![]() Gabriel de S. C. Silva
Gabriel de S. C. Silva ![]() ,
, ![]() Lais Reia,
Lais Reia, ![]() Camila S. Souza,
Camila S. Souza, ![]() Eric V. Ywamoto and
Eric V. Ywamoto and ![]() Claudio Oliveira
Claudio Oliveira
PDF: Download Here | Supplementary: S1 | Cite this article
Associate Editor: ![]() Marcelo Melo
Marcelo Melo
Section Editor: ![]() William Crampton
William Crampton
Editor-in-chief: ![]() José Birindelli
José Birindelli
Abstract
Brachyglaniini é uma tribo recentemente descrita de Heptapterinae, composta por quatro gêneros: Brachyglanis, Gladioglanis, Leptorhamdia e Myoglanis. Expedições realizadas nas bacias dos rios Aripuanã e Tapajós revelaram uma espécie ainda não descrita de Brachyglaniini, caracterizada por uma faixa de cor clara e conspícua posterior à cabeça. Nossos resultados filogenéticos fornecem fortes evidências de que esta nova espécie de Brachyglaniini pertence ao gênero Leptorhamdia e está estreitamente relacionada a L. marmorata e L. aspredinoides. Além do padrão de coloração, a nova espécie pode ser distinguida de seus congêneres por possuir 19-22 raios na nadadeira anal e 47 vértebras.
Palavras-chave: Bagre, Escudo Brasileiro, Sistemática, Taxonomia integrativa.
Introduction
Heptapteridae is a group of small Neotropical catfishes commonly known as bagrinhos, mandis, and mandizinhos, with 239 valid species classified into 23 genera (Fricke et al., 2025a). Silva et al. (2021) recognized a monophyletic component of heptapterids composed of Brachyglanis Eigenmann, 1912, Gladioglanis Ferraris & Mago-Leccia, 1989, Leptorhamdia Eigenmann, 1918, and Myoglanis Eigenmann, 1912. This clade was classified as tribe Brachyglaniini, a group with 15 valid species whose diversity predominantly occurs in rivers of the Guyana Shield, such as Essequibo, Negro, Ventuari, Orinoco and Trombetas but which also includes a few species in rivers draining the Brazilian Shield, such as Madeira and Xingu. Besides being strongly supported by molecular data, Brachyglaniini is also supported by 10 morphological synapomorphies (Bockmann, 1998; Silva et al., 2021) and can be externally easily distinguished from other heptapterids by having the principal unbranched pectoral-fin ray forming an ossified, rigid spine (Faustino-Fuster et al., 2021).
After Silva et al. (2021) described the tribe Brachyglaniini, Faustino-Fuster et al. (2021) proposed a new phylogeny of Heptapteridae using a dense sampling of Brachyglaniini from the Guiana Shield in a multi-locus approach and recovered four main clades within the tribe: (1) Brachyglanis, (2) Leptorhamdia (with Myoglanis koepckei Chang, 1999, M. aspredinoides DoNascimiento & Lundberg, 2005, and Brachyglanis nocturnus Myers, 1928), (3) Myoglanis, and (4) Gladioglanis. Although these authors identified the main clades within Brachyglaniini, they did not diagnose the clades or its major subgroups from a morphological standpoint. Currently, Leptorhamdia is composed by five valid species: L. aspredinoides from the Venturari River; L. essequibensis (Eigenmann, 1912) from the Essequibo River; L. marmorata Myers, 1928, and L. nocturna from the Negro River and L. schultzi (Miranda Ribeiro, 1964) from the Xingu River (Fricke et al., 2025b). Herein, a new species of Leptorhamdia is described based on specimens collected in the tributaries of rio Arinos, and rio Aripuanã, in the Amazon basin.
Material and methods
Morphological analysis. Measurements were taken from the left side of specimens and were made point to point to the nearest 0.1 mm with digital calipers. The nomenclature of osteology follows Bockmann, Castro (2010). Measurements and abbreviations follow Lundberg, McDade (1986). The nomenclature for lateral-line canals and branches follows Bockmann, Miquelarena (2008). Morphometrics are given as percentages of standard length (SL), except for subunits of head expressed as percentages of head length (HL). Specimens were cleared and stained (c&s) according to Taylor, Van Dyke (1985). Numbers of branchiostegal rays, gill rakers, vertebrae, ribs, and position of supporting elements of dorsal and anal fins were determined in cleared and stained specimens. Vertebral counts include those from the Weberian apparatus and the compound caudal centrum was counted as one. Morphological data for L. marmorata, L. nocturna, and L. essequibensis were obtained from images of type material, available at CAS Ichthyology Primary Types Imagebase website, and from ACSImageBase hosted at The Academy of Natural Sciences. Leptorhamdia aspredinoides data were obtained from their original descriptions (Do Nascimiento, Lundberg, 2005), and data for L. schultzi were obtained from ANSP image collections. Institutional abbreviations follow Sabaj (2020). Specimens were deposited at the Departamento de Ciências Biológicas da UNESP, São José do Rio Preto (DZSJRP), Laboratório de Biologia e Genética de Peixes, Universidade Estadual Paulista, Botucatu (LBP), and Museu de Zoologia, Universidade de São Paulo, São Paulo (MZUSP).
