![]() Carlos Arturo Amado-García1
Carlos Arturo Amado-García1 ![]() ,
, ![]() Iván Yesid López-Ardila1 and
Iván Yesid López-Ardila1 and ![]() Fernando Rondón-González1
Fernando Rondón-González1
PDF: Download Here | Cite this article
Associate Editor: ![]() José Igor da Silva
José Igor da Silva
Section Editor: ![]() Yasmim de Santana
Yasmim de Santana
Editor-in-chief: ![]() José Birindelli
José Birindelli
Abstract
Potamotrygon magdalenae es la única especie de raya de agua dulce reportada como endémica de Colombia, habitando las cuencas de los ríos Magdalena-Cauca, San Jorge, Atrato y Catatumbo. Si bien se comprenden aspectos ecológicos, reproductivos y bioquímico, su genética poblacional ha sido poco estudiada, especialmente dado las distintas amenazas que enfrenta. Para evaluar la estructura poblacional de P. magdalenae, analizamos secuencias de ADN mitocondrial del gen MT-CYB de 60 individuos muestreados en tres ciénagas de la cuenca media del río Magdalena. Nuestro análisis de secuencias de 693 pb reveló 24 sitios variables en 22 haplotipos (hd = 0.83277, π = 0.00246). Se observó un flujo génico significativo entre las ciénagas, lo que apoya la hipótesis de que P. magdalenae constituye una única población panmíctica (ΦST = -0.0083, p = 0.68035). Las pruebas de neutralidad de Tajima (D = -1.62, p < 0.05) y Fu (Fs = -4.59, p < 0.02) indican una reciente expansión poblacional tras un evento de cuellos de botella, probablemente debido a la degradación del hábitat de la especie, a pesar de las limitaciones del marcador. No se encontró diferenciación genética en el área de estudio de 200 km, y las estimaciones genéticas fueron comparables a las de otras especies de peces endémicas del río Magdalena.
Palabras clave: Citocromo B, Genética de poblaciones, Haplotipo, Panmixia, Raya de agua dulce.
Introduction
The family Potamotrygonidae (Neotropical stingrays) comprises around 40 species of fish, including both marine and freshwater organisms (Loboda et al., 2021; Fricke et al., 2024). Of all these, the subfamily Potamotrygoninae stands out, whose origin and diversification as the only living lineage of freshwater-adapted elasmobranchs within this family have been linked to historical marine incursions in northern South America (Lovejoy et al., 1998; 2006; Bloom, Lovejoy, 2011; Fontenelle et al., 2021).
Potamotrygon magdalenae (Duméril, 1865) is the only stingray species endemic to Colombia, inhabits both lentic, and lotic systems and has been recorded in the Atrato, Magdalena, Cauca, San Jorge, and Catatumbo rivers (Lasso et al., 2013). It typically occurs in shallow, turbid waters with muddy substrates (Dahl, 1971; Lasso et al., 2013). Its diet consists primarily of aquatic macroinvertebrates, particularly larval-stage insects of the order Diptera (Mejía-Falla et al., 2016; Márquez-Velásquez et al., 2019). Several aspects of its reproductive biology have been described (Pedreros-Sierra, Ramírez-Pinilla, 2015; Pedreros-Sierra et al., 2016; Anaya-López, Ramírez-Pinilla, 2017; Lizcano-Gutiérrez, Ramírez-Pinilla, 2022), along with studies on its ecological and biochemical aspects (Singer, Ballantyne, 1989; Ramos-Socha, Grijalba-Bendeck, 2011; Márquez-Velásquez et al., 2019; Pérez-Rojas et al., 2022).
However, few studies have addressed the population genetics of this species, particularly in relation to other Potamotrygon species (García et al., 2016; Batista-Morales et al., 2017; Renza-Millán et al., 2019; Rizo-Fuentes et al., 2021). Regarding the assessment of the population genetic component of other indigenous fish species distributed within the Magdalena River basin, such as striped catfish (Pseudoplatystoma magdaleniatum) and bocachico (Prochilodus magdalenae), significant levels of genetic variation have been documented utilizing microsatellite molecular markers and mitochondrial gene segments (Aguirre-Pabón et al., 2013; Berdugo, Narváez, 2014; Landínez-García et al., 2020; García-Castro et al., 2021).
As is the case with other elasmobranchs, P. magdalenae exhibits slow reproductive cycles and low offspring numbers, leading to limited population growth (Araújo et al., 2004; Mejía-Falla et al., 2009). The species faces a high risk of extinction due to several threats, including overfishing, habitat degradation and destruction, commercial fishing practices, and mining activities (Lasso et al., 2013). Given this information, the International Union for Conservation of Nature (IUCN) reports a declining trend in population size, leading to the classification of the species as Near Threatened (NT). If populations continue to decline, the species could become endangered (IUCN, 2025). The limited understanding of biological aspects may result in an underestimation of the extent to which these organisms are threatened. A similar classification was observed in the Red Book of Freshwater Fishes of Colombia, where the species is classified as NT at the national level (Mojica et al., 2012). Conversely, the National Plan of Action for the Conservation and Management of Sharks, Rays, and Chimeras in Colombia (PAN-Colombia) emphasizes the formulation and implementation of strategies to safeguard these species, given their high-priority status (Caldas et al., 2010).
To gain an understanding of the population dynamics of the endemic Colombian stingray, P. magdalenae, the present study aimed to examine aspects of its population genetics based on maternal inheritance. We collected samples from three swamps in the Magdalena River sub-basins. This study utilizes mitochondrial DNA sequences to analyze genetic diversity, population structure, and historical demographic events.
