along the Brazilian coast (Giglio et al., 2014; Almeida et al., 2024).
Our findings further emphasize the species’ vulnerability by revealing a contemporary Ne of 413.3 (95% CI: 260.8–897.1), which falls below the 500-individual threshold recommended for maintaining long-term genetic stability and adaptive potential (Frankham et al., 2014). Coupled with the species’ biological traits, such as being long-lived, late-maturing, protogynous, and hermaphroditic, and the ecological traits of showing strong site fidelity and forming reproductive aggregations, these factors make the species particularly vulnerable to environmental changes and fisheries pressures.
The lack of previous baseline data makes it challenging to assess trends in the genetic diversity of Atlantic goliath grouper populations. Further studies using genetic monitoring tools are necessary to determine whether the population is rebounding or experiencing further declines. Without such data, conservation efforts may fail to ensure the species long-term genetic stability, which is essential for its evolutionary potential and resilience to environmental stressors.
Therefore, this study provides the first baseline for understanding the species genetic diversity and population dynamics in the Southwestern Atlantic. Establishing this baseline is essential for future research and management strategies aimed at ensuring the species long-term survival. In light of these findings, we strongly suggest that: i) the current ban on the harvesting of the Atlantic goliath grouper in Brazil should be maintained; ii) coastal protection areas based on genetic data should be created, especially on mangrove and breeding sites, critical areas for the reproduction and development of Atlantic goliath grouper and several species; and iii) investments should be designated to new research concerning the genetic structure and gene flow of Atlantic goliath grouper along the Brazilian coast.
These measures are imperative to safeguard the species and ensure its long-term survival, especially considering that Brazil encompasses the largest area span of distribution for the Atlantic goliath grouper within its total distribution range, and may play an important role in its global conservation.
Acknowledgments
This research was developed and funded by Meros do Brasil Project, sponsored by Petrobras, and FAPES (FAPES N°21/2018 – UNIVERSAL). LLA thanks CAPES for the scholarships (PDSE – 88881.361817/2019–01 and PDS – 88882.385188/2019–01). MHS thanks the research fellowship provided by CNPq (Proc. 310361/2022–2) and FAPES for the Research Support Program grant (TO: 221/2019). We thank Júnio D. de Souza for the 2013 samples and fieldwork data information. A special thanks to the fishers Juliete L. Clarindo, Juliana L. Clarindo, Romildo Lopes, and Nador L. Muniz (In Memoriam) for fieldwork effort throughout the years and for facilitating our interactions with the local community and other fishers. In memoriam to Dr. Christopher Koenig, whose groundbreaking research on the Goliath Grouper not only advanced marine biology but also laid the foundation for the protection of this iconic species, leaving a legacy that will continue to inspire future generations of scientists.
References
Adamack AT, Gruber B. PopGenReport: simplifying basic population genetic analyses in R. Methods Ecol Evol. 2014; 5(4):384–87. https://doi.org/10.1111/2041-210X.12158
Almeida LL, Hostim-Silva M, Condini MV, Freitas MO, Bueno LS, Bentes B et al. Mislabeling, illegal capture, and commercialization of Atlantic goliath grouper (Epinephelus itajara) on the Brazilian coast using DNA barcoding. Neotrop Ichthyol. 2024; 22(1):e230099. https://doi.org/10.1590/1982-0224-2023-0099