Molecular analysis. The extracted DNA of one sample of the new species of Leptorhamdia, which was used for the molecular analysis: LBP 20829 (tissue 81351; Genbank number: PX974334 (COI); PX984954 (Cytb); PX984955 (Rag2)). DNA extraction followed Ivanova et al. (2006). Partial sequences of the mitochondrial genes cytochrome oxidase c subunit I (COI) and cytochrome b (Cytb), as well as the nuclear gene recombination activating gene 2 (Rag2) were amplified by polymerase chain reaction (PCR) using primers FishF6/R7 (Jennings et al., 2019), L14841/H16460 (Oliveira et al., 2011) and 164F/176R (Oliveira et al., 2011), respectively. Reactions were carried out in a 12.5 μL reaction volume containing 1.25 μL of 10× PCR buffer, 0.40 μL MgCl2 (50 mM), 0.30 μL dNTPs (2 mM), 0.25 μL of each primer (5 μM), 0.20 μL of PHT Taq DNA polymerase (Phoneutria), 2 μL DNA template (200 ng), and 7.85 μL of ddH2O. The PCR consisted of initial denaturation (5 min at 95°C) followed by 30 cycles with a chain denaturation (1 min at 95°C), primer hybridization (45 sec at 54°C), nucleotide extension (1 min at 72°C), and final extension (10 min at 72°C). All PCR products were checked using 1% agarose gel and purified with ExoSap-IT (USB Corporation) following the manufacturer’s instructions. The purified PCR products were sequenced using the Big DyeTM Terminator v. 3.1 Cycle Sequencing Ready Reaction Kit (Applied Biosystems, Austins, USA), purified through ethanol precipitation and loaded onto an ABI 3500 Genetic Analyzer (Applied Biosystems), in the Instituto de Biotecnologia (IBTEC), Universidade Estadual Paulista Júlio de Mesquita Filho, Botucatu, Brazil.
The sequences of two genes obtained in this study were assembled using the software GENEIOUS PRIME 2024.0.5 (Kearse et al., 2012). These sequences, along with available sequences from the phylogenetic study by Faustino-Fuster et al. (2021), which included three mitochondrial genes (COI, Cyt b, ND2) and two nuclear genes (RAG2 and Glyt) were aligned using muscle (Edgar, 2004) under default parameters to create a new dataset matrix comprising 40 specimens. Nucleotide variation and substitution patterns were examined using IQ-TREE v. 2.3.6(Minh et al., 2020). To evaluate the occurrence of substitution of saturation in our data, we estimated the index of substitution saturation (Iss) using the method described by Xia et al. (2003) and Xia, Lemey (2009) using DAMBE v. 7.2.1 (Xia, 2018).
The concatenated matrix was partitioned by gene (Chernomor et al., 2016) and the best-fit model of nucleotide evolution was selected with ModelFinder (Kalyaanamoorthy et al., 2017) based on the Bayesian Information Criterion (BIC) and weighted BIC (w-BIC) scores. Phylogenetic relationships were inferred through the Maximum Likelihood (ML) analysis with 1000 ultrafast bootstrap replicates (Hoang et al., 2018). All analyses were conducted using IQ-TREE v. 2.3.6(Minh et al., 2020). Pimelodus ornatus Kner, 1858 was used to root the phylogeny. Additionally, samples of Rhamdiini and Heptapterini were included in the analysis as additional related groups (Fig. S1). The final tree was visualized with the software FIGTREE v. 1.4.4 (Rambaut, 2014).
Results
Leptorhamdia kamilai,new species
urn:lsid:zoobank.org:act:4FE789F9-D7EF-4B42-A46C-E6FB75B62EA3
(Figs. 1–2; Tab. 1)
Myoglanis sp. —Casatti et al., 2020:8, tab. 2 (inventory).
Myoglanis n. sp.1 Tapajós. —Silva et al., 2021:5, fig. 1 (phylogenomic of Heptapteridae).
Holotype. DZSJRP 16293, 92.4 mm SL, Brazil, Mato Grosso State, Campos de Júlio Municipality, rio Juruena, PCH Segredo, 13°13’27.6”S 59°01’34.4”W, Jun 2012, R. J. Ilario.
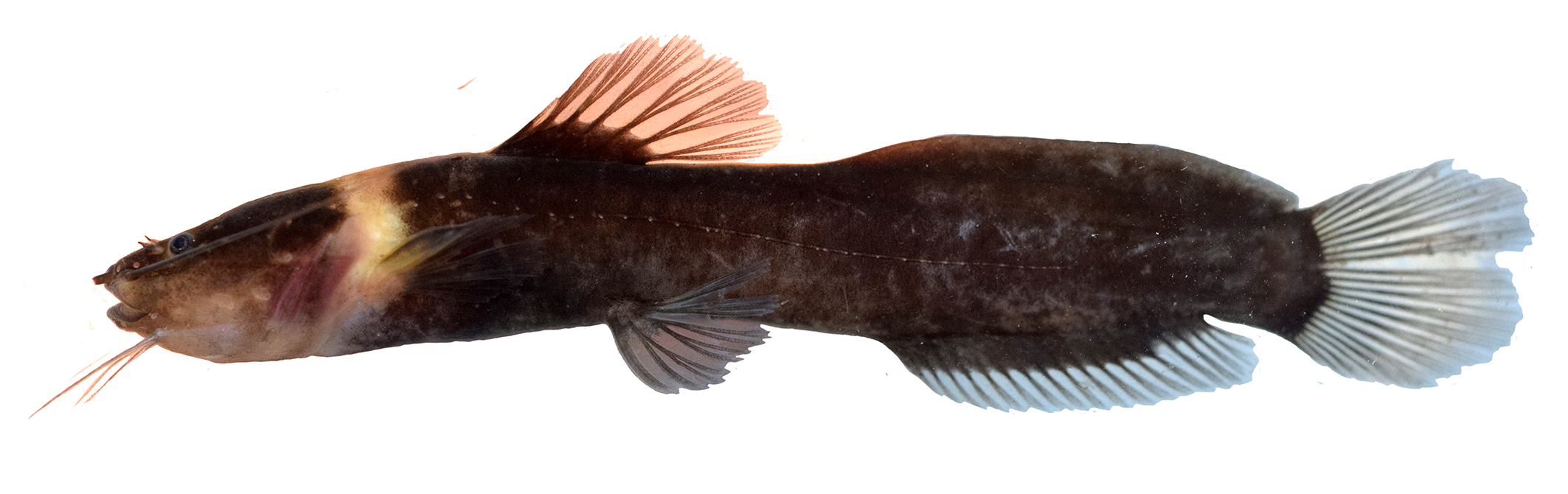
FIGURE 1| Left column: Leptorhamdia kamilai, holotype, DZSJRP 16293, 92.4 mm SL. Right column: paratypes, LBP 30639, from top to bottom: 66.3 mm SL; 47.8 mm SL; 31.4 mm SL.