Material and methods
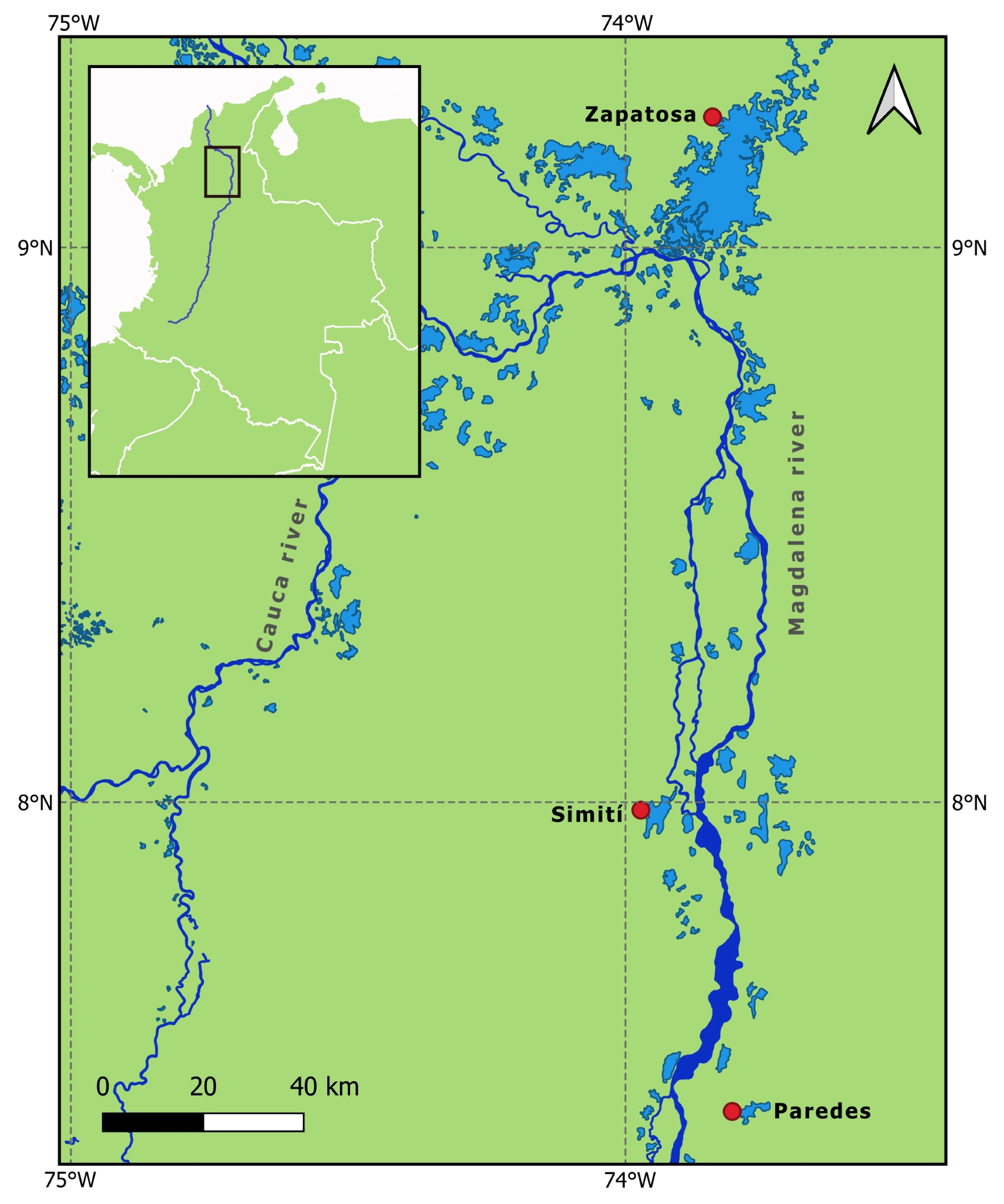
Study area and fish sampling. This study analyzed the distribution of P. magdalenae across three key swamps: the Zapatosa complex, Simití, and Paredes (Fig. 1). We assessed samples obtained from 20 individuals in each swamp. Zapatosa and Paredes swamps showed the maximum straight-line distance (199 km), while Simití and Paredes had the shortest (60 km). The surveyed swamps are primarily surrounded by tropical dry forests (García, Dister, 1990; Cruz et al., 2009), except for Simití, which lies within tropical rainforests (Vilardy et al., 2014).
FIGURE 1| Map of the sampling area for Potamotrygon magdalenae in the middle Magdalena River basin in northeastern Colombia. Swamps are shown in lighter blue, and the main stream of rivers Magdalena and Cauca are shown in darker blue. The red circles indicate the assessed swamps: the Zapatosa swamp complex (09°15’00.82”N 73°48’29.08”W), the Simití swamp (07°57’19.07”N 73°57’05.33”W), and the Paredes swamp (07°26’52.60”N 73°47’34.93”W). The WGS84 system was used.
Fishing techniques, such as trammel nets and cast nets, were employed to capture individuals (Gama et al., 2013). We supplemented our collection with specimens contributed by residents. A total of 60 specimens of P. magdalenae were meticulously identified, according to the guides of Maldonado-Ocampo et al. (2005) and Lasso et al. (2011). Blood samples of 1 mL were collected from each individual from the caudal artery using syringes and then deposited into tubes containing an equal volume of absolute ethanol. After drawing blood samples, we photographed and marked all captured individuals with plastic bands to prevent resampling during potential recapture events, then released them back into the swamps. We preserved these samples at 4 °C and stored them in the Laboratorio de Genética y Biotecnología (LGyB-UIS) at the Universidad Industrial de Santander.
DNA extraction and PCR amplification. The DNA extraction protocol involved treating each sample with a 10% SDS solution and Proteinase K before nucleic acid extraction was carried out (Green, Sambrook, 2012), with adjustments made from LGyB-UIS, which includes an additional wash with phenol:chloroform:isoamyl alcohol (25:24:1).
Before amplifying the cytochrome b (MT-CYB) fragments through Polymerase Chain Reaction (PCR), the concentration and quality were assessed using a Nanodrop 2000 spectrophotometer (Thermo Scientific, USA). The runs were then carried out on a 1% agarose gel to verify the integrity of the DNA. We diluted all samples to 100 ng/µL for optimal PCR performance.
We amplified the mitochondrial MT-CYB gene using the primers and standardized conditions described by López-Ardila et al. (2022). A 750 bp band was visualized and verified by 1% agarose gel electrophoresis using an EZ-Vision® intercalating agent (Amresco) and a GeneRuler 1 kb DNA Ladder (Thermo Scientific, USA).
The amplicons were subsequently purified and subjected to sequencing at the Servicio de Secuenciación y Análisis Molecular (SSiGMol) at the Universidad Nacional de Colombia. This service utilizes BigDye Terminator v. 3.1 (Applied Biosystems) technology. We deposited the MT-CYB sequences in GenBank under accessions PP261281 to PP261302.
Population genetic analysis. MT-CYB gene fragments were visualized and manually edited using CodonCode Aligner v. 1.2.2 (CodonCode Co.). We retained sequences with Phred scores greater than 30 (Ewing et al., 1998; Shi et al., 2016) for analysis. Subsequently, the sequences were aligned using Mega v. 10.1.8 software (Kumar et al., 2018), employing the default Muscle algorithm parameters (Edgar, 2004). Haplotype diversity (hd), nucleotide diversity (π), and the identification of polymorphic sites were determined using DnaSP v. 6.12.03 (Rozas et al., 2017). We visualized DNA sequence variations through median-joining haplotype networks (Leigh, Bryant, 2015; PopART v. 10.1.0.0).
Genetic differentiation between population pairs (ΦST) and gene flow estimation (Nm) were analyzed to assess the genetic structure of P. magdalenae across the sampled swamps. We first tested the metapopulation genetic structure hypothesis by performing an analysis of molecular variance (AMOVA) across three sampling locations. We additionally tested the isolation-by-distance hypothesis using a Mantel test. To evaluate the demographic history, neutrality tests were performed, including Tajima’s D (Tajima, 1989) and Fu’s Fs (Fu, 1997). We visualized differences in genotyped haplotype distributions through Mismatch Distribution analysis in Arlequin v. 3.5 (Excoffier, Lischer, 2010).
We used JModelTest v. 2 (Darriba et al., 2012) to determine the best-fit nucleotide substitution model for MT-CYB sequences. We constructed a Bayesian Skyline Plot (BSP) in BEAST v. 2.5 (Bouckaert et al., 2019) with 100 million generations to assess contemporary population expansion in P. magdalenae. We visualized this assessment using Tracer v. 1.7.1 was present in the studied waters with a frequency of 26%. POMACb3 and POMACb5 (each with a 3% frequency) were shared exclusively between Zapatosa and Paredes. Additionally, we identified 15 locality-excluding (private) haplotypes, each occurring at a frequency of 0.0166% within a single swamp (Tab. 1). The haplotype network showed no local clustering, indicating gene exchange between individuals from different swamps (Fig. 2).
