![]() Fábio Hiroshi Takagui1,
Fábio Hiroshi Takagui1, ![]() Patrik Viana1,
Patrik Viana1, ![]() Chrystian Aparecido Grillo Haerter1,
Chrystian Aparecido Grillo Haerter1, ![]() Jamille de Araújo Bitencourt2
Jamille de Araújo Bitencourt2 ![]() ,
, ![]() José Luís Olivan Birindelli3,
José Luís Olivan Birindelli3, ![]() Roberto Laridondo Lui4,
Roberto Laridondo Lui4, ![]() Vladimir Pavan Margarido4,
Vladimir Pavan Margarido4, ![]() Fernanda Simões Almeida5,
Fernanda Simões Almeida5, ![]() Lucia Giuliano-Caetano6,
Lucia Giuliano-Caetano6, ![]() Eliana Feldberg1 and
Eliana Feldberg1 and ![]() Jorge Ivan Rebelo Porto1
Jorge Ivan Rebelo Porto1
PDF: Download Here | Supplementary: S1 S2 S3 | Cite this article
Associate Editor: ![]() Marcelo Cioffi
Marcelo Cioffi
Editor-in-chief: ![]() Carla Pavanelli
Carla Pavanelli
Abstract
Astrodoras é um gênero monotípico de bagres que habita as principais drenagens do sistema do rio Amazonas, atualmente representado por uma única espécie reconhecida, A. asterifrons. No entanto, evidências recentes sugerem que esse gênero pode não ser monotípico e pode incluir pelo menos duas espécies não descritas. Neste estudo, integramos abordagens moleculares e citogenéticas para investigar a diversidade genética dentro de A. asterifrons. Doze indivíduos foram coletados em duas localidades da bacia do rio Negro: (a) rio Negro, próximo ao arquipélago fluvial de Anavilhanas, e (b) rio Negro, nos bancos de areia de Barcelos. Nossas análises revelaram três cariomorfos distintos: cariomorfo A e B (coexistindo próximos a Anavilhanas) e cariomorfo C (restrito a Barcelos, aproximadamente 500 km rio acima). As análises morfológicas confirmaram que os cariomorfos A e B correspondem a A. asterifrons sensu stricto, enquanto o cariomorfo C se enquadra no diagnóstico de Astrodoras sp. “sem fulcro”, uma espécie não descrita identificada em estudos anteriores. O DNA barcoding e os algoritmos de delimitação de espécies reconheceram esses três cariomorfos como Unidades Taxonômicas Operacionais Moleculares (MOTUs) independentes, juntamente com outra espécie não descrita previamente reportada como Astrodoras sp. Portanto, Astrodoras é composto de pelo menos quatro espécies que habitam a bacia amazônica, corroborando o status politípico previamente sugerido para o gênero.
Palavras-chave: Cariomorfos, Espécies crípticas, ITS, MOTUs, U2 snDNA.
Introduction
The Doradidae family comprises over 96 valid species of freshwater fish, widely distributed across most hydrographic systems in South America (Sabaj, Arce, 2021; Fricke et al., 2025). Both molecular (Moyer et al., 2004; Arce et al., 2013; Sabaj, Arce, 2021) and morphological evidence (Higuchi, 1992; Birindelli, 2014) support the monophyletic status of Doradidae, which is currently subdivided into six subfamilies: Acanthodoradinae, Agamyxinae, Astrodoradinae, Doradinae, Rhinodoradinae, and Wertheimerinae (Sabaj, Arce, 2021). Astrodoradinae was first recognized by Higuchi et al. (2007) and included the genera Anadoras, Astrodoras, Scorpiodoras, Amblydoras, Physopyxis, Merodoras, and Hypodoras. A few years later, Sousa (2010) conducted a taxonomic review of Astrodoradinae, which he considered to include two other genera, Acanthodoras and Agamyxis. Subsequently, based on mitochondrial and nuclear genes, Arce et al. (2013) also corroborated the monophyly of Astrodoradinae sensu Higuchi et al. (2007), thus excluding Acanthodoras and Agamyxis. In addition, Merodoras was considered a junior synonym of Amblydoras (Arce et al., 2013; Sabaj, Arce, 2021). More recently, Sabaj, Arce (2021) classified Acanthodoras and Agamyxis into their own subfamilies, Acanthodoradinae and Agamyxinae, further upholding the monophyly of Astrodoradinae as proposed by Higuchi et al. (2007). Currently, Astrodoradinae is composed of six genera: Anadoras, Astrodoras, Scorpiodoras, Physopyxis, Hypodoras, and Amblydoras.
Astrodoras is a monotypic genus represented solely by A. asterifrons, originally described by Kner (1853, 1855) from specimens collected in the Negro River near Manaus and the Guaporé River. Based on Kner’s (1853) line drawing of the disembodied gas bladder of the type-specimen, A. asterifrons is distinguished by having the gas bladder with terminal diverticula completely united via longitudinal septum, effecting attenuate structure with a single distal tip. Eigenmann (1925) was first to document intraspecific variation in A. asterifrons by noting that the gas bladder in his specimens from Santarém resembled Kner’s A. asterifrons, but that a single specimen from Jutaí had a gas bladder with a terminal diverticulum “consisting of two sausage-shaped tubes directed sideways” (Eigenmann, 1925:324). Roa-Fuentes et al. (2010) used Eigenmann’s (1925) character (i.e., terminal diverticulum with divergent tips) to identify a new and undescribed species of Astrodoras in the Colombian Amazon.
Sousa (2010) conducted an extensive taxonomic revision of Astrodoras and identified two putative undescribed species: Astrodoras sp. “sem fulcro” and Astrodoras sp. “fulcro”, both remain undescribed. These species occur sympatrically in several locations in the Amazon River basin, including the Catalão Lake and the Negro River at the Anavilhanas fluvial archipelago. According to Sousa (2010), these two undescribed species can be distinguished by the degree of development of the fulcra (i.e., bony plates framing the caudal peduncle, formed by the dorsal and ventral expansions of the procurrent caudal-fin rays), the shape of the upper edge of the orbit, and the shape of the gas-bladder terminal diverticula (e.g., singular in A. asterifrons and A. sp. “sem-fulcro” vs. with divergent tips in A. sp. “fulcro”). Sabaj, Arce (2021) similarly described variation in the size, shape, and articulation of the fulcra in Astrodoras and demonstrated the enlarged fulcra to be the fusion of at least two procurrent elements. Nevertheless, the diversity within Astrodoras remains unresolved, particularly regarding the status of these putative new species, which warrants further investigation using complementary approaches.
Genetic data have proven to be powerful tools in exploring species diversity (Souza-Shibatta et al., 2013; Carvalho et al., 2018; Argolo et al., 2020). Among these, DNA barcoding (i.e., the 600 bp fragment of the COI) has demonstrated high effectiveness for taxonomic identification and resolution of species-level boundaries across a wide array of taxa (Hebert et al., 2003; Ward et al., 2005; Souza-Shibatta et al., 2018). Notably, combining the DNA barcoding with cytogenetics has also provided valuable information for species delimitation in groups with complex taxonomy, cryptic species, or underestimated diversity (Prizon et al., 2017; Carvalho et al., 2018; Anjos et al., 2019). Chromosomal studies have emerged as powerful tools for characterizing species and understanding the genetic mechanisms underlying diversification. Since the late 1970s, they have provided key insights into numerous Neotropical fish groups (e.g., Bertollo et al., 1978; Moreira-Filho, Bertollo, 1991; Tenório et al., 2013; Paiz et al., 2015; Takagui et al., 2017a, 2020; Ferreira et al., 2017; Santos et al., 2021). Within Doradidae, cytogenetic data have contributed to resolving complex taxonomic questions, such as the potential placement of Franciscodoras marmoratus within the Wertheimerinae subfamily (Takagui et al., 2019), and support for Anadoras sp. “araguaia” (sensu Sousa, 2010) as a distinct species (Takagui et al., 2022). However, the only sequences available for Astrodoras are restricted to two mitochondrial and one nuclear marker that were applied in phylogenetic reconstructions (Arce et al., 2013; Sabaj, Arce, 2021), and to date no chromosomal data have been published for Astrodoras (Tab. S1).
Based on the putative cryptic diversity revealed by Eigenmann (1925) and Sousa (2010), Astrodoras may not represent a monotypic genus. Therefore, in this study, we investigate this hypothesis by applying a combination of molecular (DNA barcoding) and cytogenetic analyses (Giemsa staining, C-banding, and physical mapping of 18S rDNA, 5S rDNA, U2 snDNA, and telomeric sequences) to samples of Astrodoras collected near the Anavilhanas fluvial archipelago and at the Barcelos sandbanks, both located in the Negro River basin. Through this approach, we aim to enhance our understanding of the taxonomic status of Astrodoras,as well as to expand the knowledge of chromosomal diversity within Doradidae.