TABLE 1 | Morphometric data for holotype (H) and paratypes of Leptorhamdia kamilai. N = number of specimens; SD = Standard deviation.
| H | Tapajós (N = 14) | Aripuanã (N = 9) | ||||
Range | Mean | SD | Range | Mean | SD | ||
Standard length (mm) | 92.4 | 31.4-92.4 | – | – | 22.5-73.9 |
| – |
Percentages of standard length | |||||||
Predorsal length | 36.8 | 34.1-37.3 | 35.8 | 1.0 | 32.2-36.5 | 34.9 | 1.4 |
Dorsal-fin base | 13.5 | 11.5-15.5 | 13.4 | 1.1 | 12.5-15.5 | 14.1 | 1.0 |
Dorsal fin to adipose fin | 16.1 | 11.8-16.1 | 13.9 | 1.3 | 10.6-14.7 | 12.7 | 1.2 |
Preadipose length | 62.1 | 57.4-64.5 | 61.5 | 1.8 | 58.3-61.3 | 59.9 | 1.2 |
Adipose-fin length | 41.7 | 39.0-44.5 | 41.6 | 1.3 | 38.9-44.9 | 41.7 | 2.0 |
Prepelvic length | 42.5 | 40.4-45.5 | 42.2 | 1.4 | 40.6-43.7 | 41.8 | 0.9 |
Preanal length | 64.6 | 63.7-69.8 | 65.4 | 1.6 | 64.2-65.6 | 65.0 | 0.6 |
Anal-fin base | 28.2 | 26.1-29.6 | 28.0 | 1.1 | 25.5-29.4 | 28.1 | 1.3 |
Caudal peduncle length | 10.8 | 7.5-10.8 | 9.3 | 1.0 | 8.6-10.5 | 9.3 | 0.6 |
Caudal peduncle depth | 8.7 | 7.2-11.4 | 9.0 | 1.2 | 7.9-9.5 | 8.7 | 0.5 |
Head length | 23.2 | 22.3-26.2 | 24.1 | 1.2 | 21.7-24.3 | 23.0 | 0.9 |
Head width | 20.1 | 16.2-20.4 | 18.2 | 1.0 | 16.3-17.6 | 17.0 | 0.5 |
Head depth | 14.5 | 11.0-14,9 | 12.7 | 1.1 | 11.0-14.7 | 12.5 | 1.2 |
Maxillary-barbel length | 34.7 | 31.4-38.7 | 34.5 | 2.0 | 36.2-40.9 | 37.7 | 1.4 |
Outer mental-barbel length | 16.2 | 14.6-20.4 | 17.5 | 1.4 | 14.1-19.0 | 16.1 | 1.4 |
Inner mental-barbel length | 8.4 | 8.4-12.9 | 9.8 | 1.2 | 7.5-11.0 | 9.1 | 1.4 |
Percentages of head length | |||||||
Snout length | 32.9 | 28.3-32.9 | 31.1 | 1.8 | 30.4-34.0 | 32.9 | 1.2 |
Fleshy interorbital | 29.1 | 24.0-29.5 | 26.8 | 1.7 | 21.8-27.6 | 25.5 | 2.0 |
Width between anterior nares | 16.1 | 16.1-25.6 | 20.1 | 2.4 | 15.3-21.1 | 19.0 | 2.1 |
Width between posterior nares | 20.2 | 18.5-23.4 | 21.8 | 1.4 | 19.4-24.9 | 22.3 | 2.1 |
Paratypes. All from Brazil, Mato Grosso State. Rio Tapajós basin: DZSJRP 25393, 1, 71.5 mm SL, collected with holotype. DZSJRP 16302, 2, 53.8-91.0 mm SL, Campos de Júlio Municipality, rio Juruena, PCH Ilha Comprida, Jun 2012, R. J. Ilario. DZSJRP 21986, 2, 56.6-84.6 mm SL, Juina, rio Juruena, 11º31’49”S 58º49’33”W, 30 May 2018, G. Brejão. LBP 30639, 2, 31.4-66.3 mm SL, Nova Mutum Municipality, rio dos Patos, a tributary of rio Arinos, 13º48’4.79”S 56º1’37.4”W, 22 Aug 2021, C. S. Souza, E. V. Ywamoto, G. S. C. Silva, & L. Reia. LBP 20829, 4, 40.1-48.3 mm SL (1 c&s, 48.3 mm SL), Nova Mutum Municipality, rio dos Patos, a tributary of rio Arinos, 13º48’4.79”S 56º1’37.4”W, 29 Oct 2015, C. Araya, G. J. Costa-Silva, N. T. B. Mateussi, B. F. Melo & G. S. C. Silva. MZUSP 131552, 2, 32.9-47.8 mm SL, Nova Mutum Municipality, rio dos Patos, a tributary of rio Arinos, 13º48’4.79”S 56º1’37.4”W, 22 Aug 2021, C. S. Souza, E. V. Ywamoto, G. S. C. Silva, & L. Reia. Rio Aripuanã basin: DZSJRP 21470, 1, 30.8 mm SL, Aripuanã Municipality, 10°11’47.0”S 59°29’19.0”W, 27 Jul 2017. DZSJRP 21559, 1, 72.4 mm SL, Aripuanã Municipality, 10°11’26.0”S 59°24’34.0”W, 28 Jul 2017, G. Brejão. DZSJRP 21641, 1, 49.0 mm SL. Aripuanã Municipality, 10°08’15.0”S 59°24’57.0”W, 31 Jul 2017, G. Brejão. DZSJRP 21709, 2, 49.8-66.6 mm SL, Aripuanã Municipality, 10°15’38.0”S 59°17’06.0”W, 1 Aug, 2017, G. Brejão. DZSJRP 21755, 1, 33.2 mm SL, Aripuanã Municipality, 10°20’58.0”S 59°21’43.0”W, G. Brejão. DZSJRP 21778, 7, 24.973.9 mm SL (1 c&s 60.3 mm SL), Aripuanã Municipality, 10°21’52.0”S 59°21’38.0”W, 2, Aug 2017. G. Brejão. DZSJRP 21791, 1, 36.0 mm SL, Aripuanã Municipality, 10°25’59.0”S 59°21’41.0”W, 2 Aug 2017, G. Brejão. DZSJRP 21819, 2, 22.824.5 mm SL, Aripuanã Municipality, 10°46’49.0”S 59°13’37.0”W, 3 Aug 2017, G. Brejão. DZSJRP 21938, 1, 51.5 mm SL, Juína Municipality, 11°32’07.0”S 59°11’39.0”W, 9 Aug 2017, G. Brejão.