TABLE 1 | Haplotypes and haplotypic frequency based on a fragment of the MT-CYB gene from Potamotrygon magdalenae. The mitogenome NC_036041 was used as a reference, located at the top of the table in bold. Hap = haplotypes; f (sd) = haplotypic frequency (standard deviation); Z = Zapatosa; S = Simití; P = Paredes; Acc = Accession number in the GenBank. The Position refers to the specific cite inside of gene in which nucleotide variability is encountered. *Nomenclature used in the network in Fig. 2.
| Position |
| |||||||||||||||||||||||||
1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | ||||
4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 5 | 5 | 5 | 5 | 5 | 5 | 5 | 5 | 5 | 5 | 5 | 5 | ||||
6 | 6 | 6 | 6 | 6 | 7 | 7 | 7 | 8 | 8 | 8 | 9 | 0 | 1 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 3 | 3 | ||||
3 | 3 | 4 | 7 | 9 | 1 | 2 | 4 | 6 | 7 | 9 | 5 | 6 | 2 | 2 | 3 | 5 | 0 | 2 | 5 | 6 | 9 | 0 | 1 | ||||
1 | 7 | 7 | 2 | 5 | 1 | 5 | 7 | 7 | 7 | 7 | 1 | 2 | 1 | 5 | 3 | 8 | 9 | 4 | 0 | 1 | 7 | 2 | 4 | ||||
Haplotype (nomenclature*) | G | C | A | A | A | G | A | T | A | A | C | T | A | T | T | C | C | T | C | G | A | A | A | A | f | sd | Swamp |
POMACb1 (H1) | . | G | . | . | . | A | . | C | . | G | . | . | . | . | . | . | . | . | . | . | . | . | . | . | 0.0166 | 0.0166 | Z |
POMACb2 (H2) | . | . | . | . | . | . | . | C | . | . | . | . | . | . | . | T | . | . | . | . | . | . | . | . | 0.0333 | 0.0233 | Z |
POMACb3 (H3) | . | . | . | . | . | A | . | C | . | . | . | C | . | . | . | . | . | . | . | . | . | . | . | . | 0.0333 | 0.0233 | Z, P |
POMACb4 (H4) | . | . | . | . | . | A | . | C | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | 0.2666 | 0.0575 | Z, S, P |
POMACb5(H5) | . | G | . | . | . | A | . | C | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | 0.0333 | 0.0233 | Z, P |
POMACb6(H6) | . | . | . | T | . | A | . | C | . | . | . | . | . | . | . | . | . | . | G | . | . | . |
| . | 0.0166 | 0.0166 | Z |
POMACb7(H7) | . | . | . | . | . | . | . | C | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | 0.3166 | 0.0605 | Z, S, P |
POMACb8(H8) | A | . | . | . | . | A | . | C | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | 0.0333 | 0.0233 | Z |
POMACb9(H9) | . | . | . | . | . | A | . | C | . | . | . | C | G | . | . | . | . | . | . | . | . | . | . | . | 0.0166 | 0.0166 | Z |
POMACb10(H10) | . | G | G | . | . | . | . | C | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | 0.0166 | 0.0166 | Z |
POMACb11(H11) | . | . | . | . | T | A | G | C | . | . | . | . | . | . | . | . | . | . | . | . | G | G | G | . | 0.0166 | 0.0166 | Z |
POMACb12(H12) | . | . | . | . | . | . | . | C | . | . | . | . | . | . | . | . | . | . | . | . | . | . | G | . | 0.0166 | 0.0166 | S |
POMACb13(H13) | . | G | . | . | . | . | . | C | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | 0.0333 | 0.0233 | S |
POMACb14(H14) | . | . | . | . | . | A | . | C | . | . | T | C | . | . | . | . | . | . | . | . | . | . | . | C | 0.0166 | 0.0166 | S |
POMACb15(H15) | . | . | . | . | . | . | . | C | . | . | . | . | . | A | . | . | . | . | . | . | . | . | . | . | 0.0166 | 0.0166 | S |
POMACb16(H16) | . | . | . | . | . | A | . | C | . | . | . | . | . | . | . | . | . | . | . | A | . | . | . | . | 0.0166 | 0.0166 | S |
POMACb17(H17) | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | 0.0166 | 0.0166 | S |
POMACb18(H18) | . | . | . | T | . | A | . | C | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | . | 0.0166 | 0.0166 | S |
POMACb19(H19) | . | . | . | . | . | A | . | C | . | . | . | . | . | . | C | . | T | . | . | . | . | . | . | . | 0.0166 | 0.0166 | S |
POMACb20(H20) | . | . | . | . | . | A | . | C | . | . | . | . | . | . | . | . | T | . | . | . | . | . | . | . | 0.0166 | 0.0166 | P |
POMACb21(H21) | . | . | . | . | . | A | . | C | . | . | . | . | . | . | . | . | . | C | . | . | . | . | . | . | 0.0166 | 0.0166 | P |
POMACb22(H22) | . | . | . | . | . | A | . | C | T | . | . | . | . | . | C | . | . | . | . | . | . | . | . | . | 0.0166 | 0.0166 | P |
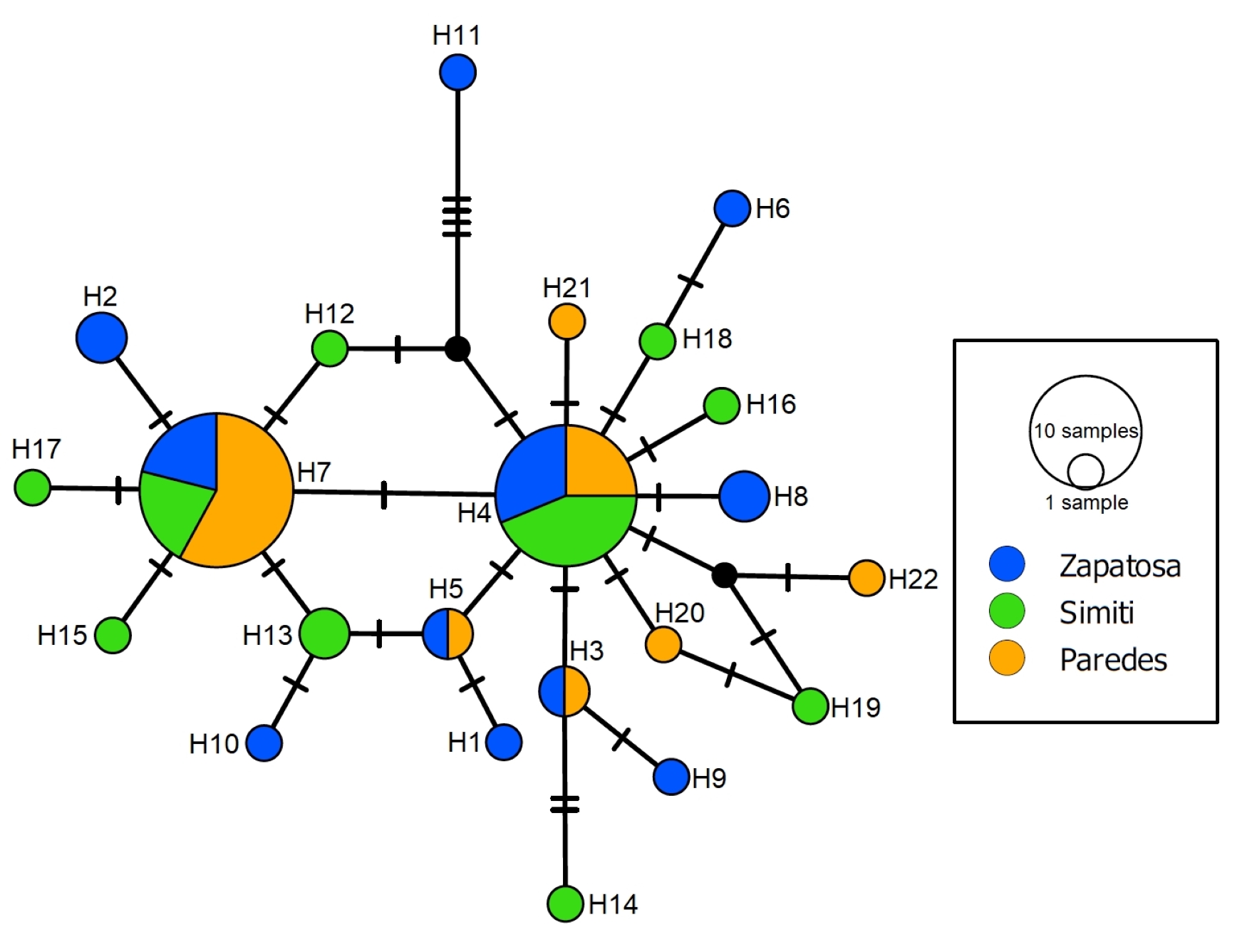
FIGURE 2| Haplotype network based on partial sequences of the MT-CYB gene of Potamotrygon magdalenae. Black circles indicate unsampled haplotypes within the study, and cross lines represent mutational changes between haplotypes.