Almeida LL, Stallings CD, Condini MV, Garcia AM, Tzadik OE, Koenig CC et al. Nonlethal stable isotope analysis reveals consistent trophic growth of juvenile Atlantic goliath grouper Epinephelus itajara in Brazilian estuaries. Bull Mar Sci. 2022; 98(1):17–26. https://doi.org/10.5343/bms.2021.0025
Antoro S, Na-Nakorn U, Koedprang W. Study of genetic diversity of orange-spotted grouper, Epinephelus coioides, from Thailand and Indonesia using microsatellite markers. Mar Biotech. 2006; 8:17–26. https://doi.org/10.1007/s10126-005-5026-0
Berry O, England P, Fairclough D, Jackson G, Greenwood JIM. Microsatellite DNA analysis and hydrodynamic modelling reveal the extent of larval transport and gene flow between management zones in an exploited marine fish (Glaucosoma hebraicum). Fish Oceanogr. 2012; 21(4):243–54. https://doi.org/10.1111/j.1365-2419.2012.00623.x
Bertoncini AA, Aguilar-Perera A, Barreiros J, Craig MT, Ferreira BP, Koenig CC. Epinephelus itajara (errata version published in 2019). The IUCN Red List of Threatened Species. 2018; 2018:e.T195409A145206345. https://doi.org/10.2305/IUCN.UK.2018-2.RLTS. T195409A145206345.en
Bueno LS, Bertoncini AA, Koenig CC, Coleman FC, Freitas MO, Leite JR et al. Evidence for spawning aggregations of the endangered Atlantic goliath grouper Epinephelus itajara in southern Brazil. J Fish Biol. 2016; 89(1):76–889. https://doi.org/10.1111/jfb.13028
Bullock LH, Murphy MD, Godcharles MF, Mitchell ME. Age, growth, and reproduction of jewfish Epinephelus itajara in the eastern Gulf of Mexico. Fish Bull. 1992; 90(2):243–49. Available from: https://spo.nmfs.noaa.gov/sites/default/files/pdf-content/1992/902/bullock.pdf
Burimski O, Vasemägi A, Gross R. Changes in the spatio-temporal genetic structure of Baltic sea trout (Salmo trutta L.) over two decades: direct and indirect effects of stocking. Conserv Gen. 2024; 25:481–97. https://doi.org/10.1007/s10592-023-01582-7
Coleman FC, Koenig CC, Huntsman GR, Musick JA, Eklund AM, McGovern JC et al. Long-lived reef fishes: the grouper-snapper complex. Fisheries. 2000; 25(3):14–21. https://doi.org/10.1577/1548-8446(2000)025<0014:LRF>2.0.CO;2
Condini MV, Garcia AM, Bastos RF, Vollrath SR, Barbosa EF, Oliveira RL et al. Hydroclimatic influence on the trophic ecology of Atlantic goliath grouper juveniles (Epinephelus itajara) in a tropical estuary using non-lethal methodologies. Mar Environ Res. 2023; 190:106103. https://doi.org/10.1016/j.marenvres.2023.106103
Condini MV, Oliveira RL, Almeida LL, Barbosa EF, Martins AC, Bertoncini AA et al . Physicochemical variables influencing the abundance of Atlantic goliath grouper juveniles (Epinephelus itajara) in a tropical Brazilian mangrove habitat. Mar Environ Res. 2024; 197:106465. https://doi.org/10.1016/j.marenvres.2024.106465
Cornuet JM, Luikart G. Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics. 1996; 144(4):2001–14. https://doi.org/10.1093/genetics/144.4.2001
Cowen RK, Sponaugle S. Larval dispersal and marine population connectivity. Annu. Rev Mar Sci. 2009; 1(1):443–66. https://doi.org/10.1146/annurev.marine.010908.163757
Damasceno JS, Farro APC, Bertoncini AA, Hostim-Silva M. Atlantic Goliath groupers as inhabitants of mangroves on the Brazilian Coast. Reg Stud Mar Sci. 2025; 84:104092. https://doi.org/10.1016/j.rsma.2025.104092
Damasceno JS, Siccha-Ramirez R, Morales MJA, Oliveira C, Torres RA, Costa EN et al. Mitochondrial DNA evidences reflect an incipient population structure in Atlantic goliath grouper (Epinephelus itajara, Epinephelidae) in Brazil. Sci Mar. 2015; 79(4):419–29. http://dx.doi.org/10.3989/scimar.04203.20A
Diaz-Suarez A, Noreikiene K, Kisand V, Burimski O, Svirgsden R, Rohtla M et al. Temporally stable small-scale genetic structure of Northern pike (Esox lucius) in the coastal Baltic Sea. Fish Res. 2022; 254:106402. https://doi.org/10.1016/j.fishres.2022.106402