Diagnosis. Leptorhamdia kamilai can be readily distinguished among its congeners by having a conspicuous light collar-band, yellowish in live specimens, posterior to the head (vs. coloration uniformly throughout body and head, lacking any distinctive bar). Additionally, L. kamilai differs from its congeners by having 19-22 anal-fin rays (vs. 17 in L. essequibensis; 15 in L. marmorata; 32-35 in L. aspredinoides; 12 in L. nocturna, 15-16 in L. schultzi) and by having 46-47 vertebrae (vs. 45 in L. essequibensis; 43 in L.marmorata; 38 in L. nocturna; 56-57 vertebrae in L. aspredinoides and 43 vertebrae in L. schultzi).
Description. Dorsal profile from snout tip to eyes convex, slightly concave in orbital region, convex from orbital region to collar-band, then straight to dorsal-fin origin, straight (descending) along dorsal-fin base, straight (ascending) from end of dorsal-fin base to origin of adipose fin, slightly concave along adipose-fin base and finishing slightly straight along caudal peduncle. Ventral profile straight from snout to posterior margin of branchiostegal membrane, then straight (ascending) to pelvic-fin origin, finishing gently concave to end of pelvic fin to caudal peduncle.
Head trapezoid in dorsal view. Adductor mandibulae muscles prominent on cheeks and extending dorsally over skull roof posterior to eyes. Eyes rounded and small, dorsally positioned, close to snout tip. Skin covering eye thin. Mouth subterminal, with upper jaw slightly longer than lower. Premaxilla and dentary with four to five irregular rows of conical teeth. Maxillary barbel surpassing vertical through dorsal-fin origin. Outer mental longer than inner mental barbel, reaching posterior margin of branchiostegal membrane. Branchiostegal membranes supported by seven rays and joined to isthmus only at anterior point. Five gill rakers along anterior border of first ceratobranchial. Total vertebrae 46-47 with 15 precaudal (including eight or nine supporting pleural ribs) and 31-32 caudal.
Laterosensory canal of the head with simple tubes ending in single pores. Supraorbital sensory canal usually with four branches and pores: s1, s2, s3, and s4. Supraorbital pore 1 medially adjacent to anterior nares. Supraorbital s2 at midway between anterior and posterior nares, s3 inside posterior nares, at the notch of the cutaneous membrane. Pores s4 paired, close together near midline, posterior at level of orbit. Infraorbital laterosensory canal with five branches and pores: i1, i2, i3, i4, and i5. Infraorbital pore i1 adjacent to anterior naris, between anterior naris and maxillary barbel; i2 laterally positioned at the midway between anterior and posterior naris. Pore i3 at vertical through anterior margin of orbit; i4 at vertical through posterior margin of orbit. Pore i5 posterior to eye. Preoperculomandibular laterosensory canal with 11 branches and pores: pm1 in the medial portion of dentary; pm2, pm3, and pm4 aligned anteriorly to inner and outer mental barbel; pm5 dorsal to outer mental barbel base; pm6 and pm7 at vertical through the anterior and posterior margin of orbit respectively. Four pores in preopercle region: pm8, pm9, pm10 and pm11.
Pectoral fin with seven branched rays preceded by a short and strong spine, with retrorse dentitions along both margins; serrae of posterior margin of spine more developed than the anterior margin. Pelvic fin i,5; first ray shorter than the posterior rays. Dorsal-fin i,6; first basal radial inserted on sixth or seventh vertebra and last basal radial anterior to neural spine of twelfth vertebra. Adipose fin elongate (40.0-44.5% of SL) with it posterior margin united to dorsal margin of caudal fin. Anal fin elongated with 19 to 22 rays, first three or four rays simple. Anal fin supported by 20-22 basal radials (2 c&s). Ventral caudal plate formed by parhypural free from hypurals 1 and 2. Dorsal caudal plate formed by uroneural, hypurals 3 and 4 fused and 5 free. Caudal fin lanceolate with i,7,8,i principal rays. Dorsal and ventral caudal-fin lobes with 5 and 9 procurrent rays, respectively.
Coloration in alcohol. Overall coloration uniform dark, mostly unpigmented on ventral half of head and trunk (Fig. 2). A conspicuous band of pale coloration, yellowish in live specimens, extending dorsally from slight posterior to supraoccipital, along the sides, to the ventral midline on branchiostegal membrane. In small specimens (< 30 mm SL) light collar band separated by a narrow-pigmented stripe at its midline portion. Adipose-, dorsal-, and anal-fin bases dark. Fin rays and interradial membranes hyaline.
FIGURE 2| Live specimens of Leptorhamdia kamilai, paratype, LBP 30639, 66.3 mm SL. Photographed by Eric V. Ywamoto.