The haplotype network revealed POMACb7 and POMACb4 in all three swamps, with POMACb3 and POMACb5 present in both the Zapatosa and Paredes swamps. Of the 15 individual haplotypes, seven were reported in the Simití swamp, five in the Zapatosa swamp, and three in the Paredes swamp (Fig. 2).
Based on the MT-CYB sequences obtained, the haplotype diversity (hd) ranged from 0.67895 (Paredes) to 0.90526 (Zapatosa) with a mean value of 0.83277 ± 0.036, while the nucleotide diversity (π) oscillated between 0.00162 (Paredes) and 0.00334 (Zapatosa) with a mean value of 0.00246 ± 0.001547 (Tab. 2).
TABLE 2 | Haplotype and nucleotide diversity by locality from MT-CYB gene sequences assessed in Potamotrygon magdalenae. Genetic diversity values with their respective standard deviations. n = number of sequences, s = polymorphic sites, H = number of haplotypes, hd = haplotype diversity, π = nucleotide diversity, D = Tajima’s neutrality test, Fs = Fu neutrality test. *Significant values – p value < 0.05. **Significant values – p value < 0.02.
Swamp | n | s | H | hd | π | D | Fs |
Zapatosa | 20 | 15 | 11 | 0.90526 ± 0.0441 | 0.00334 ± 0.002130 | -1.669* | -5.241** |
Simití | 20 | 12 | 10 | 0.85263 ± 0.0630 | 0.00247 ± 0.001679 | -1.774* | -5.419** |
Paredes | 20 | 7 | 7 | 0.67895 ± 0.1024 | 0.00162 ± 0.001228 | -1.412 | -3.122** |
General | 60 | 24 | 22 | 0.83277 ± 0.036 | 0.00246 ± 0.001547 | -1.621* | -4.594** |
Gene flow and genetic differentiation. Gene flow (Nm) of females in P. magdalenae, estimates ranged from 40 (Zapatosa-Paredes) to ∞ (Simití-Zapatosa/Simití-Paredes), indicating high migration rates. Pairwise ΦST values were low (Simití-Zapatosa: -0.014; Simití-Paredes: -0.026; Zapatosa-Paredes: 0.012) (Tab. 3). AMOVA attributed 95% of P. magdalenae genetic variation to differences within swamps. Together with these obtained values of genetic differentiation, the haplotype network presented in Fig. 2 does not show a concentration of haplotypes with in swamp. The Mantel test revealed a positive correlation between genetic and geographic distance, although it does not support the isolation by distance hypothesis in the Magdalena stingray (R² = 0.9268; p = 0.190).
TABLE 3 | Gene flow and genetic differentiation from partial sequences of the MT-CYB gene of Potamotrygon magdalenae. Above the diagonal: paired values of Nm; below the diagonal: paired values of genetic differentiation (ΦST). On the diagonal, the values of Tajima’s D and Fu’s Fs neutrality tests. *p < 0.05; **p < 0.02.
Swamp | Zapatosa | Simití | Paredes |
Zapatosa |
D
= -1.66903* | ∞ | 39.8 |
Simití | -0.01419 |
D
= -1.77436* | ∞ |
Paredes | 0.01240 | -0.02593 |
D
= -1.41981 |
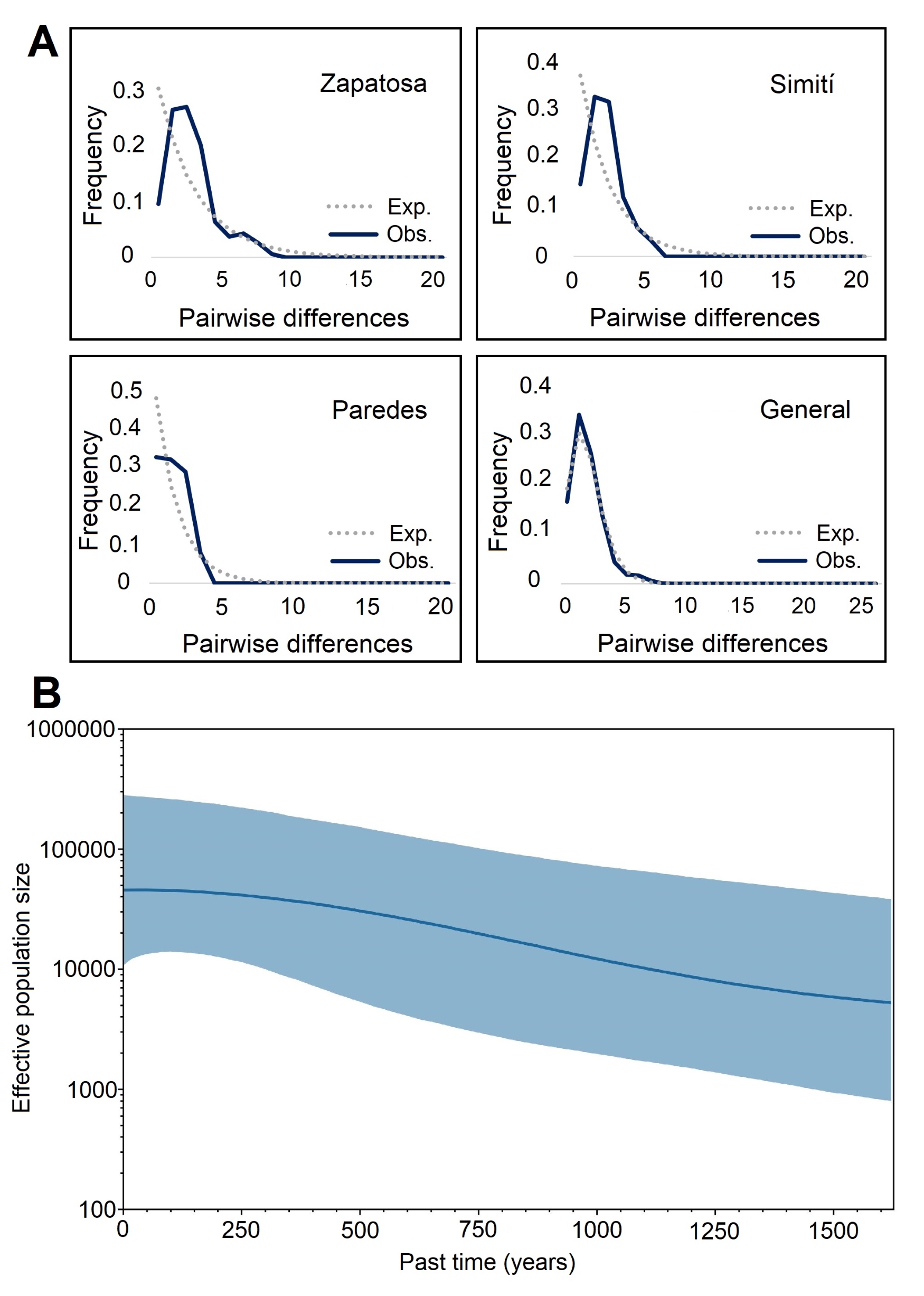
Neutrality tests and mismatch analysis support recent population expansion in P. magdalenae. Tajima’s D and Fu’s Fs values were significantly negative (Tab. 2; p < 0.02), and the unimodal mismatch distribution (Fig. 3A) matched expectations for expansion. The Bayesian skyline plot (Fig. 3B) estimated a gradual increase in effective population size over the last 1,500 years, corroborating this trend.