DiRenzo A, Peterson AC, Garza JC, Valdes AM, Slatkin M, Freimer NB. Mutational processes of simple-sequence repeat loci in human populations. PNAS. 1994; 91(8):3166–70. https://doi.org/10.1073/pnas.91.8.3166
Do C, Waples RS, Peel D, Macbeth GM, Tillett BJ, Ovenden JR. NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour. 2014; 14(1):209–14. https://doi.org/10.1111/1755-0998.12157
Earl DA, vonHoldt BM. Structure Harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour. 2012; 4:359–61. https://doi.org/10.1007/s12686-011-9548-7
Eklund AM, Schull J. A stepwise approach to investigating the movement patterns and habitat utilization of goliath grouper, Epinephelus itajara, using conventional tagging, acoustic telemetry and satellite tracking. In: Sibert JR, Nielsen JL, editors. Electronic tagging and tracking in marine fisheries, vol 1. Netherlands: Springer; 2001. p.189–216. https://doi.org/10.1007/978-94-017-1402-0_9
Ellis RD, Koenig CC, Coleman FC. Spawning-related movement patterns of goliath grouper (Epinephelus itajara) off the Atlantic Coast of Florida. Proc Gulf Caribb Res Inst. 2014; 66:395–400.
Evanno G, Regnaut S, Goudet J. Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol. 2005; 14(8):2611–20. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Excoffier L, Lischer HEL. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour. 2010; 10(3):564–67. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Félix-Hackradt FC, Hackradt CW. Populational study and monitoring of the goliath grouper, Epinephelus itajara (Lichtenstein, 1822), in the coast of Paraná, Brazil. Nat Conserv. 2008; 6(2):141–56.
Francini-Filho RB, Cordeiro MC, Omachi CY, Rocha AM, Bahiense L, Garcia GD et al. Remote sensing, isotopic composition and metagenomics analyses revealed Doce River ore plume reached the southern Abrolhos Bank Reefs. Sci Total Environ. 2019; 697:134038. https://doi.org/10.1016/j.scitotenv.2019.134038
Frankham R, Bradshaw CJ, Brook BW. Genetics in conservation management: revised recommendations for the 50/500 rules, Red List criteria and population viability analyses. Biol Conserv. 2014; 170:56–63. https://doi.org/10.1016/j.biocon.2013.12.036
Freitas MO, Abilhoa V, Giglio VJ, Hostim-Silva M, Moura RL, Francini-Filho RB et al. Diet and reproduction of the goliath grouper, Epinephelus itajara (Actinopterygii: Perciformes: Serranidae), in eastern Brazil. Acta Ichthyol Piscat. 2015; 45(1):1–11. https://doi.org/10.3750/AIP2015.45.1.01
Gerhardinger LC, Marenzi RC, Bertoncini AA, Medeiros RP, Hostim-Silva M. Local Ecological Knowledge on the Goliath Grouper Epinephelus itajara (Teleostei: Serranidae) in Southern Brazil. Neotrop Ichthyol. 2006; 4(4):441–50. https://doi.org/10.1590/S1679-62252006000400008
Giglio VJ, Bender MG, Zapelini C, Ferreira CEL. The end of the line? Rapid depletion of a large-sized grouper through spearfishing in a subtropical marginal reef. Perspect Ecol Conserv. 2017; 15(2):115–18. https://doi.org/10.1016/j.pecon.2017.03.006
Giglio VJ, Bertoncini AA, Ferreira BP, Hostim-Silva M, Freitas MO. Landings of goliath grouper, Epinephelus itajara, in Brazil: despite prohibited over ten years, fishing continues. Perspect Ecol Conserv. 2014; 12(2):118–23. https://doi.org/10.1016/j.ncon.2014.09.004
Giglio VJ, Leite JR, Freitas MO, Hostim-Silva M. Mapping goliath grouper aggregations in the southwestern Atlantic. Braz J Oceanogr. 2016; 64(4):417–20. http://dx.doi.org/10.1590/S1679-87592016122906404
Goudet J. FSTAT, a program to estimate and test gene diversities and fixation indices. 2001. Available from: http://www.unil.ch/izea/softwares/fstat.html
Hassanien HA, Al-Rashada Y. Assessment of genetic diversity and phylogenetic relationship among grouper species Epinephelus spp. from the Saudi waters of the Arabian Gulf. Saudi J Biol Sci. 2021; 28(3):1779–86. https://doi.org/10.1016/j.sjbs.2020.12.020