Material and methods
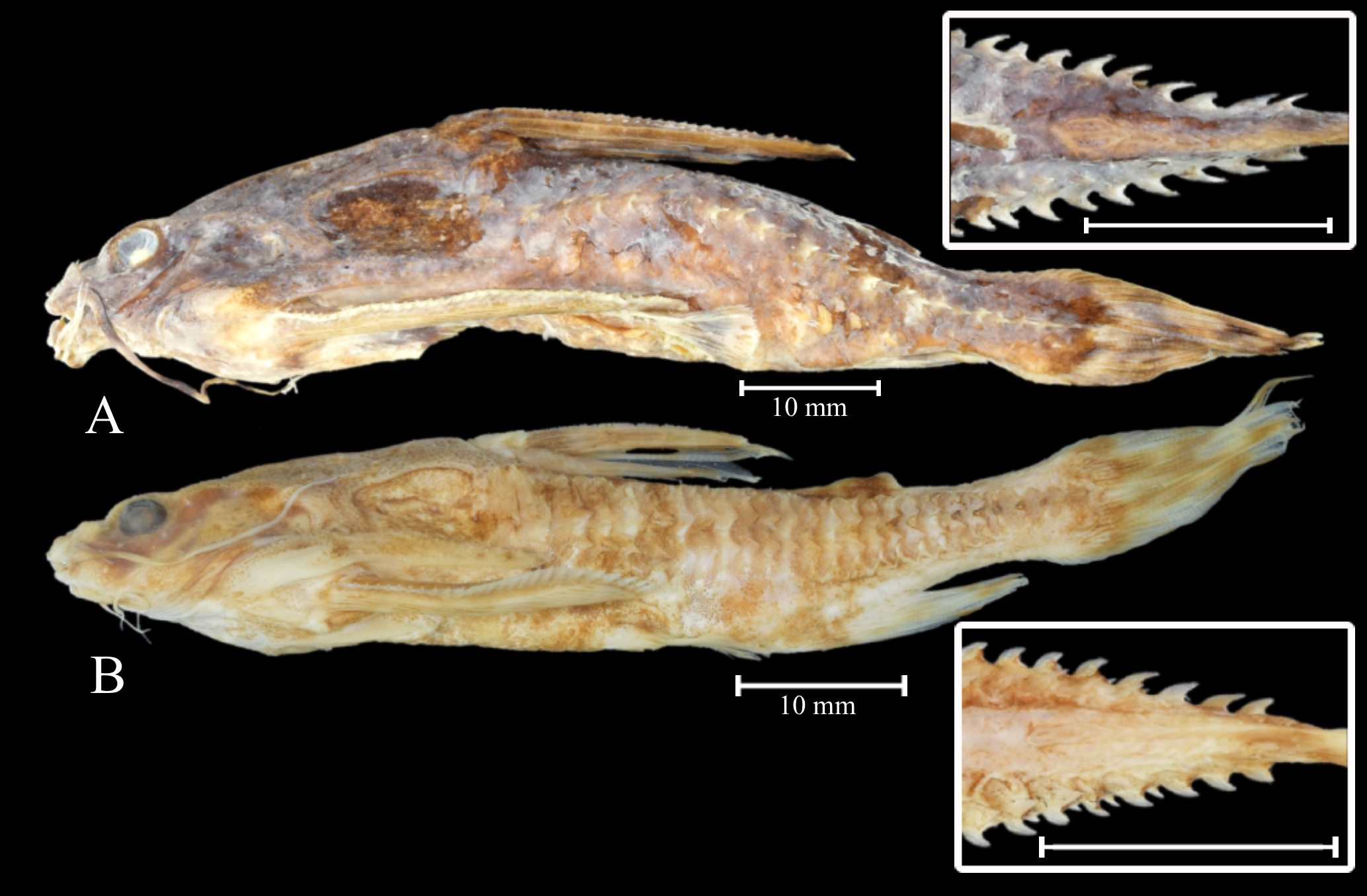
Species identification and collection sites. We analyzed 12 individuals collected from two localities along the Negro River. Eight individuals (four males and four females) (Fig. 1A), collected near the fluvial Anavilhanas archipelago, Amazonas State, Brazil (02°39’28.1”S 60°39’02.4”W), possess the procurrent caudal-fin rays well developed and the lateral border of the cranium above the orbit distinctly elevated, and thus fit the diagnosis of the nominal Astrodoras asterifrons (sensu Sousa, 2010). The remaining four individuals (three males and one female) (Fig. 1B) from the upper Negro River in the Barcelos municipality, Amazonas State, Brazil (00°57’27.41”S 62°55’10.67”W) show the procurrent caudal-fin rays not expanded and the lateral border of the cranium above the orbit only slightly elevated, and thus fit the diagnosis of the A. sp. “sem fulcro” (sensu Sousa, 2010), being herein identified as Astrodoras sp. The gas bladders of these specimens were removed during preparation and not preserved for specimen identification. The specimens are deposited in the Museu de Zoologia at the Universidade Estadual de Londrina (MZUEL) under the voucher numbers MZUEL 17800 and 17813 (Tab. S2).
FIGURE 1| Samples used in this study: A. Astrodoras asterifrons, MZUEL 17813,72.7 mm SL, from Negro River, near Anavilhanas; B. Astrodoras sp., MZUEL 17800 57.9 mm SL, Negro River at Barcelos. The caudal peduncle in dorsal view is presented in the boxes.
Mitotic chromosome preparations and chromosomal banding. Metaphasic chromosomes were obtained from the kidney cells using the air-drying technique described by Bertollo et al. (1978). Briefly, specimens were treated with a 0.02% aqueous colchicine solution (1 ml/100 g) for 30–60 min, followed by euthanasia through clove oil overdose (Griffiths et al., 2000). Subsequently, small pieces of kidney tissue were obtained using surgical scissors and tweezers, homogenized in a KCl solution (0.075 M) with a glass syringe, and incubated for 30 min at 37ºC. After hypotonic treatment with KCl, the cells were treated with a fixative solution composed of methanol and acetic acid (3:1) and stored at -20ºC. For karyotype analyses, chromosomes were stained with Giemsa 5% diluted in phosphate buffer, pH = 6.8, and classified according to arm ratio as acrocentric, subtelocentric, submetacentric, or metacentric (Levan et al., 1964). The constitutive heterochromatin (CH) pattern was detected using the acid/base/salt method described by Sumner (1972), with modifications in the staining process: the chromosomes were stained with a solution composed of 0.7 μL of propidium iodide (50 μg/mL) and 20 μL of antifading solution (Vector, Burlingame, CA, USA), according to Lui et al. (2012).
Probes for fluorescent in situ hybridization. We investigated the distribution of four repetitive DNA sequences: two rDNAs, one snDNA, and telomeric motifs (TTAGGG)n. The 18S rDNA probes were obtained from a mini-prep of Prochilodus argenteus (Hatanaka, Galetti, 2004) and labeled with biotin-16-dUTP (Bio-Nick-Translation mix, Roche Diagnostics, Mannheim, Germany). The 5S rDNA probes were obtained from a mini-prep of Megaleporinus elongatus (=Leporinus elongatus Valenciennes, 1850) (Martins, Galetti, 1999) and labeled with digoxigenin-11-dUTP (Dig-Nick-Translation mix, Roche Diagnostics). Telomeric probes were obtained through a polymerase chain reaction (PCR) using a pair of self-complementary primers (Ijdo et al., 1991) and labeled with tetramethyl-rhodamine-5-dUTP (red; Roche Diagnostics). The U2 snDNA probes were obtained through PCR using the Trachelyopterus galeatus total DNA (extracted from liver tissues) and the primers described by Bueno et al. (2013). The PCR products were sequenced and deposited in GenBank under the sequence ID: OK166555.1 (see Haerter et al., 2022). For FISH experiments, U2 snDNA probes were labeled by PCR using fluorescein-12-dUTP (Roche Diagnostics).
Fluorescence in situ hybridization (FISH) experiments. FISH experiments were adapted from Yano et al. (2017a). Briefly, slides pretreatment included the following steps: (a) two washes in 2x SSC solution for 5 min at room temperature, (b) treatment with RNAse solution (100 µg/mL in 2x SSC) for 1 h at 37ºC in a dark, moist chamber, (c) two washes with 2x SSC solution for 5 min at room temperature, and (d) dehydration in an ethanol series (70% and 100%) at -20ºC. Chromosome denaturation was performed using 70% deionized formamide in 2x SSC at 70ºC, while denaturation of the hybridization mixture (200 ng of each probe, 2x SSC, 50% deionized formamide, 10% dextran sulfate) was conducted at 90ºC in a dry block for 10 min. Hybridization was performed for 24 h at 37°C in a dark, moist chamber. Post-hybridization stringent washes included once in 15% deionized formamide/2x SSC at 42ºC for 10 min and twice in 0.5% tween/4x SSC (SSCT) at room temperature for 5 min each. The 5S rDNA probe signals were detected using anti-digoxigenin-rhodamine (red; Roche Diagnostics), while the 18S rDNA probe signals were detected using avidin fluorescein isothiocyanate (green; avidin-FITC) (Roche Diagnostics). Chromosome counterstaining was performed with fluorochrome 4’,6’-diamidino-2-phenylindole (blue; DAPI) in antifading solution (Vector).
DNA extraction and molecular analyses. Genomic DNA was extracted from the muscle tissue of Astrodoras species using the phenol-chloroform method described by Almeida et al. (2001). The mitochondrial cytochrome C oxidase subunit I (COI) gene was amplified by PCR using the primers described by Ward et al. (2005). Each PCR reaction contained 50 ng of genomic DNA, 0.25 µM of each primer, GoTaq® Master Mix (Promega), and deionized water to fill the final volume. The amplification protocol included: (a) 2 min at 94ºC; (b) 35 cycles of 1 min at 94ºC, 30 sec at 54ºC, and 90 sec at 72ºC; and (c) a final extension of 5 min at 72ºC. Amplicons were checked in a 1% agarose gel electrophoresis. For the sequencing reaction, we used the Big Dye Sequence Terminator kit (Applied Biosystems), according to the manufacturer’s instructions. The amplification conditions consisted of 25 cycles at 96ºC for 30 sec, 15 sec at 50ºC, and 4 min at 60ºC. After the reaction, the amplicons were precipitated. The reading was performed with the sequencer model ABI PRISM 3500 XL Genetic Analyzer from Applied Biosystems made by HITACHI.
Sequence alignment and molecular delimitation analysis. In this study, four sequences were generated and deposited on the Barcode of Life Data Systems (BOLD) platform under the project “Species delimitation of thorny catfishes (Doradidae-Astrodoradinae): an integrated analysis based on Chromosomal and DNA barcoding – DORBC”. These sequences were automatically assigned to a barcode index number (BIN) (i.e., a group of sequences corresponding to a single taxon), following the analytical procedures of Ratnasingham, Hebert (2013). Additionally, three sequences of Astrodoras and 16 sequences from other Astrodoradinae species previously deposited on BOLD were included in the analysis. Sequences from Wertheimeria maculata (Wertheimerinae) and Platydoras armatulus (Valenciennes, 1840)(Doradinae) were used as outgroups (Tab. S2).
The dataset was aligned on BioEdit Sequence Alignment Editor v. 7.1.9. (Hall, 1999) with the ClustalW tool (Thompson et al., 1994). Intra- and interspecific genetic distance matrices were generated in MEGA v. 6 (Tamura et al., 2011). The Neighbor-Joining (NJ) tree was generated based on the Kimura-2-Parameter (K2P) nucleotide substitution model (Kimura, 1980) with 1,000 bootstrap replicates. Phylogenetic reconstructions were performed using maximum likelihood (ML) and Bayesian inference (BI) methods in RAxML v. 8.2 (Stamatakis, 2014) and MrBayes v. 3.2.6 (Ronquist, Huelsenbeck, 2003), respectively, through the CIPRES Science Gateway v. 3.3 (Miller et al., 2010). For BI, we used two rows of four chains (10 × 10⁶ generations) with sampling at each 1,000 trees, assuming 10% of burn-in. Branch support values were calculated using posterior probabilities (BI) and 1,000 bootstrap replicates (ML).
The barcodes obtained in the present study were submitted to the analysis of three algorithms based on single-locus data commonly employed for species delimitation: barcode index numbers (BIN) (Ratnasingham, Hebert, 2013), single General Mixed Yule-Coalescent analysis (GMYC) (Pons et al., 2006), and Bayesian Poisson Tree Process (bPTP) (Zhang et al., 2013). BIN analyses were performed by comparing the input taxa with all others that share identical BINs, including those uploaded by different users based on the Refined Single Linkage (RESL) algorithm. This system divided the COI sequences uploaded in the BOLD systems into Molecular Operational Taxonomic Units (MOTUs) independently of their predefined taxonomic classification, providing a framework for assessing congruence between DNA barcodes and species designations (Ratnasingham, Hebert, 2013). The sGMYC analysis was performed for mtDNA lineages delimitation (Fujisawa, Barraclough, 2013), available on https://species.h-its.org/gmyc/, and using an ultrametric tree as input, obtained in the BEAST v. 1.8.2 software (Drummond et al., 2012). For bPTP analysis, we used a maximum likelihood phylogenetic tree constructed in the RAxML-HPC BlackBox v. 8.2.10 as the input file. The bPTP adds Bayesian inference support values to the branches of the inputted tree, enhancing reliability (https://species.h-its.org/).