FIGURE 3| Demographic history of Potamotrygon magdalenae using partial MT-CYB gene sequences. A. Distribution of nucleotide differences between haplotype pairs. X-axis: paired nucleotide differences, Y-axis: difference frequency. B. Bayesian Skyline Plot analysis. X-axis: effective population size, Y-axis: time in years. The solid blue line represents the mean, and the blue shaded area indicates the 95% highest posterior density.
Results
NOME DE ESPÉCIE,
A
Discussion
This study expands current knowledge on the population genetics of the Colombian freshwater stingray, Potamotrygon magdalenae, using 693 bp of the mitochondrial gene MT-CYB sequences. Our findings expand the limited genetic knowledge of P. magdalenae, a species that has been previously addressed in only four studies (Torres et al., 2022).
The genetic parameters estimated in this research indicate that P. magdalenae exhibits high haplotype diversity and low nucleotide diversity. This characteristic pattern, consistent with the findings of Hilsdorf, Hallerman (2017) for Neotropical fishes, suggests a recent population expansion following a historical bottleneck event.
This demographic bottleneck could reflect a combination of natural factors (Aguirre et al., 2021), overfishing (Restrepo et al., 2021), water pollution (Gómez-Martínez et al., 2024), and abortions of pregnant females (Lizcano-Gutiérrez, Ramírez-Pinilla, 2022), as well as historical processes that have shaped the species population. However, it is important to investigate how anthropogenic processes may impact the species and the Magdalena River ecosystem in the future. Pollution and contamination represent threats to aquatic biodiversity, particularly to ichthyofauna (Gallo-Vélez et al., 2022), which further exacerbates the challenges faced by P. magdalenae.
This high haplotype diversity and low nucleotide diversity patterns align with findings reported for other potamotrygonid stingrays across northern South America, i.e., Paratrygon aiereba (Frederico et al., 2012; Rizo-Fuentes et al., 2021) and multiple Potamotrygon species (P. motoro, Renza-Millán et al., 2019; P. orbignyi, Batista-Morales et al., 2017; P. scobina, Toffoli, 2006).
Ramírez-Hernández (2018) reported significantly lower genetic diversity (h = 0.47 ± 0.59; π= 0.001 ± 0.0002) using 549-bp MT-COI sequences in P. magdalenae (Tab. 2). This discrepancy from the pattern described previously may reflect sampling dissimilarity between studies and unequal sample distribution across localities, as well as properties intrinsic to the marker (Fratini et al., 2016). For studies assessing the same population, it is recommended to use between 10 and 20 samples to obtain reliable estimates of genetic structure (Grünwald et al., 2017).
In P. magdalenae, the gene sequence analysis of MT-CYB revealed a latitudinal gradient, where genetic diversity decreased from north to south, with peak diversity observed in the northern Zapatosa swamp (Fig. 1; Tab. 2). This spatial pattern supports the hypothesis proposed by Lovejoy et al. (1998) regarding the incursion of marine stingrays into northern South American freshwater systems during the Miocene. According to this hypothesis, marine-derived elasmobranchs underwent adaptive freshwater colonization through introgressive processes (Smith, 1992). The widespread distribution of the POMACb7 haplotype (C14747T), which shows minimal sequence variation across all sampled swamps, suggests it may represent the ancestral lineage of Magdalena basin populations. The prevalence and conservation of this haplotype across the species’ range will indicate long-term stability in the face of historical environmental changes.
Studies of genetic diversity in other Magdalena River endemic fishes, such as Prochilodus magdalenae (Landínez-García et al., 2020; García-Castro et al., 2021) and Pseudoplatystoma magdaleniatum (Aguirre-Pabón et al., 2013), P. magdalenae exhibits high genetic diversity maintained through the river’s role as a migratory corridor (García-Castro et al., 2024). In this sense, the Nm estimated and AMOVA results (95% within-swamp variation; Tab. 3) support the idea of the Magdalena River as a migratory corridor for P. magdalenae, in addition to favoring the idea of panmixia along the 200 km of the study area, despite the non-significant isolation-by-distance pattern (Mantel test: R² = 0.93, p = 0.19) (Diniz-Filho et al., 2013). These findings contrast with the MT-COI-based study by Ramírez-Hernández (2018), which did not report any geographic structure.
Key factors likely contribute to this genetic homogeneity, such as life history traits (Toffoli, 2006; Ramírez-Hernández, 2018), historical factors (Frederico et al., 2012; Batista-Morales et al., 2017; Renza-Millán et al., 2019), and anthropogenic impacts (Lasso et al., 2016), all of which could be responsible for the absence of genetic structure in P. magdalenae, as confirmed by the haplotype network. In addition to shared haplotypes, POMACb4 and POMACb7 are found in all swamps, while POMACb3 and POMACb5 are detected in two swamps, indicating female-mediated gene flow even across different swamps. This pattern of gene flow suggests that while private haplotypes may exist, they do not restrict overall dispersal within the population. These results contrast with those of Ramírez-Hernández (2018), who identified many private haplotypes.
Different evidence suggests recent population expansion in P. magdalenae. Genetic diversity metrics (Tab. 2), neutrality tests (Tab. 3), a star-like haplotype network topology (Fig. 2), and demographic reconstructions, including a unimodal mismatch distribution and a Bayesian Skyline Plot showing an expansion over approximately 1,500 years (Fig. 3) (Drummond et al., 2005). We also emphasize that these signals should be interpreted with caution, and that a multilocus approach would be advisable to overcome the potential influence of factors such as sample size and differences in gene-specific evolutionary rates, which may reflect distinct evolutionary histories (Zamora-Mejías et al., 2022). Our findings provide evidence in support of the population expansion approach, also noted for the MT-COI gene (Ramírez-Hernández, 2018). This same trend has been observed in other endemic fish species from the Magdalena River (Aguirre et al., 2013).
The results led to the conclusion based on its maternal inheritance of MT-CYB that P. magdalenae: i) has not shown significant genetic differentiation and have high contemporary gene flow; ii) post-bottleneck expansion is supported by Bayesian Skyline Plot (≈1500-year) and iii) parallels genetics and demographic patterns compared to other Magdalena River migratory fishes (Prochilodus magdalenae, Pseudoplatystoma magdaleniatum). The validation of these primers, coupled with evidence of panmictic gene flow, establishes molecular tools to monitor population dynamics and genetics in endemic stingrays.