Heemstra PC, Randall JE. Groupers of the world. FAO Fisheries Synopsis. Rome: Food and Agriculture Organization. 1993; 16:382. Available from: https://www.fao.org/4/t0540e/t0540e00.htm
Herrera M, Nanninga GB, Planes S, Jones GP, Thorrold SR, Saenz-Agudelo P et al. Seascape and life-history traits do not predict self-recruitment in a coral reef fish. Biol Lett. 2016; 12(8):4–07. https://doi.org/10.1098/rsbl.2016.0309
Hostim-Silva M, Bertoncini AA, Gerhardinger LC, Machado LF. The “Lord of the Rock’s” conservation program in Brazil: the need for a new perception of marine fishes. Coral Reefs. 2005; 24(1):74. https://doi.org/10.1007/s00338-004-0437-3
Instituto Brasileiro do Meio Ambiente e dos Recursos Naturais Renováveis (IBAMA). Portaria IBAMA N° 121, de 20 de setembro de 2002. [Internet]. 2002. Available from: https://www.icmbio.gov.br/cepsul/images/stories/legislacao/Portaria/2002/p_ibama_121_2002_moratoria5anospescadomero_alterada_p_ibama_42_2007.pdf
Instituto Brasileiro do Meio Ambiente e dos Recursos Naturais Renováveis (IBAMA). Portaria IBAMA No 42, de 19 de setembro de 2007. [Internet] 2007. Available from: https://www.icmbio.gov.br/cepsul/images/stories/legislacao/Portaria/2007/p_ibama_42_2007_defesomero_altera_p_ibama_121_2002.pdf
Instituto Chico Mendes de Conservação da Biodiversidade e Ministério do Meio Ambiente (ICMBio/MMA). Livro Vermelho da Fauna Brasileira Ameaçada de Extinção: Vol. VI – Peixes. [Internet] 2018. Available from: https://www.gov.br/icmbio/pt-br/centrais-de-conteudo/publicacoes/publicacoes-diversas/livro_vermelho_2018_vol1.pdf
Jackson AM, Semmens BX, Mitcheson YS, Nemeth RS, Heppell SA, Bush PG et al. Population structure and phylogeography in Nassau grouper (Epinephelus striatus), a mass-aggregating marine fish. PLoS ONE. 2014; 9(5):e97508. https://doi.org/10.1371/journal.pone.0097508
Jeannot LL, Mouronvalle C, Peyran C, Olabarria C, Beger M, Blanco A et al. Deriving connectivity from relatedness: broad-scale isolation-by-distance in the shanny Lipophrys pholis. Mar Ecol Progr Ser. 2024; 731:9–26. https://doi.org/10.3354/meps14459
Jones OR, Wang J. COLONY: a program for parentage and sibship inference from multilocus genotype data. Mol Ecol Resour. 2010; 10(3):551–55. https://doi.org/10.1111/j.1755-0998.2009.02787.x
Jørgensen HBH, Hansen MM, Bekkevold D, Ruzzante DE, Loeschke V. Marine landscapes and population genetic structure of herring (Clupea harengus) in the Baltic Sea. Mol Ecol. 2005; 14(10):3219–34. https://doi.org/10.1111/j.1365–294X.2005.02658.x
Kalinowski ST, Taper ML, Marshall TC. Revising how the computer program Cervus accommodates genotyping error increases success in paternity assignment. Mol Ecol. 2007; 16(5):1099–106. https://doi.org/10.1111/j.1365-294X.2007.03089.x
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012; 28(12):1647–49. https://doi.org/10.1093/bioinformatics/bts199
Kinghorn B, Kinghorn S. Pedigree viewer version 5.5c. University of New England, Australia. 2009. Available from: https://bkinghor.une.edu.au/pedigree.htm
Koenig C, Bueno L, Coleman F, Cusick J, Ellis R, Kingon K et al. Diel, lunar, and seasonal spawning patterns of the Atlantic goliath grouper, Epinephelus itajara, off Florida, United States. Bull Mar Sci. 2017; 93(2):391–406. https://doi.org/10.5343/bms.2016.1013
Koenig CC, Coleman FC. Population density, demographics, and predation effects of adult goliath grouper. Tallahassee. 2009. Available from: https://www.reef.org/sites/default/files/SEDAR23_RD_19_GoliathFINAL.pdf
Koenig CC, Coleman FC, Eklund AM, Schull J, Ueland J. Mangroves as essential nursery habitat for goliath grouper (Epinephelus itajara). Bull Mar Sci. 2007; 80(3):567–86.