Results
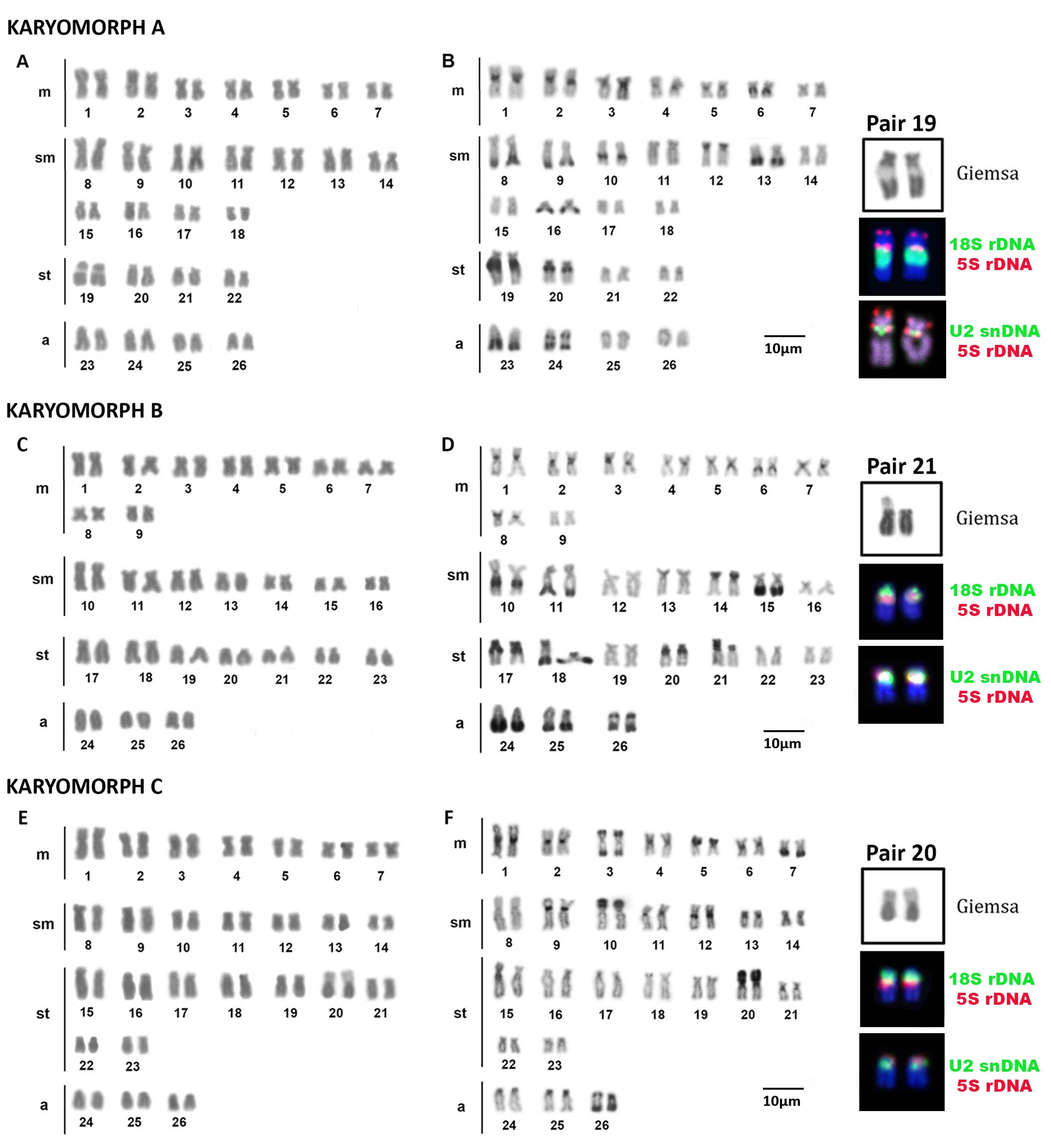
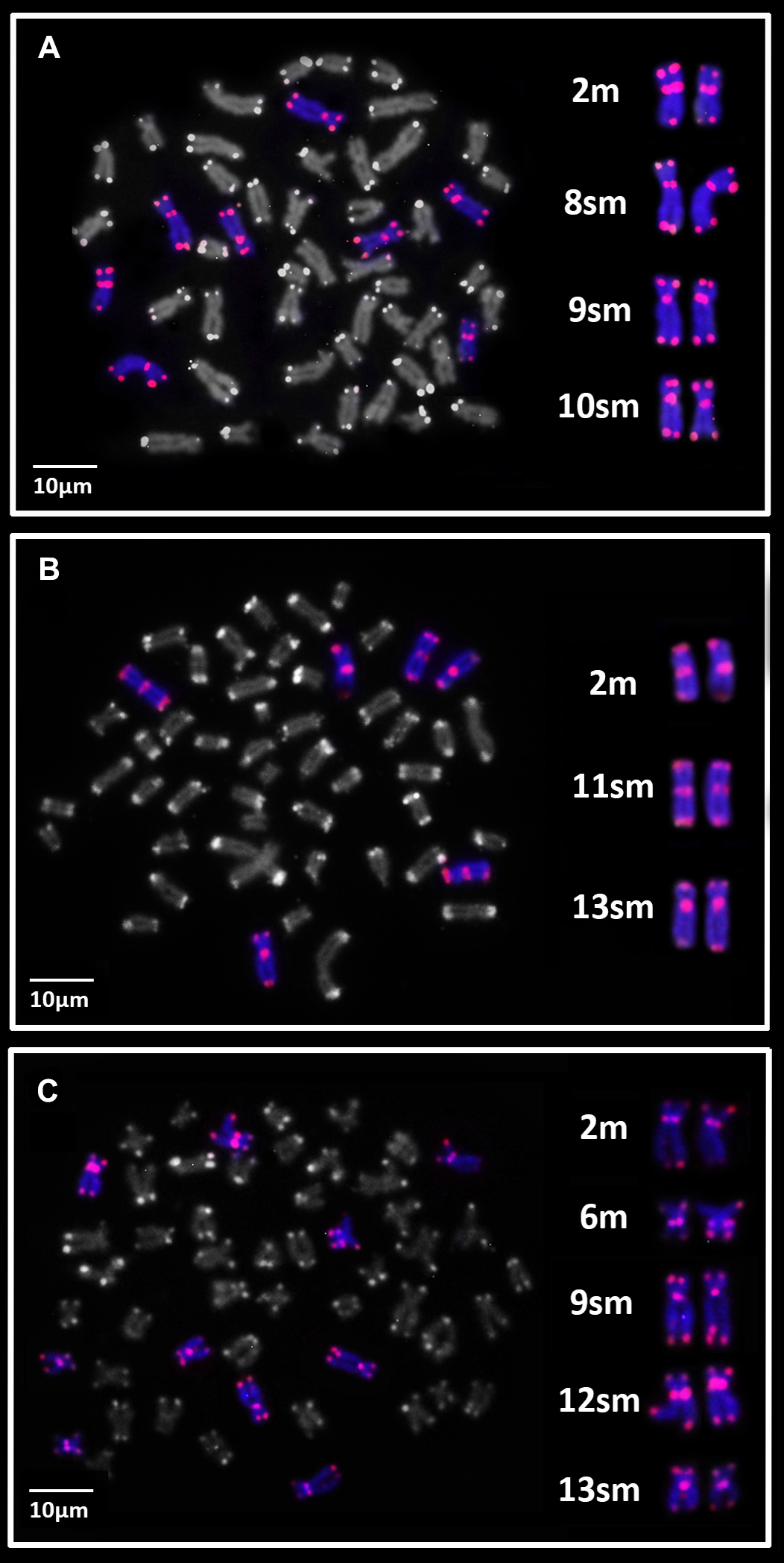
Astrodoras asterifrons. Three of the eight specimens collected in the Negro River, near the Anavilhanas archipelago, exhibited 2n = 52 chromosomes and a karyotype composed of 14m + 22sm + 8st + 8a chromosomes for both sexes. These individuals were designated as Karyomorph A in this study (Fig. 2A). C-banding revealed CH blocks in pericentromeric regions of chromosome pairs 1m, 2m, 3m, 4m, and 5m; interstitial blocks on the long arm of the chromosome pairs 6m, 10sm, 19st, and 20st; terminal blocks on the short arm of chromosome pair 12sm; and terminal blocks on the long arm of chromosome pairs 8sm, 9sm, 13sm, 16sm, and 23a. The chromosome pair 24a exhibited interstitial CH on the long arm (Fig. 2B). FISH with 18S rDNA probes revealed a single site on the long arm of chromosome pair 19st (Fig. 2B – box). FISH with 5S rDNA probes revealed two sites on chromosome pair 19st: one site on the terminal position of the short arm and another site on the proximal position of the long arm (Fig. 2B – box). FISH with telomeric probes (TTAGGG)n showed fluorescent signals at the terminal positions of all chromosomes. Additionally, interstitial telomeric sites (ITSs) were observed in the chromosome pairs 2m, 8sm, 9sm, and 10sm (Fig. 3A). The U2 snDNA sites were detected in the long arm of the chromosome pair 19st (Fig. 2B – box).
FIGURE 2| Karyotypes of Astrodoras asterifrons sensu lato after conventional and molecular cytogenetics methods: karyomorph A after (A) Giemsa staining (B) C-banding; karyomorph B after (C) Giemsa staining (D) C-banding; karyomorph C (E) Giemsa staining (F) C-banding. The boxes contain the chromosome pairs with the 18S (green) and 5S rDNA sites (red), and with the U2snDNA (green) and 5S rDNA sites (red).
The remaining five specimens also exhibited 2n = 52 chromosomes, with a karyotype composed of 18m + 14sm + 14st + 6a chromosomes, and no sexual dimorphism in chromosome number or morphology was observed. These specimens were designated as Karyomorph B in this study (Fig. 2C). C-banding revealed CH blocks in pericentromeric regions of pairs: 1m, 2m, 3m, 4m, 5m, 6m, 7m, and 8m; interstitial blocks on the long arm of chromosome pairs: 10sm, 17st, and 20st; terminal blocks on the short arm of chromosome pairs 14sm, 18sm, 17st, and 21st; and terminal blocks on the long arm of chromosome pairs: 11sm, 15sm, 18st, 24a, and 25a. The pair 26 exhibited proximal and terminal CH on the long arm (Fig. 2D). FISH with 18S rDNA probes revealed two terminal sites on the short arm of pair 21st (Fig. 2D – box). 5S rDNA probes revealed a single proximal site in the short arm of pair 21st (Fig. 2D – box). FISH with telomeric probes (TTAGGG)n evidenced fluorescent signals in terminal positions of all chromosomes and ITSs in the pairs 2m, 11sm, and 13sm (Fig. 3B). The U2 snDNA sites were detected on the long arm of the chromosome pair 21st, the same chromosomes carrying 18S and 5S rDNA sites (Fig. 2D – box).