This study provides further insights into the genetic and demographic structure of Potamotrygon magdalenae, offering valuable guidance for conservation strategies within the Magdalena River basin. While mitochondrial markers enable the detection of maternal lineages and recent demographic events, their uniparental inheritance and limited genomic scope constrain the ability to reconstruct a population’s genetic history fully. To overcome these limitations, future research should integrate nuclear and male parental inherited markers using a multilocus approach and extend sampling efforts to include poorly characterized portions of the Magdalena and Atrato rivers, particularly in the southern and northern basin areas influenced by anthropogenic interventions. Such expanded analyses would help clarify and complement our understanding of population structure and gene flow, considering the distinct evolutionary dynamics revealed by each marker type.
Acknowledgments
We would like to thank the various local fishermen from each of the swamps for their assistance with the fishing procedures and their hospitality.
References
Aguirre WE, Alvarez-Mieles G, Anaguano-Yancha F, Burgos Morán R, Cucalón RV, Escobar-Camacho D et al. Conservation threats and future prospects for the freshwater fishes of Ecuador: A hotspot of Neotropical fish diversity. J Fish Biol. 2021; 99(4):1158 –89. https://doi.org/10.1111/jfb.14844
Aguirre-Pabón JC, Narváez JC, Castro LF. Mitochondrial DNA variation of the bocachico Prochilodus magdalenae (Characiformes, Prochilodontidae) in the Magdalena River Basin, Colombia. Aquat Conserv Mar Freshw Ecosyst. 2013; 23(4):594–605. https://doi.org/10.1002/aqc.2339
Anaya-López P, Ramírez-Pinilla M. Clasper gland morphology and development in Potamotrygon magdalenae (Elasmobranchii: Potamotrygonidae). J Morphol. 2017; 278(3):369–79. https://doi.org/10.1002/jmor.20647
Araújo M, Charvet-Almeida P, Almeida M, Pereira H. Freshwater stingrays (Potamotrygonidae): status, conservation and management challenges. Information document AC. 2004; 20(8):1–06.
Batista-Morales A, Lasso CA, Morales-Betancourt M. Phylogeography and genetic structure of the species complex of the freshwater stingray, Potamotrygon orbignyi (Castelnau, 1855), among Amazonas and Orinoco rivers [Master dissertation]. Bogotá, Colombia: Universidad de los Andes; 2017.
Berdugo GO, Narváez Barandica JC. Genetic diversity and population structure of bocachico Prochilodus magdalenae (Pisces, Prochilodontidae) in the Magdalena River basin and its tributaries, Colombia. Genet Mol Biol. 2014; 37(1):37–45. https://doi:10.1590/s1415-47572014000100008
Bloom DD, Lovejoy NR. The biogeography of marine incursions in South America. In: Albert JS, Reis RE, editors. Historical biogeography of neotropical freshwater fishes. Berkeley: University of California Press; 2011. p.137–44.
Bouckaert R, Vaughan TG, Barido-Sottani J, Duchêne S, Fourment M, Gavryushkina A et al. BEAST 2.5: an advanced software platform for Bayesian evolutionary analysis. PLoS Comput Biol. 2019; 15(4):e1006650. https://doi.org/10.1371/journal.pcbi.1006650
Caldas J, Castro-Gonzalez E, Puentes V, Rueda M. Plan de acción nacional para la conservación y manejo de tiburones, rayas y quimeras de Colombia (PAN Tiburones Colombia) [Internet]. Bogotá: Instituto Colombiano Agropecuario; 2010. Available from: http://minagricultura.gov.co
Cruz M, Estupiñán A, Jiménez-Escobar N, Sánchez N, Galeano G, Linares E. Etnobotánica de la región tropical del Cesar, Complejo Ciénaga de Zapatosa. In: Colombia diversidad Biótica VIII: media y baja montaña de la Serranía del Perijá. Bogotá, Colombia: Universidad Nacional de Colombia; 2009. p.417–74.
Dahl G. Los peces del norte de Colombia. Bogotá: Instituto de Desarrollo de los Recursos Naturales Renovables; 1971.
Darriba D, Taboada GL, Doallo R, Posada D. jModelTest 2: more models, new heuristics and parallel computing. Nat Methods. 2012; 9(8):772. https://doi.org/10.1038/nmeth.2109
Diniz-Filho JA, Soares TN, Lima JS, Dobrovolski R, Landeiro VL, Campos Telles MP et al. Mantel test in population genetics. Genet Mol Biol. 2013; 36(4):475–85. https://doi.org/10.1590/S1415-47572013000400002
Drummond AJ, Rambaut A, Shapiro B, Pybus OG. Bayesian coalescent inference of past population dynamics from molecular sequences. Mol Biol Evol. 2005; 22(5):1185–92. https://doi.org/10.1093/molbev/msi103
Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004; 32(5):1792–97. https://doi.org/10.1093/nar/gkh340
Ewing B, Hillier L, Wendl MC, Green P. Base-calling of automated sequencer traces using phred. I. Accuracy assessment. Genome Res. 1998; 8(3):175–85. https://doi.org/10.1101/gr.8.3.175
Excoffier L, Lischer HEL. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour. 2010; 10(3):564–67. https://doi.org/10.1111/j.1755-0998.2010.02847.x
Fratini S, Ragionieri L, Deli T, Harrer A, Marino IAM, Cannicci S et al. Unravelling population genetic structure with mitochondrial DNA in a notional panmictic coastal crab species: sample size makes the difference. BMC Evol Biol. 2016; 16:150. https://doi.org/10.1186/s12862-016-0720-2
Frederico RG, Farias IP, Araújo MLG, Charvet P, Alves-Gomes JA. Phylogeography and conservation genetics of the Amazonian freshwater stingray Paratrygon aiereba Müller & Henle, 1841 (Chondrichthyes: Potamotrygonidae). Neotrop Ichthyol. 2012; 10(1):71–80. https://doi.org/10.1590/S1679-62252012000100007
Fontenelle JP, Marques FPL, Kolmann MA, Lovejoy NR. Biogeography of the neotropical freshwater stingrays (Myliobatiformes: Potamotrygoninae) reveals effects of continent-scalepaleogeographic changeand drainage evolution. J Biogeogr. 2021; 48(6):1406–19. https://doi.org/10.1111/jbi.14086
Fricke R, Eschmeyer WN, Fong JD. Eschmeyer’s catalog of fishes: genera, species, references [Internet]. San Francisco: California Academy of Science; 2024. Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp
Fu YX. Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics. 1997;147(2):915–25. https://doi.org/10.1093/genetics/147.2.915
Gallo-Velez D, Restrepo JC, Newton A. A socio-ecological assessment of land-based contamination and pollution: The Magdalena delta, Colombia. Front Mar Sci. 2022; 9:1057426. https://doi.org/10.3389/fmars.2022.1057426
Gama CS, Araújo MLG, Lasso CA, Morales-Betancourt MA, Sánchez-Duarte P, Guzmán A. Métodos para la captura y análisis de datos bioecológicos de las rayas de agua dulce (Potamotrygonidae). In: Lasso CA, Rosa RS, Sánchez-Duarte P, Morales-Betancourt MA, Agudelo-Córdoba E, editors. Rayas de agua dulce (Potamotrygonidae) de Suramérica. Bogotá, Colombia: Instituto Alexander von Humboldt; 2013. p.65–84.