Koenig CC, Coleman FC, Kingon K. Pattern of recovery of the goliath grouper Epinephelus itajara population in the southeastern US. Bull Mar Sci. 2011; 87(4):891–911. https://doi.org/10.5343/bms.2010.1056
Kopelman NM, Mayzel J, Jakobsson M, Rosenberg NA, Mayrose I. Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol Ecol Res. 2015; 15(5):1179–91. https://doi.org/10.1111/1755-0998.12387
Lara MR, Schull J, Jones DL, Allman R. Early life history stages of goliath grouper Epinephelus itajara (Pisces: Epinephelidae) from Ten Thousand Islands, Florida. Endanger Species Res. 2009; 7(3):221–28. https://doi.org/10.3354/esr00193
Larsson LC, Laikre L, André C, Dahlgren TG, Ryman N. Temporally stable genetic structure of heavily exploited Atlantic herring (Clupea harengus) in Swedish waters. Heredity. 2010; 104(1):40–51. https://doi.org/10.1038/hdy.2009.98
Leão ZMAN, Kikuchi RKP, Testa V. Corals and coral reefs of Brazil. In: Cortes J, editor. Latin America coral reefs. Elsevier, Oxford; 2003. p.9–52. https://doi.org/10.1016/B978-044451388-5/50003-5
Liu BJ, Zhang BD, Xue DX, Gao TX, Liu JX. Population structure and adaptive divergence in a high gene flow marine fish: the small yellow croaker (Larimichthys polyactis). PLoS ONE. 2016; 11(4):e0154020. https://doi.org/10.1371/journal.pone.0154020
Locatelli ACP, Bastos RF, Oliveira MA, Ferreira BP. Scientometric analysis and literature synthesis of 60 years of science on the Atlantic goliath grouper (Epinephelus itajara). J Fish Biol. 2023; 102(4):740–56. https://doi.org/10.1111/jfb.15312
López A, Vera M, Planas M, Bouza C. Conservation genetics of threatened Hippocampus guttulatus in vulnerable habitats in NW Spain: temporal and spatial stability of wild populations with flexible polygamous mating system in captivity. PLoS ONE. 2015; 10:e0117538. https://doi.org/10.1371/journal.pone.0117538
Malinowski CR, Coleman FC, Koenig CC, Locascio JV, Murie, DJ. Are Atlantic goliath grouper, Epinephelus itajara, establishing more northerly spawning sites? Evidence from the northeast Gulf of Mexico. Bull Mar Sci. 2019; 95(3):371–91. https://doi.org/10.5343/bms.2018.0062
Malinowski CR, Perrault JR, Coleman FC, Koenig CC, Stilwell JM, Cray C et al. The iconic Atlantic goliath grouper (Epinephelus itajara): a comprehensive assessment of health indices in the southeastern United States population. Front Vet Sci. 2020; 7:635. https://doi.org/10.3389/fvets.2020.00635
Mann DA, Locascio JV, Coleman FC, Koenig CC. Goliath grouper (Epinephelus itajara) sound production and movement patterns on aggregation sites. Endanger Species Res. 2009; 7(3):229–36. https://doi.org/10.3354/esr00109
Melero Y, Oliver MK, Lambin X. Relationship type affects the reliability of dispersal distance estimated using pedigree inferences in partially sampled populations: A case study involving invasive American mink in Scotland. Mol Ecol. 2017; 26(15):4059–71. https://doi.org/10.1111/mec.14154
Ministério do Meio Ambiente e Ministério da Pesca e Aquicultura (MMA/MPA). Instrução Normativa Interministerial no 13, de 16 de outubro de 2012. [Internet] 2012. Available from: https://www.icmbio.gov.br/cepsul/images/stories/legislacao/Instrucao_normativa/2012/in_inter_mpa_mma_13_2012_defesomero.pdf