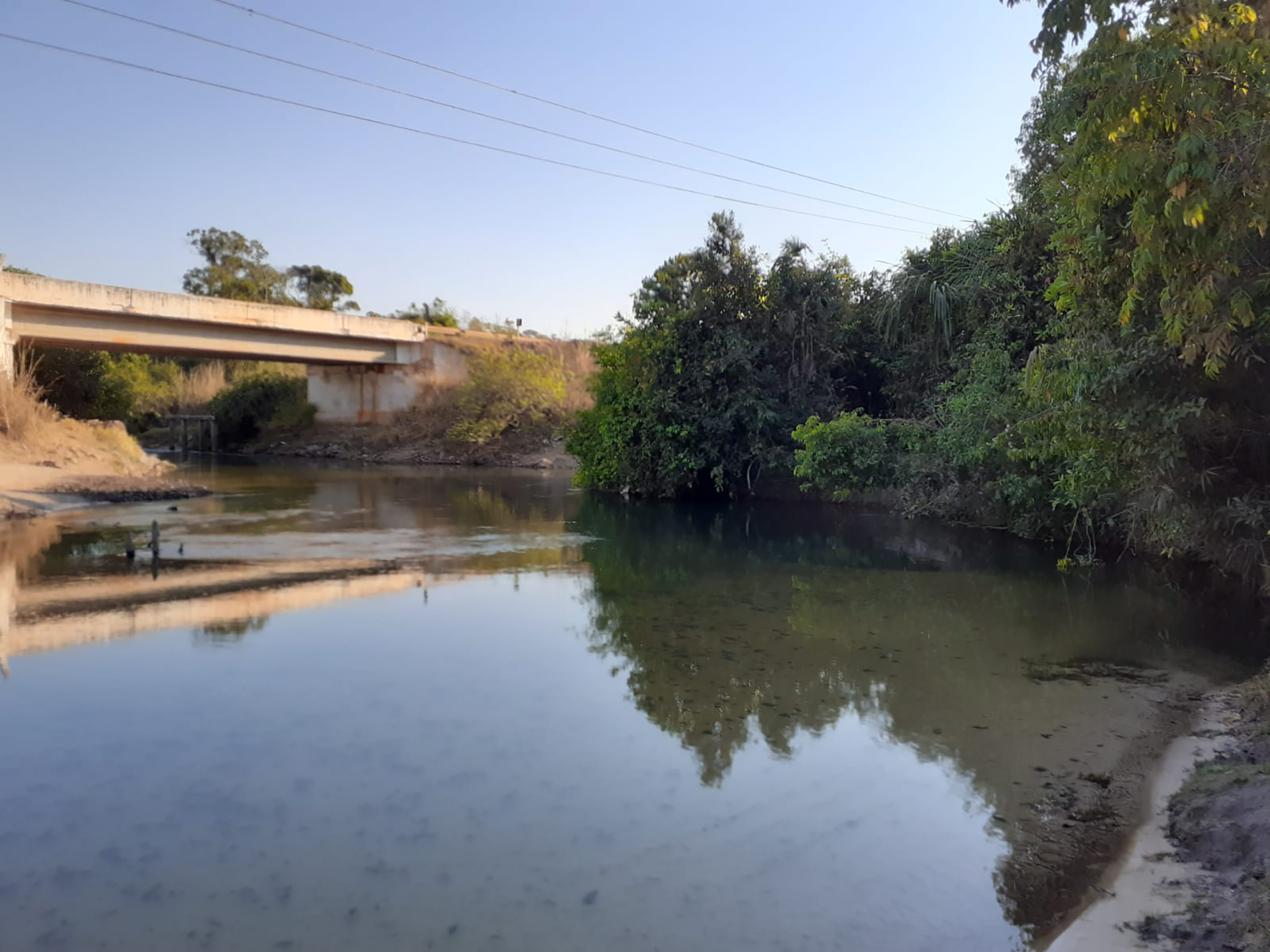
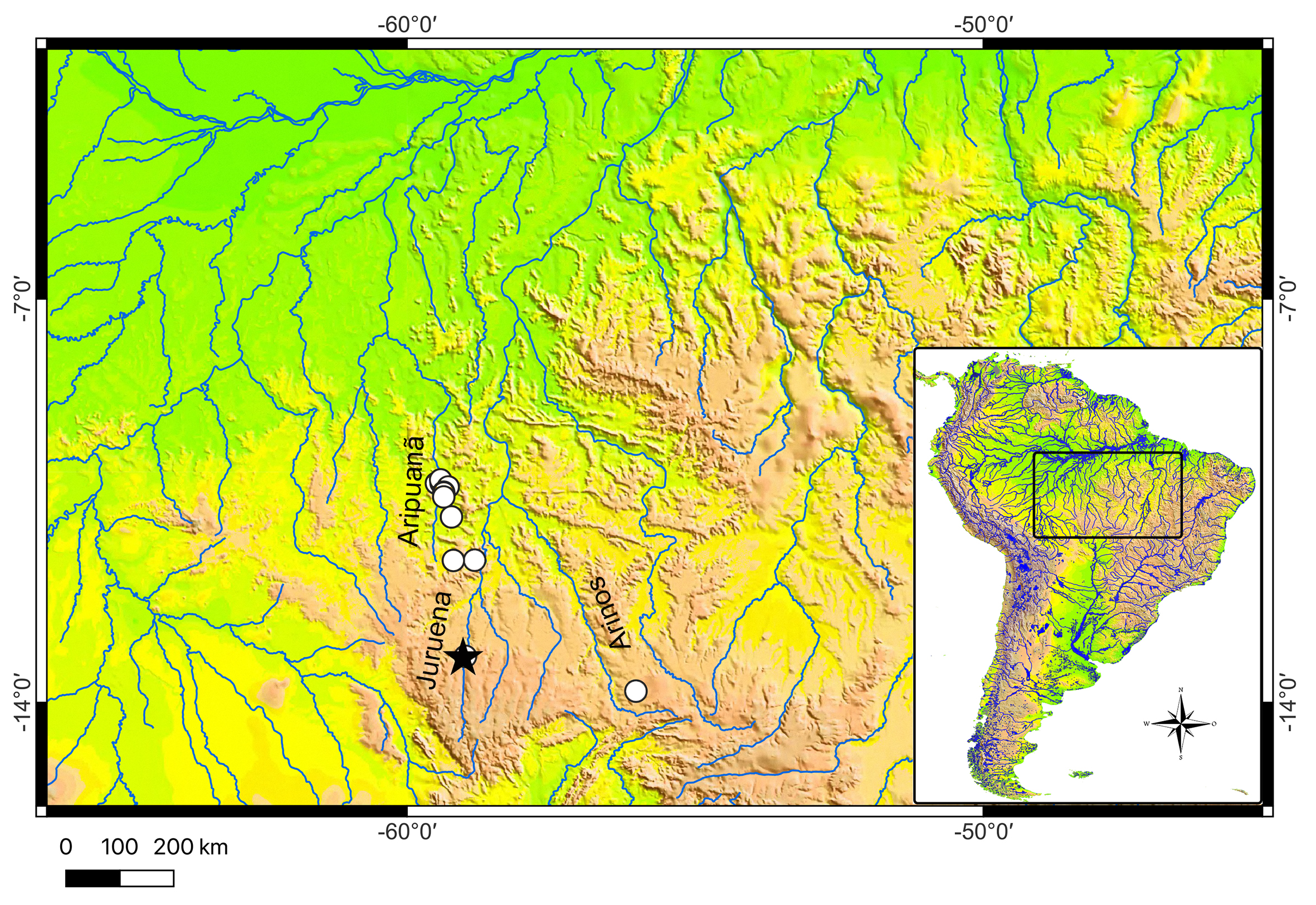
Geographicaldistribution. The new species Leptorhamdia kamilai is known from tributaries of the upper rio Arinos and Juruena in the rio Tapajós basin and from tributaries of the rio Aripuanã in the rio Madeira basin, Mato Grosso State, central Brazil (Fig. 3). The new species was collected about 1.5 m deep, hidden beneath a leaf layer and muddy bottom (Fig. 4).