Garcia DA, Lasso CA, Morales M, Caballero SJ. Molecular systematics of the freshwater stingrays (Myliobatiformes: Potamotrygonidae) of the Amazon, Orinoco, Magdalena, Esequibo, Caribbean, and Maracaibo basins (Colombia – Venezuela): evidence from three mitochondrial genes. Mitochondrial DNA A. 2016; 27(6):4479–91. https://doi.org/10.3109/19401736.2015.1101536
Garcia LC, Dister E. La planicie de inundación del Medio-bajo Magdalena: restauración y conservación de habitats. Interciencia. 1990; 15(6):396–410.
García-Castro KL, Márquez EJ. Temporal analysis of genetic diversity and gene flow in the threatened catfish Pseudoplatystoma magdaleniatum from a dammed neotropical river. PLoS One. 2024;19(4):e0301577. https://doi.org/10.1371/journal.pone.0301577
García-Castro KL, Rangel-Medrano JD, Landínez-García RM, Márquez EJ. Population genetics of the endangered catfish Pseudoplatystoma magdaleniatum (Siluriformes: Pimelodidae) based on species-specific microsatellite loci. Neotrop Ichthyol. 2021; 19(1):e200120. https://doi.org/10.1590/1982-0224-2020-0120
Gómez-Martínez D, Londoño-Cruz E, Mejía-Falla PA. Upper thermal tolerance and population implications for the Magdalena River stingray Potamotrygon magdalenae. J Fish Biol. 2024; 104(6):1698–708. https://doi.org/10.1111/jfb.15668
Green MR, Sambrook J. Molecular cloning: a laboratory manual. 4th ed. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 2012.
Grünwald NJ, Everhart SE, Knaus BJ, Kamvar ZN. Best practices for population genetic analyses. Phytopathology. 2017; 107(9):1000–10. https://doi.org/10.1094/PHYTO-12-16-0425-RVW
Hilsdorf A, Hallerman E. Genetic resources of Neotropical fishes. Cham: Springer International Publishing; 2017. https://doi.org/10.1007/978-3-319-55838-7
International Union for Conservation of Nature (IUCN). The IUCN Red List of Threatened Species. Version 2025-1 [Internet]. Gland; 2025. Available from: https://www.iucnredlist.org
Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol Biol Evol. 2018; 35(6):1547–49. https://doi.org/10.1093/molbev/msy096
Landínez-García RM, Narváez JC, Márquez EJ. Population genetics of the freshwater fish Prochilodus magdalenae (Characiformes: Prochilodontidae), using species-specific microsatellite loci. PeerJ. 2020; 8:e10327. https://doi.org/10.7717/peerj.10327
Lasso CA, Agudelo E, Córdoba L, Jiménez-Segura F, Ramírez-Gil H, Morales-Betancourt M et al. Catálogo de los recursos pesqueros continentales de Colombia. Bogotá, Colombia: Instituto Alexander von Humboldt; 2011.
Lasso CA, Rosa RS, Morales-Betancourt M, Garrone-Neto D, Carvalho MR. Rayas de agua dulce (Potamotrygonidae) de Sudamérica. Parte II: Colombia, Brasil, Perú, Bolivia, Paraguay, Uruguay y Argentina. Bogotá, Colombia: Instituto Alexander von Humboldt; 2016.
Lasso CA, Rosa RS, Sánchez-Duarte P, Morales-Betancourt MA, Agudelo-Córdoba E. IX. Rayas de agua dulce (Potamotrygonidae) de Suramérica. Parte I. Colombia, Venezuela, Ecuador, Perú, Brasil, Guyana, Surinam y Guayana Francesa: diversidad, bioecología, uso y conservación. Bogotá: Instituto de Investigación de Recursos Biológicos Alexander von Humboldt; 2013.
Leigh JW, Bryant D. POPART: Full-feature software for haplotype network construction. Methods Ecol Evol. 2015; 6(9):1110–16. https://doi.org/10.1111/2041-210X.12410
Lizcano-Gutiérrez SA, Ramírez-Pinilla MP. Caracterización morfológica de los estadios embrionarios de la raya dulceacuícola Potamotrygon magdalenae (Elasmobranchii: Potamotrygonidae). Rev Acad Colomb Cienc Exactas Fis Nat. 2022; 46(180):697–713. https://doi.org/10.18257/raccefyn.1650
Loboda TS, Lasso CA, Rosa RDS, Carvalho MRD. Two new species of freshwater stingrays of the genus Paratrygon (Chondrichthyes: Potamotrygonidae) from the Orinoco basin, with comments on the taxonomy of Paratrygon aiereba. Neotrop Ichthyol. 2021; 19(2):e200083. https://doi.org/10.1590/1982-0224-2020-0083
López-Ardila I, Martínez-Pérez F, Rondón-González F. Aplicación del modelo de pérdida de ADN para el diseño de cebadores en Potamotrygon magdalenae (Potamotrygonidae). Acta Biol Colomb. 2021; 27(1):97–103. https://doi.org/10.15446/abc.v27n1.87401
Lovejoy NR, Albert JS, Crampton WGR. Miocene marine incursions and marine/freshwater transitions: evidence from Neotropical fishes. J South Am Earth Sci. 2006;21(1–2):5–13. https://doi.org/10.1016/j.jsames.2005.07.009
Lovejoy NR, Bermingham E, Martin AP. Marine incursion into South America. Nature. 1998; 396(6710):421–22. https://doi.org/10.1038/24757
Maldonado-Ocampo JA, Ortega-Lara A, Usma JS, Galvis G, Villa-Navarro FA, Vásquez L et al. Peces de los Andes de Colombia. Bogotá, Colombia: Instituto Alexander von Humboldt; 2005.
Márquez-Velásquez V, Rosa RS, Galindo E, Navia AF. Feeding habits and ecological role of the freshwater stingray Potamotrygon magdalenae (Duméril 1865) (Myliobatiformes: Potamotrygonidae), combining gut-content and stable isotope analysis. Environ Biol Fish. 2019; 102:1119–36. https://doi.org/10.1007/s10641-019-00897-0
Mejía-Falla P, Pérez-Rojas J, Gómez-Martínez D, Muñoz-Osorio L, Galindo E, López-García J et al. Avances en el conocimiento de Potamotrygon magdalenae (Dumeril, 1865), raya dulceacuícola endémica de Colombia. In: Lasso C, Rosa R, Morales-Betancourt M, Garrone-Neto D, Carvalho M, editors. XV. Rayas de agua dulce (Potamotrygonidae) de Suramérica. Parte II: Colombia, Brasil, Perú, Bolivia, Paraguay, Uruguay y Argentina. Bogotá: Instituto de Investigación de Recursos Biológicos Alexander von Humboldt; 2016. p.305–23.
Mejía-Falla P, Ramírez-Luna V, Usma-Oviedo J, Muñoz-Osorio L, Maldonado-Ocampo J, Sanabria A et al. Estado del conocimiento de las rayas dulceacuícolas de Colombia. In: Puentes V, Navia A, Mejía-Falla P, Caldas J, Diazgranados M, Zapata-Padilla L, editors. Avances en el conocimiento de tiburones, rayas y quimeras de Colombia. Bogotá: Fundación SQUALUS; 2009. p.197–245.
Mojica JI, Usma-Oviedo JS, Alvarez-León R, Lasso CA. Libro rojo de peces dulceacuícolas de Colombia. Bogotá: Instituto de Investigación de Recursos Biológicos Alexander von Humboldt; 2012.