Ministério do Meio Ambiente (MMA). Portaria n° 445, de 17 de dezembro de 2014. [Internet] 2014. Available from: https://www.icmbio.gov.br/cepsul/images/stories/legislacao/Portaria/2014/p_mma_445_2014_lista_peixes_amea%C3%A7ados_extin%C3%A7%C3%A3o.pdf
Ministério do Meio Ambiente e Ministério da Pesca e Aquicultura (MMA/MPA). Portaria Interministerial no 13, de 2 de outubro de 2015. [Internet] 2015. Available from: https://www.icmbio.gov.br/cepsul/images/stories/legislacao/Portaria/2015/p_mpa_mma_13_2015_proibe_pesca_mero.pdf
Moreno-Santos RA, Condini MV, Almeida LL, Soeth M, Bertoncini AA, Hostim-Silva M. Unveiling the first steps of juvenile Atlantic goliath grouper Epinephelus itajara (Perciformes: Epinephelidae) in a tropical estuary at the southwestern Atlantic coast. Mar Biol Res. 2022; 18(5–6):372–79. https://doi.org/10.1080/17451000.2022.2119251
Murie DJ, Parkyn DC, Koenig CC, Coleman FC, Malinowski CR, Cusick JA et al. Age, growth, and functional gonochorism with a twist of diandric protogyny in goliath grouper from the Atlantic Coast of Florida. Fishes. 2023; 8(8):412. https://doi.org/10.3390/fishes8080412
Nance HA, Klimley P, Galván-Magaña F, Martínez-Ortíz J, Marko PB. Demographic processes underlying subtle patterns of population structure in the scalloped hammerhead shark, Sphyrna lewini. PLOS ONE. 2011; 6(7):e21459. https://doi.org/10.1371/journal.pone.0021459
Østergaard S, Hansen MM, Loeschcke V, Nielsen EE. Long‐term temporal changes of genetic composition in brown trout (Salmo trutta L.) populations inhabiting an unstable environment. Mol Ecol. 2003; 12(11):3123–35. https://doi.org/10.1046/j.1365-294X.2003.01976.x
Petrou EL, Fuentes-Pardo AP, Rogers LA, Orobko M, Tarpey C, Jiménez-Hidalgo I et al. Functional genetic diversity in an exploited marine species and its relevance to fisheries management. Proc Biol Sci. 2021; 288(1945):20202398. https://doi.org/10.1098/rspb.2020.2398
Pritchard JK, Stephens M, Donnelly P. Inference of population structure using multilocus genotype data. Genetics. 2000; 155(2):945–59. https://doi.org/10.1093/genetics/155.2.945
Raymond M, Rousset F. GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered. 1995; 86:248–49. https://doi.org/10.1093/oxfordjournals.jhered.a111573
Robitzch V, Saenz-Agudelo P, Berumen ML. Travel with your kin ship! Insights from genetic sibship among settlers of a coral damselfish. Ecol Evol. 2020; 10(15):8265–78. https://doi.org/10.1002/ece3.6533
Rousset F. Genepop’007: a complete reimplementation of the Genepop software for Windows and Linux. Mol Ecol Resour. 2008; 8(1):103–06. https://doi.org/10.1111/j.1471-8286.2007.01931.x
Sadovy Y. Grouper stocks of the western central Atlantic: the need for management and management needs. Proceedings of the 43rd Gulf and Caribbean Fisheries Institute. 1994; 43–64. Available from: https://sedarweb.org/documents/s23rd24-grouper-stocks-of-the-western-central-atlantic-the-need-for-management-and-management-needs/
Sadovy Y, Eklund AM. Synopsis of biological information on the Nassau grouper, Epinephelus striatus (Bloch 1792), and the jewfish, E. itajara (Lichtenstein 1822). NOAA Technical Report, NMFS 146, and FAO Fisheries Synopsis. 1999; 157:65. Available from: https://repository.library.noaa.gov/view/noaa/3090