FIGURE 3| Metaphase plates of Astrodoras asterifrons sensu lato showing the (TTAGGG)n distribution pattern (red). A. karyomorph A; B. karyomorph B; C. karyomorph C.
Astrodoras sp. All specimens from the Negro River in Barcelos exhibited 2n = 52 chromosomes, with a karyotype composed of 14m + 14sm + 18st + 6a for both sexes (Fig. 2E). These individuals were designated as Karyomorph C in this study. C-banding revealed CH blocks on pericentromeric regions of the chromosome pairs: 1m, 2m, 4m, 6m, 9sm, 11sm, 12sm, 13sm, 21st, 24a, and 25a; terminal blocks on the long arm of the chromosome pairs 5m, 10sm, and 20st; and terminal CH blocks on the long arm of the chromosome pair 7m. The chromosome pair 3m had CH blocks in terminal position of both chromosomal arms, while the chromosome pair 26a exhibited proximal and terminal CH blocks on the long arm (Fig. 2F). FISH with 18S rDNA probes revealed single, terminal sites on the short arm of chromosome pair 20st (Fig. 2F – box). FISH with 5S rDNA probes also revealed sites on the short arm of chromosome pair 20st but in a proximal position (Fig. 2F – box). FISH with telomeric probes (TTAGGG)n evidenced fluorescent signals in terminal positions of all chromosomes and ITSs on the chromosome pairs 2m, 6m, 9sm, 12sm, and 13sm (Fig. 3C). The U2 snDNA sites were detected on the long arm of the chromosome pair 20st, which also carries the 18S and 5S rDNA clusters (Fig. 2F – box).
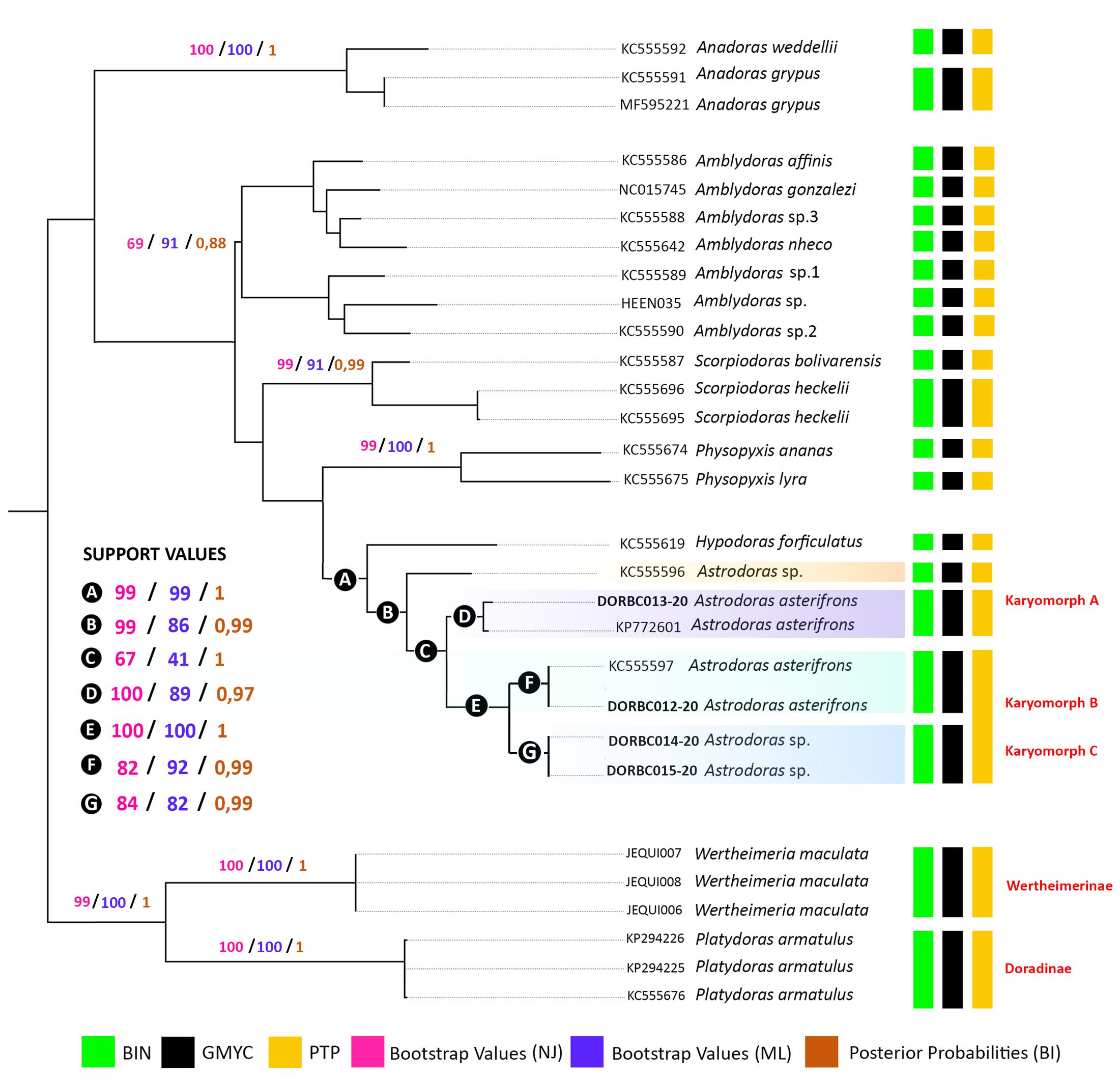
Molecular species delimitation analysis based on a fragment of the mtDNA COI gene. The COI sequences of obtained from seven specimens of Astrodoras resulted in a 604 bp dataset, with no deletions, insertions, or stop codons. The NJ, ML, and BI methods produced similar topologies, revealing seven major clades with high support values (above 90% bootstrap and 0.9 posterior probability): Wertheimeria and Platydoras (outgroups), Scorpiodoras, Physopyxis, Anadoras, Amblydoras affinis, Amblydoras sp., and Hypodoras + Astrodoras. The Astrodoras specimens from our sample were subdivided into three well-supported clades, corresponding to the three karyomorphs identified in cytogenetic analyses (A, B, and C). The specimens of Astrodoras asterifrons from near Anavilhanas correspond to Karyomorph A and B, while Astrodoras sp. corresponds to Karyomorph C.
To estimate genetic distances, two grouping approaches were applied based on morphological classification and molecular analyses results. The first approach considered all sequences of A. asterifrons of our sample and from available databases as a single group that was named as A. cf. asterifrons, revealing an intraspecific distance of 0.055 (5.5%) (Tab. S3). A second morphological grouping (named as A. asterifrons), excluding one sequence from the Madeira River (05°48’33.0”S 61°18’01.1”W) identified as Astrodoras sp. (sensu Roa-Fuentes et al., 2010) in Sabaj, Arce (2021), demonstrated an intraspecific distance of 0.046 (4.6%).
Using the second grouping approach, based on the subdivisions identified in the phylogenetic reconstructions, which also correspond to the karyomorphs identified in our sample through cytogenetic analyses, Astrodoras samples were divided into three groups: group 1 (karyomorph A + KP772601), group 2 (karyomorph B+ KC555597), and group 3 (karyomorph C).The interspecific genetic distances ranged from 2.9% (between groups 2 and 3) to 6.5% (between groups 1 and 3) and 7% (between groups 1 and 2). The intraspecific distances ranged from 0.7% in group 3 and 0.9% in group 1 to 1.4% in group 2 (Tab. S3).
Species delimitation algorithms yielded slightly different MOTU counts: BIN and GMYC identified 20 MOTUs, whereas bPTP identified 19. Despite this variation, all algorithms consistently recovered the three karyomorphs (A, B, and C) as distinct clades with strong support (above 90%; Fig. 4).
FIGURE 4| Consensus tree (NJ neighbor-joining, ML maximum likelihood and BI Bayesian inference) based on COI sequences from samples of Astrodoras asterifrons sensu lato (the same specimens cytogenetically analyzed), species of Astrodoradinae, Wertheimeria maculata and Platydoras armatulus (outgroup). The vertical-colored bars represent the results obtained after the species delimitation analyses.
Discussion
Karyotypic data for Doradidae have been predominantly reported for species within the subfamilies Doradinae (Milhomem et al., 2008; Takagui et al., 2021) and Wertheimerinae (Eler et al., 2007; Takagui et al., 2019). In contrast, cytogenetic information for Astrodoradinae remains scarce, limited to the three species of Anadoras, all presenting 2n = 56 chromosomes (Baumgärtner et al., 2018; Takagui et al., 2022). Our study expands the chromosomal dataset to A. asterifrons, revealing a 2n = 52 chromosomes, the lowest reported to date for Doradidae. Moreover, we identified three distinct karyomorphs within Astrodoras. Karyomorphs A and B were collected in sympatry in the Negro River, near Anavilhanas, while Karyomorph C was sampled in the Negro River near Barcelos City (500 km upstream from Anavilhanas). All karyomorphs exhibited 2n = 52 chromosomes, with variations in karyotype composition, CH distribution, rDNA and U2 snDNA site organization, and the number of interstitial telomeric sequences (ITS). Species delimitation analyses based on molecular data support the interpretation of these karyomorphs as independent evolutionary units, thus reinforcing the hypothesis of a polytypic status for Astrodoras.
Ancestral state reconstruction suggests that 2n = 58 constitutes the plesiomorphic condition for Doradidae. Thus, chromosome diversification in the thorny catfishes can be explained by three main evolutionary pathways: (a) maintenance of the ancestral 2n = 58 chromosomes; (b) reduction in 2n via chromosome fusions; and (c) increase in 2n through chromosome fissions (see Takagui et al., 2024). The specimens of A. asterifrons in our sample exhibit the smallest 2n reported among thorny catfishes so far (2n = 52), corresponding to the chromosomal diversification pathway (b) 2n reduction through chromosome fusions. This type of rearrangement may occur in two situations: loss of the capping function while telomeric repeats are still present or the loss of telomeric repeats that maintain the telomere functional (Slijepcevic, 1998; Murnane, 2006). According to Ocalewicz (2013), pericentromeric and interstitial telomeric sites may represent relicts of the ancient chromosomal fusion events and can be classified into six types: heterochromatic (het-ITSs), short (s-ITSs), large ITSs in restricted euchromatic regions (restricted eu-ITSs), long subtelomeric ITSs, fusion ITSs, and pericentromeric ITSs (Lin, Yan, 2008; Ruiz-Herrera et al., 2008; Schmid, Steinlein, 2016).