FIGURE 3| Geographic distribution of Leptorhamdia kamilai in Brazilian Shield. Black star: holotype; white circles: paratypes.
FIGURE 4| Habitat of Leptorhamdia kamilai,rio dos Patos, tributary of the upper Arinos River in Tapajós River basin.
Conservation status. Leptorhamdia kamilai was collected at several sampling sites in the Upper Tapajós and Aripuanã river basins, with no imminent threats to the species detected. Therefore, we recommend that L. kamilai be categorized as Least Concern (LC) under IUCN’s criteria (IUCN, 2024).
Etymology. The specific name, kamilai, honors our dear friend Dr. Kamila Mayumi Duarte Kuabara, an entomologist and lab manager of the Ornithology Department at The Academy of Natural Sciences of Drexel University. A noun in a genitive case.
Phylogeny. The concatenation resulted in a matrixof Heptapteridae with five loci and 4150 pb. No saturation was observed in the matrix, since the values of Iss (Index of Substitution Saturation) were lower than Iss.c (Critical Index of Substitution Saturation). According to BIC the best model fitted each gene were TPM2u+F+I+G4 (LogL -6087.950) for COI; TVM+F+I+R3 (LogL -9138.688) for Cyt b; TPM3+G4 (LogL -3158.490) for Glyt; TPM2+G4 (LogL -3601.665) for RAG2; TPM2u+F+I+R3 (LogL -8437.546) for ND2. The ML tree (LogL -30419.782) and Consensus tree (LogL -30419.809) returned identical topology and values of bootstrap. Our results recovered the same major clades identified in a recent phylogenomic study of Heptapteridae (Faustino-Fuster et al., 2021; Silva et al., 2021). Additionally, our results found Leptorhamdia kamilai nested in the tribe Brachyglaniini and sister group to L. marmorata+L. aspredinoides, two species from Guyana Shield (Figs. 5, S1).
FIGURE 5| Maximum likelihood tree of Heptapteridae showing the relationships among Brachyglaniini species. Black nodes indicate support between 90% and 100% bootstrap values; gray nodes indicate nodal support between 80% and 89%.
Discussion
Our phylogenetic analysis revealed that the new species of Brachyglaniini described here is part of the clade composed of Leptorhamdia species, sister to L. marmorata+L. aspredinoides, justifying its allocation within this genus. Additionally, L. kamilai shares one of the three morphological characters proposed by Bockman, Slobodian (2018) to define Leptorhamdia: a premaxillary tooth plate formed as a compact triangular or rectangular structure, bearing a distinct posterolateral projection (vs. forming an arch, lacking a posterolateral projection in Myoglanis species). The other two diagnostic characters proposed by Bockman, Slobodian (2018) to distinguish Leptorhamdia from Myoglanis were not found in the new species: adipose fin very low, with the same depth along its entire length and adipose fin disconnected from caudal fin, with a distinct rounded posterior margin forming a lobe. These latter characters are absent in the new species, suggesting that they may not be suitable for defining the genus.
The new species described here, L. kamilai, is known from two distinct and adjacent drainages: the Juruena and Aripuanã river basins. Although representatives of this species occur in allopatry, specimens from both basins exhibit no morphological differences. Dagosta, de Pinna (2019) pointed out that the Aripuanã and Juruena Rivers share several ichthyofaunal elements, such as Hemigrammus silimoni Britski & Lima, 2008, Moenkhausia levidorsa Benine, 2002, and the genus Utiaritichthys Miranda Ribeiro, 1937, and argued that recent geomorphological events have caused biotic dispersal in the headwater of both rivers. Although the specimens of L. kamilai from the Juruena and Aripuanã are highly similar in morphological terms, genetic analyses of samples from the Aripuanã River are still pending, and it is not possible to assess potential genetic disparity between them.
During two expeditions to the rio dos Patos (the type locality), we were only able to collect eight specimens of L. kamilai. The difficulty in capturing this species is because these fish live buried in the leaf litter at the bottom of the river, at a depth of about 1.5 m. To collect specimens of L. kamilai, it was necessary to dive to the bottom of the riverbed and remove the layer of mud, where the species is associated with the leaf litter. Among the members of Brachyglaniini, only L. kamilai and Myoglanis koepckei have been reported to inhabit the leaf litter banks (Lima et al., 2005); all other members of the tribe are found in rapids, associated with rock-bottomed, fast-flowing rivers (Bockman, Slobodian, 2018).