Sánchez-Velásquez JJ, Pinedo-Bernal PN, Reyes-Flores LE, Yzásiga-Barrera C, Zelada-Mázmela E. Genetic diversity and relatedness inferred from microsatellite loci as a tool for broodstock management of fine flounder Paralichthys adspersus. Aquac Fish. 2022; 7(6):664–74. https://doi.org/10.1016/j.aaf.2021.06.008
Schunter C, Carreras-Carbonell J, Planes S, Sala E, Ballesteros E, Zabal M, Harmelin JG et al. Genetic connectivity patterns in an endangered species: The dusky grouper (Epinephelus marginatus). J Exp Mar Biol Ecol. 2011; 401(1–2):126–33. https://doi.org/10.1016/j.jembe.2011.01.021
Seyoum S, Tringali MD, Barthel BL, Puchulutegui C, Davis MC, Collins AB et al. Isolation and characterization of 29 polymorphic microsatellite markers for the endangered Atlantic goliath grouper (Epinephelus itajara), and the Pacific goliath grouper (E. quinquefasciatus). Conserv Genet Resour. 2013; 5:729–32. http://dx.doi.org/10.1007/s12686-013-9892-x
Silva-Oliveira GC, Rêgo PS, Schneider H, Sampaio I, Vallinoto M. Genetic characterization of populations of the critically endangered Goliath grouper (Epinephelus itajara, Serranidae) from the Northern Brazilian coast through analyses of mtDNA. Genet Mol Biol. 2008; 31:988–95. http://dx.doi.org/10.1590/S1415-47572008005000016
Snead AA, Tatarenkov A, Avise JC, Taylor DS, Turner BJ, Marson K et al. Out to sea: ocean currents and patterns of asymmetric gene flow in an intertidal fish species. Front Genet. 2023; 14:1206543. http://dx.doi.org/10.3389/fgene.2023.1206543
Taboun ZS, Walter RP, Ovenden JR, Heath DD. Spatial and temporal genetic variation in an exploited reef fish: The effects of exploitation on cohort genetic structure. Evol Appl. 2021; 14(5):1286–300. http://dx.doi.org/10.1111/eva.13198
Tognella MMP, Leopoldo RVS, Oliveira CP, Pascoalini SS, Silva ED. Diversidade estrutural das florestas de mangue da costa central e norte do Espírito Santo: contribuições para entendimento de funções ecossistêmicas. Enciclopédia Biosfera. 2020; 17(33):178–93. https://doi.org/10.18677/EnciBio_2020C16
Tringali, MD. Reproductive success dynamics could limit precision in close-kin mark–recapture abundance estimation for Atlantic goliath grouper (Epinephelus itajara). Fishes. 2023; 8(5):254. https://doi.org/10.3390/fishes8050254
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P. MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes. 2004; 4(3):535–38. https://doi.org/10.1111/j.1471-8286.2004.00684.x
Wang J. Computationally efficient sibship and parentage assignment from multilocus marker data. Genetics. 2012; 191(1):183–94. https://doi.org/10.1534/genetics.111.138149
Wang J, Santure AW. Parentage and sibship inference from multilocus genotype data under polygamy. Genetics. 2009; 181(4):1579–94. https://doi.org/10.1534/genetics.108.100214
Waples RS. A generalized approach for estimating effective population size from temporal changes in allele frequency. Genetics. 1989; 121(2):379–91. https://doi.org/10.1093/genetics/121.2.379
Waples RS, Do C. Linkage disequilibrium estimates of contemporary Ne using highly variable genetic markers: a largely untapped resource for applied conservation and evolution. Evol Appl. 2010; 3(3):244–62. https://doi.org/10.1111/j.1752-4571.2009.00104.x