In Astrodoras, the hypothesis of chromosomal fusion is further corroborated by FISH analyses using telomeric probes, which revealed ITSs in all examined specimens, varying from three sites in karyomorph B and four sites in karyomorph A to five sites in karyomorph C. Despite this variation, all karyomorphs share at least three chromosome pairs (metacentrics and submetacentrics) carrying ITSs, suggesting that these chromosomal fusions might represent an ancestral event in Astrodoras. However, karyomorphs A and C exhibit additional ITSs, which might represent remnants of other types of structural rearrangements, such as pericentric inversions (reviewed in Vicari et al., 2022). Alternatively, some of these additional ITSs could also represent Het-ITS, that is, TTAGGG repeats associated with heterochromatic regions (e.g., Milhomem et al., 2008; Ruiz-Herrera et al., 2008; Scacchetti et al., 2011; Marajó et al., 2018; Campos et al., 2020; Viana et al., 2020; Vicari et al., 2022).
Chromosomal fusions have been reported only in a few species of Doradidae. For instance, Baumgärtner et al. (2016) identified two proximal ITSs on the long arm of the chromosome pair 19 in Trachydoras paraguayensis (Eigenmann & Ward, 1907) (2n = 56), suggesting that the 2n reduction in this species was the result of the fusion of acrocentric and submetacentric chromosomes. In contrast, Anadoras species lacks ITSs despite their reduced 2n = 56 chromosomes, making it difficult to determine the type of fusion (in tandem or centric) or to identify the fused chromosome pairs (Takagui et al., 2022). In the sister family Auchenipteridae, ITSs have been identified in three species so far: Ageneiosus inermis (Linnaeus, 1766) which has one ITS (Lui et al., 2013); Centromochlus schultzi Rössel, 1962, which has two ITSs(Kowalski et al., 2024); and Centromochlus heckelii (De Filippi, 1853) which exhibits several ITSs (Haerter et al., 2025). In A. inermis, the ITS was attributed to a single end-to-end chromosomal fusion, while in C. schultzi, the heterochromatic ITSs (het-ITSs) might be either a result of pericentric inversions or repetitive DNA amplification (Kowalski et al., 2024). Interestingly, the origin of ITSs in C. heckelii is possibly related to multiple mechanisms, such as chromosomal fusions, inversions, transpositions, or association with satellite DNA (Haerter et al., 2025).
C-banding revealed distinct CH distribution patterns across the Astrodoras karyomorphs. They share certain chromosomal pairs, including the chromosome pairs carrying the 18S rDNA, a submetacentric pair with CH in the entire short arm (pairs 12, 13, and 10), and an acrocentric chromosome pair with two CH blocks on the long arm (pairs 24, 26, and 26). However, Karyomorph C exhibited fewer CH blocks compared to the other two karyomorphs. Thus, the CH distribution may constitute an important chromosomal marker to differentiate them. Variation in CH has been fundamental to understanding cytotaxonomic issues and some diversification pathways in Doradidae (Baumgärtner et al., 2018; Takagui et al., 2019, 2021, 2022). For instance, in the subfamily Wertheimerinae, CH arrangements are highly divergent among the three studied species, likely influenced by the saltatory activity of retroelements, particularly from the Rex3 group (Takagui et al., 2019). In the Doradinae, the CH distribution is crucial to karyotypically discriminate Centrodoras brachiatus (Cope, 1872), Pterodoras granulosus (Valenciennes, 1821), and Oxydoras niger (Valenciennes, 1821), which share the same diploid number, karyotype composition, and rDNA distribution patterns (Takagui et al., 2021). Furthermore, C-positive regions were crucial during the differentiation of the ZZ/ZW sex chromosome system reported in Nemadoras trimaculatus (= Tenellus trimaculatus (Boulenger, 1898)) (sensu Sabaj, Arce, 2021) from the Araguaia River (Takagui et al., 2017a) and in the evolution of supernumerary chromosomes (B chromosomes) of Doras punctata (=Ossancora punctata (Kner, 1853)), Platydoras armatulus, and Pterodoras granulosus (Takagui et al., 2017b).
Physical mapping with 18S and 5S rDNA probes showed sites in the same chromosome pair across all Astrodoras karyomorphs but with some notable differences in the arrangement of these repetitive DNAs. In karyomorphs B and C, the rDNA loci are syntenic on the short arm of the chromosome pairs 20/21st. On the other hand, in Karyomorph A, the 18S rDNA locus is also located on a subtelocentric pair, but in an interstitial position on the long arm. It is noteworthy that simple, terminal 18S rDNA sites represent the putative plesiomorphic condition in Doradidae, inferred through ancestral state reconstruction (Takagui et al., 2021). Similarly, most Doradidae species exhibit the 5S rDNA in a syntenic arrangement with the 18S rDNA cluster (Baumgärtner et al., 2018; Takagui et al., 2019, 2021, 2022), a feature that is absent in most Auchenipteridae species (see Kowalski et al., 2024). Thus, the rDNA pattern observed in Karyomorph A might represent an apomorphic trait, likely resulting from a pericentric inversion. In this scenario, the inversion would have encompassed the entire 18S rDNA cluster and part of the 5S rDNA cluster, leading to a fragmented 5S rDNA site. As a result, one portion of the 5S rDNA cluster remains at the terminal position of the short arm, while the other portion was relocated to an interstitial position on the long arm alongside the 18S rDNA site. This inversion hypothesis is also supported by the U2 snDNA mapping, which revealed interspersed U2/5S rDNA sites on the short arms of the chromosome pair 20 and 21 of karyomorphs B and C, respectively, whereas the karyomorph A exhibits the U2 snDNA loci associated only with the 5S fraction situated on the long arm, near to the 18S rDNA sites.
The organization and structure of snDNAs remain poorly understood in Neotropical fishes. In Doradidae, cytogenetic mapping of U2 snDNA has been conducted in two species, Acanthodoras cf. polygrammus (Linnaeus, 1758) and Acanthodoras cataphractus (Linnaeus, 1758), which possess two and three sites, respectively (Takagui et al., 2024). Both species exhibit a syntenic arrangement of the U2 snDNA and 5S rDNA loci, a pattern consistent with that observed in Anadoras species in this study. Among neotropical fishes, this syntenic arrangement is relatively common and has been reported in characiforms Piabarchus stramineus (Eigenmann, 1908) (Piscor et al., 2018), Triportheus (Yano et al., 2017b), and ten species of Characidium (Scacchetti et al., 2015), as well as the gymnotiform Eigenmannia aff. trilineata (Araya-Jaime, 2017). Interestingly, this syntenic arrangement is absent in Auchenipteridae species (see Haerter et al., 2022). As noted by Takagui et al. (2024), such differences may serve as important cytotaxonomic markers, offering insights into karyotype differentiation and evolutionary processes in Doradidae and Auchenipteridae.
The taxonomic diagnosis of the Astrodoras speciesremains challenging. Although previous studies (Eigenmann, 1925; Roa-Fuentes et al., 2010; Sousa, 2010) identified putative undescribed taxa within Astrodoras, these species have not yet been formally described. Our cytogenetic analyses revealed three distinct karyomorphs within Astrodoras, characterized by differences in CH distribution, the positions of 18S rDNA, 5S rDNA, and U2 snDNA sites, and the number of chromosomes carrying ITSs. The species delimitation analyses indicate that these karyomorphs represent independent evolutionary units, supported by high phylogenetic support values (NJ, BI, and ML). These units are also recognized by BIN and GMYC, two of the most reliable algorithms for species delimitation and the identification of MOTUs in fish (e.g., Costa-Silva et al., 2015; Carvalho et al., 2018; Anjos et al., 2019; Argolo et al., 2020). Additional evidence for the distinction of these karyomorphs is provided by the absence of hybrids between karyomorphs A and B, despite their sympatric occurrence.
The intraspecific genetic distance for the morphology-based grouping was high (4.6%), exceeding by more than twice the commonly accepted threshold value (2%) for fish species delimitation using the DNA barcoding method (Pereira et al., 2011, 2013). When considering the three MOTUs in Astrodoras, the interspecific distances (2.9%, 6.5%, and 7%) are consistent with previously reported values for congeneric species (Pereira et al., 2011, 2013) (Tab. S3). Despite including only two specimens, phylogenetic analyses performed by Arce et al. (2013) and Sabaj, Arce (2021) had already recognized the non-monotypic status of Astrodoras, identifying at least two distinct taxonomic units. Considering all available Astrodoras sequences, including those from Arce et al. (2013) and Sabaj, Arce (2021), the molecular analyses recovered at least four distinct lineages: (1) ‘Astrodoras sp. (sensu Roa-Fuentes et al., 2010)’ from Arce et al. (2013) and Sabaj, Arce (2021); (2) karyomorph B + ‘A. asterifrons’ from Arce et al. (2013) and Sabaj, Arce (2021); (3) karyomorph A + A. asterifrons (KP772601); and (4) karyomorph C. Thus, integration of cytogenetic markers and COI-based species delimitation analyses further supports the polytypic status of Astrodoras.
Currently, it is widely known that the Amazon drainage system exhibits seasonal oscillations of water levels that induce the emergence and disappearance of many landscape formations, such as rivers, lakes, streams, beaches, floodplains, and flooded forests (Albert, Reis, 2011). This vast hydrographic system has an old and complex geomorphological history, marked by successive episodes of geomorphological changes and marine incursions (Hoorn et al., 2010, 2017; Wesselingh, Hoorn, 2009; Cassemiro et al., 2023). This highly complex scenario favors vicariant events and allopatric speciation (Dagosta, de Pinna, 2017), which is congruent with some Doradidae genera that were already the focus of taxonomic revisions, such as Physopyxis (Sousa, Rapp Py-Daniel, 2005) and Scorpidoras (Sousa, Birindelli, 2011). The region where these A. asterifrons samples were collected has historically served as a basin divider, particularly due to the presence of the Purus Arch (Lundberg et al., 1998). Consequently, vicariant events associated with past geomorphological shifts in the region may underlie the chromosomal and molecular divergence observed among thekaryomorphs. In this scenario, the sympatric occurrence in the Central Amazon River floodplains might be a result of the Late Miocene rise of the Vaupes structural arch and concomitant breaching of the Purus arch, which allowed the secondary contact among several freshwater fish species (Ivanyisky, Albert, 2014). Thus, it is possible that the distinct sympatric lineages that compose Astrodoras emerged through allopatric speciation, giving rise to sympatric populations by secondary contact.