Pedreros-Sierra T, Arrieta-Prieto D, Mejía-Falla P. Reproductive system of females of the Magdalena river endemic stingray Potamotrygon magdalenae: anatomical and functional aspects. J Morphol. 2016; 277(5):680–97. https://doi.org/10.1002/jmor.20527
Pedreros-Sierra T, Ramírez-Pinilla M. Morphology of the reproductive tract and acquisition of sexual maturity in males of Potamotrygon magdalenae (Elasmobranchii: Potamotrygonidae). J Morphol. 2015; 276(3):273–89. https://doi.org/10.1002/jmor.20337
Pérez-Rojas J, Mejía-Falla P, Navia A, Tarazona A, Pardo-Carrasco S. Hematology and blood biochemistry profile of the freshwater stingray Potamotrygon magdalenae as a tool for population assessment in artificial environments. Braz J Biol. 2022; 82:e233780. https://doi.org/10.1590/1519-6984.233780
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol. 2018; 67(5):901–04. https://doi.org/10.1093/sysbio/syy032
Ramírez-Hernández AM. Variabilidad genética de la raya de agua dulce Potamotrygon magdalenae (Elasmobranchii: Potamotrygonidae) en el Río Magdalena, Colombia [Master dissertation]. México: Universidad Nacional Autónoma de México; 2018.
Ramos-Socha HB, Grijalba-Bendeck M. Bioecología de la raya de agua dulce Potamotrygon magdalenae (Duméril, 1865) (Myliobatiformes) en la ciénaga de Sabayo, Guaimaral. Colombia. Rev UDCA Actual Divulg Cient. 2011; 14(2):109–18.
Renza-Millán M, Lasso CA, Morales-Betancourt MA, Villa F, Caballero SJ. Mitochondrial DNA diversity and population structure of the ocellate freshwater stingray Potamotrygon motoro (Müller & Henle, 1841) (Myliobatiformes: Potamotrygonidae) in the Colombian Amazon and Orinoco Basins. Mitochondrial DNA A. 2019; 30(3):466–73. https://doi.org/10.1080/24701394.2018.1546300
Restrepo J, Cárdenas-Rozo AL, Paniagua-Arroyave JF, Jiménez-Segura LF. Aspectos físicos de la cuenca del río Magdalena, Colombia: geología, hidrología, sedimentos, conectividad, ecosistemas acuáticos e implicaciones para la biota. In: Jiménez-Segura LF, Lasso CA, editors. Peces de la cuenca del río Magdalena, Colombia: diversidad, conservación y uso sostenible. Bogotá, Colombia: Instituto Alexander von Humboldt; 2021. p.41–82.
Rizo-Fuentes MA, Correa-Cárdenas CA, Lasso CA, Morales-Betancourt MA, Barragán-Barrera DC, Caballero S. Phylogeography, genetic diversity and population structure of the freshwater stingray, Paratrygon aiereba (Müller & Henle, 1841) (Myliobatiformes: Potamotrygonidae) in the Colombian Amazon and Orinoco basins. Mitochondrial DNA Part A. 2021; 32(1):20–33. https://doi.org/10.1080/24701394.2020.1844679
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins S et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol Biol Evol. 2017; 34(12):3299–302. https://doi.org/10.1093/molbev/msx248
Singer T, Ballantyne J. Absence of extrahepatic lipid oxidation in a freshwater elasmobranch, the dwarf stingray Potamotrygon magdalenae: evidence from enzyme activities. J Exp Zool. 1989; 251(3):355–60. https://doi.org/10.1002/jez.1402510312
Shi H, Li W, Xu X. Learning the comparing and converting method of sequence phred quality score. In: Proceedings of the International Conference on Management, Education, Information and Control (MEICI 2016). Amsterdam: Atlantis Press; 2016. p.260–63. https://doi.org/10.2991/meici-16.2016.55
Smith GR. Introgression in fishes: significance for paleontology, cladistics, and evolutionary rates. Syst Biol. 1992; 41(1):41–57. https://doi.org/10.1093/sysbio/41.1.41
Tajima F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics. 1989; 123(3):585–95. https://doi.org/10.1093/genetics/123.3.585
Toffoli D. História evolutiva de espécies do gênero Potamotrygon Garman, 1877 (Potamotrygonidae) na Bacia Amazônica [Master Dissertation]. Manaus: Universidade Federal do Amazonas; 2006.
Torres Y, Faria VV, Charvet P. Current status and future perspectives of Neotropical freshwater stingrays (Potamotrygoninae, Myliobatiformes) genetics. Environ Biol Fish. 2022; 105:1111–27. https://doi.org/10.1007/s10641-022-01320-x
Vilardy SP, Cortés-Duque J. Los Humedales de Cantagallo, San Pablo y Simití: una propuesta para su delimitación desde el enfoque de los sistemas socioecológicos. Bogotá, Colombia: Instituto Alexander von Humboldt; 2014.
Zamora-Mejías D, Trejo-Salazar RE, Eguiarte LE, Ojeda M, Rodríguez-Herrera B, Morales-Malacara JB et al. Traveler mites: population genetic structure of the wing mites Periglischrus paracaligus (Acari: Mesostigmata: Spinturnicidae). J Med Entomol. 2022; 59(4):1198–210. https://doi.org/10.1093/jme/tjac059
Authors
![]() Carlos Arturo Amado-García1
Carlos Arturo Amado-García1 ![]() ,
, ![]() Iván Yesid López-Ardila1 and
Iván Yesid López-Ardila1 and ![]() Fernando Rondón-González1
Fernando Rondón-González1
[1] Grupo de Investigación en Microbiología y Genética, Escuela de Biología, Facultad de Ciencias, Universidad Industrial de Santander, Carrera 27 calle 9, 680002, Bucaramanga, Santander, Colombia. (CAAG) carlos.amado2@correo.uis.edu.co, (IYLA) ivan2228062@correo.uis.edu.co (corresponding author), (FRG) ferongon@uis.edu.co.
Authors’ Contribution 

Carlos Arturo Amado-García: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Writing-original draft, Writing review and editing.
Iván Yesid López-Ardila: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Writing-original draft, Writing review and editing.
Fernando Rondón-González: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Project administration, Resources, Supervision, Writing-original draft, Writing review and editing.
Ethical Statement
Not applicable.
Competing Interests
The author declares no competing interests.
Data availability statement
The authors confirm that the data supporting the findings of this study are available within the article.
AI statement
The authors did not use any AI-assisted technologies in the creation of this manuscript or its figures.
Funding
Programa de apoyo a la movilidad del Portafolio de programas de la Vicerrectoría de Investigación y Extensión de la Universidad Industrial de Santander.
How to cite this article
Amado-García CA, López-Ardila IY, Rondón-González F. Mitochondrial genetic variation of endemic stingray Potamotrygon magdalenae (Myliobatiformes: Potamotrygonidae) in the middle Magdalena River basin, Colombia. Neotrop Ichthyol. 2026; 24(1):e250127. https://doi.org/10.1590/1982-0224-2025-0127
Copyright
This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Distributed under
Creative Commons CC-BY 4.0

© 2025 The Authors.
Diversity and Distributions Published by SBI
![]() Accepted November 1, 2025
Accepted November 1, 2025
![]() Submitted July 15, 2025
Submitted July 15, 2025
![]() Epub 16 March, 2026
Epub 16 March, 2026