Among members of Brachyglaniini, only Leptorhamdia kamilai exhibits a conspicuous light collar-band posterior to the head, which is regarded as an autapomorphy for this species. Similar pigmentation patterns are also observed in other heptapterid taxa, including species of Pariolius (P. armillatus and P. maldonadoi) and Chasmocranus stricto sensu (C. brevior and C. longior). According to recent phylogenetic hypotheses for Heptapteridae (Silva et al., 2021), L. kamilai (there named as Myoglanis n. sp. 1 Tapajós), Pariolius and Chasmocranus are phylogenetically distant, indicating that the light collar-band represents a homoplastic condition that evolved independently multiple times in the family. However, the selective pressures driving the convergent evolution of such pigmentation patterns within the group remain to be investigated.
Comparative material examined. Brazil. Leptorhamdia shultzi: ANSP 197846, 1, 88.1 mm SL; ANSP 200078, 1, 90.0 mm SL. Peru. Myoglanis koepckei: ANSP 167657, 6, 33.3-55.3 mm SL. Venezuela. Leptorhamdia sp.: ANSP 192588, 1, 65.6 mm SL.
Acknowledgments
We are grateful to Bruno Melo, Guilherme Costa-Silva, Cristian Araya and Nadayca Mateussi for their help during fieldwork, and specially thank Gabriel Brejão (UNESP), for collecting most of the type material. Thanks to Mark Sabaj (ANSP) and Francisco Langeani (UNESP) for the curatorial assistance and Marcelo Melo (USP) for the contribution with comments and revision of the manuscript. We also thank to Jefferson Crispim (UNESP) for the x-rays of L. schultzi.
References
Bockmann FA. Análise filogenética da família Heptapteridae (Teleostei, Ostariophysi, Siluriformes) e redefinição de seus gêneros. [PhD Thesis]. São Paulo: Universidade de São Paulo; 1998. Available from: https://repositorio.usp.br/item/001000413
Bockmann FA, Castro R. The blind catfish from the caves of Chapada Diamantina, Bahia, Brazil (Siluriformes: Heptapteridae): description, anatomy, phylogenetic relationships, natural history, and biogeography. Neotrop Ichthyol. 2010; 8(4):673–706. https://doi.org/10.1590/S1679-62252010000400001
Bockmann FA, Miquelarena AM. Anatomy and phylogenetic relationships of a new catfish species from northeastern Argentina with comments on the phylogenetic relationships of the genus Rhamdella Eigenmann and Eigenmann 1888 (Siluriformes, Heptapteridae). Zootaxa. 2008; 1780(1):1–54. https://doi.org/10.11646/zootaxa.1780.1.1
Bockmann FA, Slobodian V. Family Heptapteridae – Three-barbeled catfishes. In: Sleen PV, Albert JS, editors. Field guide to the fishes of the Amazon, Orinoco and Guianas. New Jersey: Princeton University Press; 2018. p.233–52.
Casatti L, Brejão GL, Carvalho FR, Silva HP, Pérez-Mayorga MA, Manzotti AR et al. Stream fish from recently deforested basins in the Meridional Amazon, Mato Grosso, Brazil. Biota Neotrop. 2020; 20(1):e20190744. https://doi.org/10.1590/1676-0611-BN-2019-0744
Chernomor O, von Haeseler A, Minh BQ. Terrace aware data structure for phylogenomic inference from supermatrices. Syst Biol. 2016; 65:997–1008. https://doi.org/10.1093/sysbio/syw037
Dagosta FCP, de Pinna M. The fishes of the Amazon: distribution and biogeographical patterns, with a comprehensive list of species. Bull Am Mus Nat Hist. 2019; 2019(431):1–163. https://doi.org/10.1206/0003-0090.431.1.1
Donascimiento C, Lundberg JG. Myoglanis aspredinoides (Siluriformes: Heptapteridae), a new catfish from the Río Ventuari, Venezuela. Zootaxa. 2005; 1009(1):37–49. https://doi.org/10.11646/zootaxa.1009.1.4
Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004; 32:1792–97. https://doi.org/10.1093/nar/gkh340
Faustino-Fuster DR, Mesa-Vargas V, Lovejoy NR, Lujan NK. Multi-locus phylogeny with dense Guiana Shield sampling supports new suprageneric classification of the neotropical three-barbeled catfishes (Siluriformes: Heptapteridae). Mol Phylogenet Evol. 2021; 162:107186. https://doi.org/10.1016/j.ympev.2021.107186
Fricke, R, Eschmeyer, WN, Fong JD. Eschmeyer’s catalog of fishes: genera, species by family/subfamily. [Internet]. San Francisco:California Academy of Science; 2025a. Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/SpeciesByFamily.asp
Fricke R, Eschmeyer WN, Van der Laan R. Eschmeyer’s catalog of fishes: genera, species, references. [Internet]. San Francisco: California Academy of Science; 2025b. Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp
Hoang DT, Chernomor O, von Haeseler A, Minh BQ, Vinh LS. UFBoot2: improving the ultrafast bootstrap approximation. Mol Biol Evol. 2018; 35(2):518–22. https://doi.org/10.1093/molbev/msx281
International Union for Conservation of Nature (IUCN). Standards and petitions committee. Guidelines for using the IUCN Red List categories and criteria. Version 16 [Internet]. Gland; 2024. Available from: http://www.iucnredlist.org/documents/RedListGuidelines.pdf
Ivanova NV, Dewaard JR, Hebert PDN. An inexpensive, automation-friendly protocol for recovering high-quality DNA. Mol Ecol Resour. 2006; 6(4):998–1002. https://doi.org/10.1111/j.1471-8286.2006.01428.x
Jennings WB, Ruschi PA, Ferraro G, Quijada CC, Silva-Malanski ACG, Prosdocimi F et al. Barcoding the Neotropical freshwater fish fauna using a new pair of universal COI primers with a discussion of primer dimers and M13 primer tails. Genome. 2019; 62(2):77–83. https://doi.org/10.1139/gen-2018-0145
Kalyaanamoorthy S, Minh BQ, Wong TKF, von Haeseler A, Jermiin LS. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods. 2017; 14:587–89. https://doi.org/10.1038/nmeth.4285
Kearse M, Moir R, Wilson A, Stone-Havas S, Cheung M, Sturrock S et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012; 28(12):1647–49. https://doi.org/10.1093/bioinformatics/bts199
Lima FCT, Ramos L, Barreto T, Cabalzar A, Tenório G, Barbosa A et al. Peixes do alto Tiquié: ictiologia e conhecimentos dos Tukano e Tuyuka. In: Cabalzar A, Lima FCT, Lopes MC, editors. Peixes e gente no alto rio Tiquié: conhecimento Tukano e Tuyuka, ictiologia, etnologia. São Paulo: Instituto Socioambiental; 2005. p.111–282.