The integration of cytogenetic and molecular evidence supports the hypothesis of a polytypic status of Astrodoras. The specimens analyzed in this study exhibited significant chromosomal and molecular divergence, even in cases of sympatric occurrence within the Central Amazon floodplains. These findings underscore the urgency of re-evaluating the taxonomic framework of Astrodoras through the inclusion of additional molecular markers, expanded geographic sampling, and integrative approaches to fully resolve its diversity and evolutionary history.
Acknowledgments
We thank Jansen Zuanon (INPA) for assisting in fish collection; the Instituto Nacional de Pesquisas da Amazônia (INPA) and Universidade Estadual de Londrina (UEL) for providing laboratory infrastructure; and the Instituto Chico Mendes de Conservação da Biodiversidade (ICMBio) for granting the permit to collect biological material. We also thank the Postgraduate Program in Genetics, Conservation, and Evolutionary Biology (PPG GCBEv) for providing logistical and financial support. This study is the result of joint efforts with our esteemed colleague Fábio H. Takagui, who passed away prematurely at the age of 33 (1 June 1989 – 18 April 2023). Fábio was deeply loved by his friends and family and was a remarkable cytogeneticist. We dedicate this study to his beloved mother, Rosania.
References
Albert JS, Bart HL Jr., Reis RE. Species richness and cladal diversity. In: Albert JS, Reis RE, editors. Historical biogeography of Neotropical freshwater fishes. Berkeley: University of California Press; 2011. p.89–104.
Almeida FS, Fungaro MHP, Sodré LMK. RAPD and isoenzyme analysis of genetic variability in three allied species of catfish (Siluriformes, Pimelodidae) from the Tibagi River. J Zool. 2001; 253(1):113–20. https://doi.org/10.1017/S0952836901000103
Anjos MS, Bitencourt JA, Nunes LA, Sarmento-Soares LM, Carvalho DC, Armbruster JW et al. Species delimitation based on integrative approach suggests reallocation of genus in Hypostomini catfish (Siluriformes, Loricariidae). Hydrobiologia. 2019; 847:563–78. https://doi.org/10.1007/s10750-019-04121-z
Arce MH, Reis RE, Geneva AJ, Sabaj Pérez MH. Molecular phylogeny of thorny catfishes (Siluriformes: Doradidae). Mol Phylogenet Evol. 2013; 67(3):560–77. https://doi.org/10.1016/j.ympev.2013.02.021
Araya-Jaime C, Mateussi NTB, Utsunomia R, Costa-Silva GJ, Oliveira C, Foresti F. ZZ/Z0: the new system of sex chromosomes in Eigenmannia aff. trilineata (Teleostei: Gymnotiformes: Sternopygidae) characterized by molecular cytogenetics and DNA barcoding. Zebrafish. 2017; 14:464–70. https://dx.doi.org/10.1089/zeb.2017.1422
Argolo LA, López-Fernández H, Batalha-Filho H, Affonso PRAM. Unraveling the systematics and evolution of the “Geophagus” brasiliensis (Cichliformes: Cichlidae) species complex. Mol Phylogenet Evol. 2020; 150:106855. https://doi.org/10.1016/j.ympev.2020.106855
Baumgärtner L, Paiz LM, Margarido VP, Portela-Castro ALB. Cytogenetics of the thorny catfish Trachydoras paraguayensis (Eigenmann & Ward, 1907) (Siluriformes, Doradidae): evidence of pericentric inversions and chromosomal fusion. Cytogenet Genome Res. 2016; 149(3):201–06. https://doi.org/10.1159/000448126
Baumgärtner L, Paiz LM, Takagui FHT, Lui RL, Moreira-Filho O, Giuliano-Caetano L et al. Comparative cytogenetics analysis on five genera of thorny catfish (Siluriformes, Doradidae): chromosome review in the family and inferences about chromosomal evolution integrated with phylogenetics proposals. Zebrafish. 2018; 15:270–78. https://doi.org/10.1089/zeb.2017.1554
Bertollo LAC, Takahashi CS, Moreira-Filho O. Cytotaxonomic considerations on Hoplias lacerdae (Pisces, Erythrinidae). Rev Bras Genet. 1978; 1:103–20.
Birindelli JLO. Phylogenetic relationships of the South American Doradoidea (Ostariophysi: Siluriformes). Neotrop Ichthyol. 2014; 12(3):451–64. https://doi.org/10.1590/1982-0224-20120027
Bueno D, Palacios-Gimenez OM, Cabral-de-Mello DC. Chromosomal mapping of repetitive DNAs in the grasshopper Abracris flavolineata reveal possible ancestry of the B chromosome and H3 histone spreading. PLoS ONE. 2013; 8:e66532. https://doi.org/10.1371/journal.pone.0066532
Campos AS, Favarato RM, Feldberg E. Interspecific cytogenetic relationships in three Acestrohynchus species (Acestrohynchinae, Characiformes) reveal the existence of possible cryptic species. Comp Cytogen. 2020; 14(1):27–42. https://doi.org/10.3897/CompCytogen.v14i1.33483
Carvalho ML, Costa-Silva GJS, Melo S, Ashikaga FY, Shimabukuro-Dias CK, Scachetti PC. The non-monotypic status of the Neotropical fish genus Hemiodontichthys (Siluriformes, Loricariidae) evidenced by genetic approaches. Mitochondr DNA A. 2018; 29:1224–30. https://doi.org/10.1080/24701394.2018.1431230
Cassemiro FAS, Albert JS, Antonelli A, Menegotto A, Wüest RO, Cerezer F et al. Landscape dynamics and diversification of the megadiverse South American freshwater fish fauna. PNAS. 2023; 120(2):e2211974120. https://doi.org/10.1073/pnas.2211974120
Costa-Silva GJ, Rodriguez MS, Roxo FF, Foresti F, Oliveira C. Using different methods to access the difficult task of delimiting species in a complex Neotropical hyperdiverse group. PLoS ONE. 2015; 10(9):e0135075. https://doi.org/10.1371/journal.pone.0135075
Dagosta FCP, de Pinna M. Biogeography of Amazonian fishes: deconstructing river basins as biogeographic units. Neotrop Ichthyol. 2017; 15(3):e170034. https://doi.org/10.1590/1982-0224-20170034
Drummond AJ, Suchard MA, Xie D, Rambaut A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol. 2012; 29(8):1969–73. https://doi.org/10.1093/molbev/mss075
Eigenmann CH. A review of the Doradidae, a family of South American Nematognathi, or catfishes. Trans Amer Philos Soc. 1925; 22(5):280–365.
Eler ES, Dergam JA, Venere PC, Paiva LC, Miranda GA, Oliveira AA. The karyotypes of the thorny catfishes Wertheimeria maculata Steindachner, 1877 and Hassar wilderi Kindle, 1895 (Siluriformes: Doradidae) and their relevance in doradids chromosomal evolution. Genetica. 2007; 130:99–103. https://doi.org/10.1007/s10709-006-0023-4
Ferreira M, Garcia C, Matoso DA, Jesus IS, Cioffi MB, Bertollo LAC et al. The Bunocephalus coracoideus species complex (Siluriformes, Aspredinidae): signs of a speciation process through chromosomal, genetic and ecological diversity. Front Genet. 2017; 8:120. https://doi.org/10.3389/fgene.2017.00120
Fricke R, Eschmeyer WN, Van der Laan R. Eschmeyer’s catalog of fishes: genera, species, references [Internet]. San Francisco: California Academy of Science; 2025. Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp
Fujisawa T, Barraclough TG. Delimiting species using single-locus data and the Generalized Mixed Yule Coalescent approach: a revised method and evaluation on simulated data sets. Syst Biol. 2013; 62(5):707–24. https://doi.org/10.1093/sysbio/syt033
Griffiths SP. The use of clove oil as an anaesthetic and method for sampling intertidal rockpool fishes. J Fish Biol. 2000; 57(6):1453–64. https://doi.org/10.1111/j.1095-8649.2000.tb02224.x
Haerter CAG, Margarido VP, Blanco DR, Traldi JB, Feldberg E, Lui RL. Contributions to Trachelyopterus (Siluriformes: Auchenipteridae) species diagnosis by cytotaxonomic autapomorphies: from U2 snRNA chromosome polymorphism to rDNA and histone gene synteny. Org Divers Evol. 2022; 22:747–58. https://doi.org/10.1007/s13127-022-00560-0
Haerter CAG, Viana PF, Takagui FHT, Tonello S, Margarido VP, Blanco DR et al. A variant W chromosome in Centromochlus heckelii (Siluriformes, Auchenipteridae) and the role of repeated DNA in its heteromorphism. Genet Mol Biol. 2025; 48:e20240071. https://doi.org/10.1590/1678-4685-GMB-2024-0071
Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser. 1999; 41:95–98.
Hatanaka T, Galetti PM. Mapping of the 18S and 5S ribosomal RNA genes in the fish Prochilodus argenteus Agassiz, 1829 (Characiformes, Prochilodontidae). Genetica. 2004; 122:239–44. https://doi.org/10.1007/s10709-004-2039-y
Hebert PD, Cywinska A, Ball SL, Dewaard JR. Biological identifications through DNA barcodes. Proc Biol Sci. 2003; 270(1512):313–21. https://doi.org/10.1098/rspb.2002.2218
Higuchi HA. Phylogeny of South American thorny catfishes (Osteichthyes: Siluriformes, Doradidae) [Master Dissertation]. Cambridge (MA): Harvard University; 1992.
Higuchi H, Birindelli JL, Sousa L, Britski H. Merodoras nheco, new genus and species from Rio Paraguay basin, Brazil (Siluriformes, Doradidae), and nomination of the new subfamily Astrodoradinae. Zootaxa. 2007; 1446:31–42. https://doi.org/10.5281/zenodo.176125
Hoorn C, Bogotá-A GR, Romero-Baez M, Lammertsma EI, Flantua SGA, Dantas EL et al. The Amazon at sea: onset and stages of the Amazon River from a marine record, with special reference to Neogene plant turnover in the drainage basin. Glob Planet Change. 2017; 153:51–65. https://doi.org/10.1016/j.gloplacha.2017.02.005
Hoorn C, Wesselingh FP, ter Steege H, Bermúdez MA, Mora A, Sevink J et al. Amazonia through time: Andean uplift, climate change, landscape evolution and biodiversity. Science. 2010; 330(6006):927–31. https://doi.org/10.1126/science.1194585
Ijdo JM, Wells RA, Baldini A, Reeders ST. Improved telomere detection using a telomere repeat probe (TTAGGG)n generated by PCR. Nucleic Acids Res. 1991; 19:4780.