Weng Z, Yang Y, Wang X, Wu L, Hua S, Zhang H et al. Parentage analysis in gian grouper (Epinephelus lanceolatus) using microsatellite and SNP markers from genotyping-by-sequencing data. Genes. 2021; 12(7):1042. https://doi.org/10.3390/genes12071042
Wenne R. Microsatellites as molecular markers with applications in exploitation and conservation of aquatic animal populations. Genes. 2023; 14(4):808. https://doi.org/10.3390/genes14040808
Zapelini C, Giglio VJ, Carvalho RC, Bender MG, Gerhardinger LC. Assessing fishing experts knowledge to improve conservation strategies for an endangered grouper in the southwestern Atlantic. J Ethnobiol. 2017; 37(3):478–93. https://doi.org/10.2993/0278-0771-37.3.478
Authors
![]() Lorena Lopes Almeida1,2,3
Lorena Lopes Almeida1,2,3 ![]() ,
, ![]() Maurício Hostim-Silva1,2,4,5 and
Maurício Hostim-Silva1,2,4,5 and ![]() Ana Paula Cazerta Farro1,3,4
Ana Paula Cazerta Farro1,3,4
[1] Programa de Pós-Graduação em Biologia Animal (PPGBAN), Universidade Federal do Espírito Santo (UFES), Departamento de Ciências Biológicas, Av. Fernando Ferrari, 514, Goiabeiras, 29075-910 Vitória, ES, Brazil. (LLA) lorenalopesbio@gmail.com (corresponding author), (MHS) mhostim@gmail.com, (APCF) farro.ana@gmail.com.
[2] Laboratório de Ecologia de Peixes Marinhos (LEPMAR), Centro Universitário Norte do Espírito Santo, (CEUNES), Departamento de Ciências Agrárias e Biológicas, Rodovia BR-101, km 60, Litorâneo, 29932-540 São Mateus, ES, Brazil.
[3] Laboratório de Genética e Conservação Animal, Centro Universitário Norte do Espírito Santo (CEUNES), Departamento de Ciências Agrárias e Biológicas, Rodovia BR-101, km 60, Litorâneo, 29932-540 São Mateus, ES, Brazil.
[4] Programa de Pós-Graduação em Oceanografia Ambiental (PPGOAM), Universidade Federal do Espírito Santo (UFES), Av. Fernando Ferrari, 514, Goiabeiras, 29075-910 Vitória, ES, Brazil.
[5] Instituto Meros do Brasil (PMB), Benjamin Constant 67, Conj. 1104 10° andar, 80060-020 Curitiba, PR, Brazil.
Authors’ Contribution 

Lorena Lopes Almeida: Conceptualization, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Writing-original draft, Writing-review and editing.
Maurício Hostim-Silva: Conceptualization, Funding acquisition, Methodology, Project administration, Resources, Supervision, Writing-review and editing.
Ana Paula Cazerta Farro: Conceptualization, Funding acquisition, Investigation, Methodology, Project administration, Resources, Supervision, Visualization, Writing-review and editing.
Ethical Statement
All sample collections were performed under license number SISBIO 15080–7.
Competing Interests
The author declares no competing interests.
How to cite this article
Almeida LL, Hostim-Silva M, Farro APC. Temporal genetic monitoring of juvenile Atlantic goliath grouper Epinephelus itajara (Perciformes: Epinephelidae) on the Brazilian central coast. Neotrop Ichthyol. 2025; 23(2):e250008. https://doi.org/10.1590/1982-0224-2025-0008
Copyright
This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Distributed under
Creative Commons CC-BY 4.0

© 2025 The Authors.
Diversity and Distributions Published by SBI
![]() Accepted April 3, 2025
Accepted April 3, 2025
![]() Submitted January 27, 2025
Submitted January 27, 2025
![]() Epub July 11, 2025
Epub July 11, 2025