Lundberg JG, McDade LA. On the South American catfish Brachyrhamdia imitator Myers (Siluriformes, Pimelodidae), with phylogenetic evidence for a large intrafamilial lineage. Acad Nat Sci Phila. 1986; 463:1–24.
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, von Haeseler A et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol. 2020; 37(5):1530–34. https://doi.org/10.1093/molbev/msaa015
Oliveira C, Avelino GS, Abe KT, Mariguela TC, Benine RC, Ortí G et al. Phylogenetic relationships within the speciose family Characidae (Teleostei: Ostariophysi: Characiformes) based on multilocus analysis and extensive ingroup sampling. BMC Evol Biol. 2011; 11:275. https://doi.org/10.1186/1471-2148-11-275
Rambaut A, Suchard MA, Xie D, Drummond AJ. Tracer v1.6. 2014. Available from: http://beast.bio.ed.ac.uk/Tracer
Sabaj MH. Codes for natural history collections in ichthyology and herpetology. Copeia. 2020; 108(3):593-669. https://doi.org/10.1643/ASIHCODONS2020
Silva GSC, Roxo FF, Melo BF, Ochoa LE, Bockmann FA, Sabaj MH et al. Evolutionary history of Heptapteridae catfishes using ultraconserved elements (Teleostei, Siluriformes). Zool Scr. 2021; 50(5):543–54. https://doi.org/10.1111/zsc.12493
Taylor WR, Van Dyke GC. Revised procedures for staining and clearing small fishes and other vertebrates for bone and cartilage study. Cybium. 1985; 9(2):107–19. Available from: https://sfi-cybium.fr/en/node/2423
Xia X, Xie Z, Salemi M, Chen L, Wang Y. An index of substitution saturation and its application. Mol Phylogenet Evol. 2003; 26(1):1–07. https://doi.org/10.1016/S1055-7903(02)00326-3
Xia X, Lemey P. Assessing substitution saturation with DAMBE. In: Lemey P, Salemi M, Vandamme AM, editors. The phylogenetic handbook: a practical approach to phylogenetic analysis and hypothesis testing. Cambridge: Cambridge University Press; 2009. p.615–30.
Xia X. DAMBE7: new and improved tools for data analysis in molecular biology and evolution. Mol Biol Evol. 2018; 35(6):1550–52. https://doi.org/10.1093/molbev/msy073
Authors
![]() Gabriel de S. C. Silva
Gabriel de S. C. Silva ![]() ,
, ![]() Lais Reia,
Lais Reia, ![]() Camila S. Souza,
Camila S. Souza, ![]() Eric V. Ywamoto and
Eric V. Ywamoto and ![]() Claudio Oliveira
Claudio Oliveira
[1] Departamento de Biologia Estrutural e Funcional, Instituto de Biociências – Câmpus de Botucatu, Universidade Estadual Paulista, Rua Prof. Dr. Antonio C. W. Zanin, 250, 18618-689, Botucatu, SP, Brazil. (GSCS) gabriel_biota@hotmail.com (corresponding author), (LR) laisreia@gmail.com, (EVY) ericywamoto@gmail.com, (CO) claudio.oliveira@unesp.br.
Authors’ Contribution 

Gabriel de S. C. Silva: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Resources, Writing-original draft.
Lais Reia: Data curation, Formal analysis, Methodology, Software, Writing-review and editing.
Camila S. Souza: Data curation, Funding acquisition, Methodology, Writing-review and editing.
Eric V. Ywamoto: Data curation, Methodology, Writing-review and editing.
Claudio Oliveira: Data curation, Funding acquisition, Project administration, Supervision, Writing-review and editing.
Ethical Statement
Not applicable.
Competing Interests
The author declares no competing interests.
Data availability statement
All molecular markers generated and analyzed in this study are deposited in GenBank. Accession numbers are provided in the manuscript.
AI statement
The authors declare that AI tools were used exclusively to perform minor English language corrections, without any influence on the scientific content, data analysis, or interpretation of the results.
Funding
This research was supported by FAPESP process 2021/12979–8 and 2022/13025–0 (GSCS), 2020/13433–6 (CO), 17/06551–0 (CSS), 2023/09902–9 (LR), CNPq process #151174/2023–7 (LR), #306054/2006–0 (CO) and 441128/2020–3 (CO), and Pro-Reitoria de Pesquisa da Universidade Estadual Paulista Júlio de Mesquita Filho (Prope-UNESP).
Supplementary Material
Supplementary material S1
How to cite this article
Silva GSC, Reia L, Souza CS, Ywamoto EV, Oliveira C. Phylogenetic placement and description of a new species of Leptorhamdia (Siluriformes: Heptapteridae) from the Amazon basin. Neotrop Ichthyol. 2026; 24(1):e250037. https://doi.org/10.1590/1982-0224-2025-0037
Copyright
This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Distributed under
Creative Commons CC-BY 4.0

© 2025 The Authors.
Diversity and Distributions Published by SBI
![]() Accepted January 9, 2026
Accepted January 9, 2026
![]() Submitted March 6, 2025
Submitted March 6, 2025
![]() Epub April 27, 2026
Epub April 27, 2026