Ivanyisky SJ, Albert JS. Systematics and biogeography of Sternarchellini (Gymnotiformes: Apteronotidae): diversification of electric fishes in large Amazonian rivers. Neotrop Ichthyol. 2014; 12(3):565–84. https://doi.org/10.1590/1982-0224-20130159
Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980; 16:111–20. https://doi.org/10.1007/BF01731581
Kowalski S, Haerter CAG, Perin DP, Takagui FHT, Viana PF, Feldberg E et al. Karyotypic characterization of Centromochlus schultzi Rössel 1962 (Auchenipteridae, Centromochlinae) from the Xingu River basin: new inferences on chromosomal evolution in Centromochlus. Genet Mol Biol. 2024; 47:e20230105. https://doi.org/10.1590/1678-4685-GMB-2023-0105
Levan A, Fredga K, Sandberg AA. Nomenclature for centromeric position on chromosomes. Hereditas. 1964; 52(2):201–22. https://doi.org/10.1111/j.1601-5223.1964.tb01953.x
Lin KW, Yan J. Endings in the middle: current knowledge of interstitial telomeric sequences. Mutat Res. 2008; 658:95–110. https://doi.org/10.1016/j.mrrev.2007.08.006
Lui RL, Blanco DR, Martinez JF, Margarido VP, Venere PC, Moreira-Filho O. The role of chromosomal fusion in the karyotypic evolution of the genus Ageneiosus (Siluriformes: Auchenipteridae). Neotrop Ichthyol. 2013; 11(2):327–34. https://doi.org/10.1590/S1679-62252013005000004
Lui RL, Blanco DR, Moreira-Filho O, Margarido VP. Propidium iodide for making heterochromatin more evident in the C-banding technique. Biotech Histochem. 2012; 87(7):433–38. https://doi.org/10.3109/10520295.2012.696700
Lundberg JG, Marshall LG, Guerrero J, Horton B, Malabarba MCSL, Wesselingh F. The stage for neotropical fish diversification: a history of tropical South America rivers. In: Malabarba LR, Reis RE, Vari RP, Lucena CAS, Lucena ZMS, editors. Phylogeny and classification of Neotropical fishes. Porto Alegre: Edipucrs; 1998. p.13–48.
Marajó L, Viana PF, Ferreira M, Py-Daniel LHR, Feldberg E. Cytogenetics of two Farlowella species (Loricariidae: Loricariinae): implications on the taxonomic status of the species. Neotrop Ichthyol. 2018; 16(4):e180029. https://doi.org/10.1590/1982-0224-20180029
Martins C, Galetti Jr. PM. Chromosomal localization of 5S rDNA genes in Leporinus fish (Anostomidae, Characiformes). Chromosome Res. 1999; 7:363–67. https://doi.org/10.1023/A:1009216030316
Miller MA, Pfeiffer W, Schwartz T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees [Internet]. 2010. Available from: https://www.phylo.org/sub_sections/portal/sc2010_paper.pdf
Milhomem SSR, Souza ACP, Nascimento AL, Carvalho Jr. JR, Feldberg E, Pieczarka JC et al. Cytogenetic studies in fishes of the genera Hassar, Platydoras and Opsodoras (Doradidae, Siluriformes) from Jarí and Xingú rivers, Brazil. Genet Mol Biol. 2008; 31:256–60. https://doi.org/10.1590/S1415-47572008000200017
Moreira-Filho O, Bertollo LAC. Astyanax scabripinnis (Pisces, Characidae): a species complex. Braz J Genet. 1991; 14:331–57.
Moyer G, Burr BM, Krajewski C. Phylogenetic relationships of thorny catfishes (Siluriformes: Doradidae) inferred from molecular and morphological data. Zool J Linn Soc. 2004; 140(4):551–75. https://doi.org/10.1111/j.1096-3642.2004.00114.x
Murnane JP. Telomeres and chromosome instability. DNA Repair (Amst). 2006; 5(9–10):1082–92. https://doi.org/10.1016/j.dnarep.2006.05.030
Ocalewicz K. Telomeres in fishes. Cytogenet Genome Res. 2013; 141(2–3):114–25. https://doi.org/10.1159/000354278
Paiz LM, Baumgärtner L, Graça WJ, Margarido VP. Basic cytogenetics and physical mapping of ribosomal genes in four Astyanax species (Characiformes, Characidae) collected in Middle Paraná River, Iguassu National Park: considerations on taxonomy and systematics of the genus. Comp Cytogenet. 2015; 9(1):51–65. https://doi.org/10.3897/CompCytogen.v9i1.9002
Pereira LHG, Hanner R, Foresti F, Oliveira C. Can DNA barcoding accurately discriminate megadiverse Neotropical freshwater fish fauna? BMC Genetics. 2013; 14:20. https://doi.org/10.1186/1471-2156-14-20
Pereira LHG, Maia GMG, Hanner R, Foresti F, Oliveira C. DNA barcodes discriminate freshwater fishes from the Paraíba do Sul River Basin, São Paulo, Brazil. Mitochondrial DNA. 2011; 22:71–79. https://doi.org/10.3109/19401736.2010.532213
Piscor D, Fernandes CA, Parise-Maltempi PP. Conserved number of U2 snDNA sites in Piabina argentea, Piabarchus stramineus and two Bryconamericus species (Characidae, Stevardiinae). Neotrop Ichthyol. 2018; 16(1):e170066. https://doi.org/10.1590/1982-0224-20170066
Pons J, Barraclough TG, Gomez-Zurita J, Cardoso A, Duran DP, Hazell S et al. Sequence-based species delimitation for the DNA taxonomy of underscribed insects. Syst Biol. 2006; 55(4):595–609. https://doi.org/10.1080/10635150600852011
Prizon AC, Bruschi DP, Borin-Carvalho LA, Cius A, Barbosa LM, Ruiz HB et al. Hidden diversity in the populations of the armored catfish Ancistrus Kner, 1854 (Loricariidae, Hypostominae) from the Paraná River basin revealed by molecular and cytogenetic data. Front Genet. 2017; 8:185. https://doi.org/10.3389/fgene.2017.00185
Ratnasingham S, Hebert PDA. DNA-based registry for all animal species: the Barcode Index Number (BIN) system. PLoS ONE. 2013; 8(7):e66213. https://doi.org/10.1371/journal.pone.0066213
Roa-Fuentes CA, Alonso JC, Alfonso A, Sabaj Pérez M. Pisces, Siluriformes, Doradidae, Astrodoras Bleeker, 1862: first record in the Colombian Amazon. Check List. 2010; 6(4):485–87. https://doi.org/10.15560/6.4.485
Ronquist F, Huelsenbeck JP. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003; 19(12):1572–74. https://doi.org/10.1093/bioinformatics/btg180
Ruiz-Herrera A, Nergadze SG, Santagostino M, Giulotto E. Telomeric repeats far from the ends: mechanisms of origin and role in evolution. Cytogenet Genome Res. 2008; 122(3–4):219–28. https://doi.org/10.1159/000167807
Sabaj MH, Arce HM. Towards a complete classification of the Neotropical thorny catfishes (Siluriformes: Doradidae). Neotrop Ichthyol. 2021; 19(4):e210064. https://doi.org/10.1590/1982-0224-2021-0064
Santos DP, Felicetti D, Baumgärtner L, Margarido VP, Blanco DR, Moreira-Filho O et al. Contributions to the taxonomy of Trachelyopterus (Siluriformes): comparative cytogenetic analysis in three species of Auchenipteridae. Neotrop Ichthyol. 2021; 19(1):e200115. https://doi.org/10.1590/1982-0224-2020-0115
Scacchetti PC, Pansonato-Alves JC, Utsunomia R, Oliveira C, Foresti F. Karyotypic diversity in four species of the genus Gymnotus Linnaeus, 1758 (Teleostei, Gymnotiformes, Gymnotidae): physical mapping of ribosomal genes and telomeric sequences. Comp Cytogenet. 2011; 5(3):223–35. https://doi.org/10.3897/CompCytogen.v5i3.1375
Scacchetti PC, Utsunomia R, Pansonato-Alves JC, Vicari MR, Artoni RF, Oliveira C et al. Chromosomal mapping of repetitive DNAs in Characidium (Teleostei, Characiformes): genomic organization and diversification of ZW sex chromosomes. Cytogenet Genome Res. 2015; 146(2):136–43. https://doi.org/10.1159/000437165
Schmid M, Steinlein C. Chromosome banding in Amphibia. XXXIV. Intrachromosomal telomeric DNA sequences in Anura. Cytogenet Genome Res. 2016; 148(2–3):211–26. https://doi.org/10.1159/000446298
Slijepcevic P. Telomeres and mechanisms of Robertsonian fusion. Chromosoma. 1998; 107:136–40. https://doi.org/10.1007/s004120050289
Sousa LM. Revisão taxonômica e filogenia de Astrodoradinae (Siluriformes, Doradidae). [PhD Thesis]. São Paulo: Universidade de São Paulo; 2010.
Sousa LM, Birindelli JLO. Taxonomic revision of the genus Scorpiodoras (Siluriformes: Doradidae) with resurrection of Scorpiodoras calderonensis and description of a new species. Copeia. 2011; 2011(1):121–40. https://doi.org/10.1643/CI-10-038
Sousa LM, Rapp Py-Daniel LH. Description of two new species of Physopyxis and redescription of P. lyra (Siluriformes: Doradidae). Neotrop Ichthyol. 2005; 3(4):625–36. https://doi.org/10.1590/S1679-62252005000400019
Souza-Shibatta L, Pezenti LF, Ferreira DG, Almeida FS, Sofia SH, Shibatta AO. Cryptic species of the genus Pimelodella (Siluriformes: Heptapteridae) from the Miranda River, Paraguay River basin, Pantanal of Mato Grosso do Sul, Central Brazil. Neotrop Ichthyol. 2013; 11(1):101–19. https://doi.org/10.1590/S1679-62252013000100012
Souza-Shibatta L, Tonini JFR, Abrahão VP, Jarduli LR, Oliveira C, Malabarba LR et al. Reappraisal of the systematics of Microglanis cottoides (Siluriformes, Pseudopimelodidae), a catfish from southern Brazil. PLoS ONE. 2018; 13(7):e0199963. https://doi.org/10.1371/journal.pone.0199963
Stamatakis A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 2014; 30(9):1312–13. https://doi.org/10.1093/bioinformatics/btu033
Sumner AMT. A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res. 1972; 75:304–06.
Takagui FH, Baumgärtner L, Baldissera JN, Lui RL, Margarido VP, Fonteles SBA et al. Chromosomal diversity of thorny catfishes (Siluriformes-Doradidae): a case of allopatric speciation among Wertheimerinae species of São Francisco and Brazilian eastern coastal drainages. Zebrafish. 2019; 16:477–85. https://doi.org/10.1089/zeb.2019.1769
Takagui FH, Baumgärtner L, Venturelli NB, Paiz LM, Viana P, Dionísio JF et al. Unraveling the karyotypic evolution and cytotaxonomy of armored catfishes (Loricariinae) with emphasis in Sturisoma, Loricariichthys, Loricaria, Proloricaria, Pyxiloricaria and Rineloricaria. Zebrafish. 2020; 17:319–32. https://doi.org/10.1089/zeb.2020.1893
Takagui FH, Baumgärtner L, Viana P, Lima MCC, Bitencourt JA, Venere PC et al. Karyotype evolution of talking thorny catfishes Anadoras (Doradidae, Astrodoradinae): a process mediated by structural rearrangements and intense reorganization of repetitive DNAs. Cytogenet Genome Res. 2022; 162(1–2):64–75. https://doi.org/10.1159/000523747
Takagui FH, Dias AL, Birindelli JLO, Swarça AC, Rosa R, Lui RL et al. First report of B chromosomes in three neotropical thorny catfishes (Siluriformes, Doradidae). Comp Cytogenet. 2017a; 11(1):55–64. https://doi.org/10.3897/CompCytogen.v11i1.10496
Takagui FH, Moura LF, Ferreira DC, Centofante L, Vitorino CA, Bueno V et al. Karyotype diversity in Doradidae (Siluriformes, Doradoidae) and presence of the heteromorphic ZZ/ZW sex chromosome system in the family. Zebrafish. 2017b; 14:236–43. https://doi.org/10.1089/zeb.2016.1368
Takagui FH, Viana P, Baumgärtner L, Bitencourt JA, Margarido VP, Lui RL et al. Reconstruction of the Doradinae (Siluriformes-Doradidae) ancestral diploid number and NOR pattern, reveals new insights about the karyotypic diversification of the neotropical thorny catfishes. Genet Mol Biol. 2021; 44(4):e20200068. https://doi.org/10.1590/1678-4685-GMB-2020-0068
Takagui FH, Viana P, Haerter CAG, Zuanon J, Birindelli JLO, Lui RL et al. Chromosomal analysis of two Acanthodoras species (Doradidae, Siluriformes): insights into the oldest thorny catfish clade and its karyotype evolution. J Fish Biol. 2024; 105(4):1109–19 https://doi.org/10.1111/jfb.15863
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol. 2011; 28(10):2731–39. https://doi.org/10.1093/molbev/msr121
Tenório RCCO, Vitorino CA, Souza IL, Oliveira C, Venere PC. Comparative cytogenetics in Astyanax (Characiformes: Characidae) with focus on the cytotaxonomy of the group. Neotrop Ichthyol. 2013; 11(3):553–64. https://doi.org/10.1590/S1679-62252013000300008
Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994; 22(22):4673–80. https://doi.org/10.1093/nar/22.22.4673
Vicari MR, Bruschi DP, Cabral-de-Mello DC, Nogaroto V. Telomere organization and the interstitial telomeric sites involvement in insects and vertebrates chromosome evolution. Genet Mol Biol. 2022; 45:e20220071. https://doi.org/10.1590/1678-4685-GMB-2022-0071
Viana PF, Feldberg E, Cioffi MB, Carvalho VT, Menezes S, Vogt RC et al. The Amazonian red side-necked turtle Rhinemys rufipes (Spix, 1824) (Testudines, Chelidae) has a GSD sex-determining mechanism with an ancient XY sex microchromosome system. Cells. 2020; 9(9):2088. https://doi.org/10.3390/cells9092088
Ward RD, Zemlak TS, Innes BH, Last PR, Hebert PD. DNA barcoding Australia’s fish species. Philos Trans R Soc Lond B Biol Sci. 2005; 360:1847–57. https://doi.org/10.1098/rstb.2005.1716
Yano CF, Bertollo LAC, Cioffi MB. Fish-FISH: molecular cytogenetics in 684 fish species. In: Liehr T, editor. Fluorescence in situ hybridization (FISH) – application guide. 2nd ed. Berlin: Springer; 2017a. p.429–44.
Yano CF, Bertollo LAC, Rebordinos L, Merlo MA, Liehr T, Portela-Bens S et al. Evolutionary dynamics of rDNAs and U2 small nuclear DNAs in Triportheus (Characiformes, Triportheidae): high variability and particular syntenic organization. Zebrafish. 2017b; 14:146–54. https://doi.org/10.1089/zeb.2016.1351
Zhang J, Kapli P, Pavlidis P, Stamatakis A. A general species delimitation method with applications to phylogenetic placements. Bioinformatics. 2013; 29:2869–76. https://doi.org/10.1093/bioinformatics/btt499
Wesselingh FP, Hoorn C. Amazonia: landscape and species evolution: a look into the past. Oxford: Blackwell Publishing Ltd; 2009. https://doi.org/10.1002/9781444306408
Authors
![]() Fábio Hiroshi Takagui1,
Fábio Hiroshi Takagui1, ![]() Patrik Viana1,
Patrik Viana1, ![]() Chrystian Aparecido Grillo Haerter1,
Chrystian Aparecido Grillo Haerter1, ![]() Jamille de Araújo Bitencourt2
Jamille de Araújo Bitencourt2 ![]() ,
, ![]() José Luís Olivan Birindelli3,
José Luís Olivan Birindelli3, ![]() Roberto Laridondo Lui4,
Roberto Laridondo Lui4, ![]() Vladimir Pavan Margarido4,
Vladimir Pavan Margarido4, ![]() Fernanda Simões Almeida5,
Fernanda Simões Almeida5, ![]() Lucia Giuliano-Caetano6,
Lucia Giuliano-Caetano6, ![]() Eliana Feldberg1 and
Eliana Feldberg1 and ![]() Jorge Ivan Rebelo Porto1
Jorge Ivan Rebelo Porto1
[1] Laboratório de Genética Animal, Instituto Nacional de Pesquisas da Amazônia, André Araújo, 2936, 69060-001, Manaus, AM, Brazil. (FHT) fabiotakagui@hotmail.com, (PFV) patrik.biologia@gmail.com, (CAGH) chrystianhaerter@gmail.com, (EF) feldberg@inpa.gov.br, (JIRP) jorge.porto@inpa.gov.br.
[2] Laboratório de Citogenética, Departamento de Ciências Biológicas, Universidade Estadual do Sudoeste da Bahia, Avenida Jose Moreira Sobrinho, s/n, 42205-490, Jequié, BA, Brazil. (JAB) millebitencourt@gmail.com.
[3] Museu de Zoologia, Departamento de Biologia Animal e Vegetal, Universidade Estadual de Londrina, Rodovia Celso Garcia Cid, PR-445, km 380, 86057-970, Londrina, PR, Brazil. (JLOB) josebirindelli@uel.br.
[4] Laboratório de Citogenética e Genética de Peixes, Centro de Ciências Biológicas, Universidade Estadual do Oeste do Paraná, Campus Cascavel, Rua Universitária, 1619, 85819-110, Cascavel, PR, Brazil. (RLL) roberto.lui@unioeste.br (corresponding author), (VPM) vladimir.margarido@unioeste.br.
[5] Laboratório de Genética Animal e Ecologia, Departamento de Biologia Geral, Universidade Estadual de Londrina, Rodovia Celso Garcia Cid, PR-445, km 380, 86057-970, Londrina, PR, Brazil. (FSA) fernandasa@uel.br.
[6] Laboratório de Citogenética Animal, Departamento de Biologia Geral, Universidade Estadual de Londrina, Rodovia Celso Garcia Cid, PR-445, km 380, 86057-970, Londrina, PR, Brazil. (LGC) luciagiulianocaetano@gmail.com.
Authors’ Contribution 

Fábio Hiroshi Takagui: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Writing-original draft, Writing-review and editing.
Patrik Viana: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Validation, Visualization, Writing-review and editing.
Chrystian Aparecido Grillo Haerter: Formal analysis, Investigation, Methodology, Validation, Visualization, Writing-review and editing.
Jamille de Araújo Bitencourt: Data curation, Formal analysis, Investigation, Methodology, Validation, Visualization, Writing-review and editing.
José Luís Olivan Birindelli: Data curation, Formal analysis, Investigation, Methodology, Validation, Visualization, Writing-review and editing.
Roberto Laridondo Lui: Data curation, Funding acquisition, Investigation, Resources, Validation, Visualization, Writing-review and editing.
Vladimir Pavan Margarido: Data curation, Funding acquisition, Investigation, Resources, Supervision, Validation, Visualization, Writing-review and editing.
Fernanda Simões Almeida: Data curation, Investigation, Validation, Visualization, Writing-review and editing.
Lucia Giuliano-Caetano: Data curation, Funding acquisition, Investigation, Resources, Supervision, Validation, Visualization, Writing-review and editing.
Eliana Feldberg: Data curation, Funding acquisition, Investigation, Resources, Supervision, Validation, Visualization, Writing-review and editing.
Jorge Ivan Rebelo Porto: Conceptualization, Data curation, Funding acquisition, Investigation, Project administration, Resources, Supervision, Validation, Visualization, Writing-review and editing.
Ethical Statement
The experiments followed ethical conducts according to the Ethics Committee for Animal Use of the Universidade Estadual de Londrina, under the protocol number 60/2017. The specimens were collected under the authorization of the Instituto Chico Mendes de Conservação da Biodiversidade (ICMBio; permit number 85306–1).
Competing Interests
The author declares no competing interests.
Data availability statement
The cytogenetic data that support this study are partially included in the article. Unfortunately, the raw data with full metaphase plates hybridized with 5S rDNA, 18S rDNA, and U2 snDNA probes are no longer accessible. Further details can be requested from the corresponding author. The genetic datasets are deposited in the BOLD system and listed in the supplementary material.
AI statement
The authors declare that no artificial intelligence was used in this study.
Funding
This study was financially supported by the Fundação de Amparo à Pesquisa do Estado do Amazonas (FAPEAM) through a postdoctoral scholarship granted to Fábio H. Takagui (PRODOC/FAPEAM, nº 003/2022); by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES); the Fundação Araucária (FA); and the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), through a research productivity grant (process nº 302872/2018–3). The fish collection expeditions were supported by resources from the Postgraduate Program in Genetics and Molecular Biology at Universidade Estadual de Londrina (UEL) and the Instituto Nacional de Pesquisas da Amazônia (INPA).
Supplementary Material
Supplementary material S1
Supplementary material S2
Supplementary material S3
How to cite this article
Takagui FH, Viana P, Haerter CAG, Bitencourt JA, Birindelli JLO, Lui RL, Margarido VP, Almeida FS, Giuliano-Caetano L, Feldberg E, Porto JIR. Cytogenetic and COI-based analyses support a polytypic status of Astrodoras (Doradidae: Siluriformes). Neotrop Ichthyol. 2026; 24(1):e250120. https://doi.org/10.1590/1982-0224-2025-0120
Copyright
This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Distributed under
Creative Commons CC-BY 4.0

© 2025 The Authors.
Diversity and Distributions Published by SBI
![]() Accepted October 1, 2025
Accepted October 1, 2025
![]() Submitted July 7, 2025
Submitted July 7, 2025
![]() Epub March 6, 2026
Epub March 6, 2026