![]() Lígia Carla Balini1
Lígia Carla Balini1 ![]() ,
, ![]() Rafael Fernando de Melo1,
Rafael Fernando de Melo1, ![]() Ana Luiza de Brito Portela-Castro1,2,
Ana Luiza de Brito Portela-Castro1,2, ![]() Cláudio Henrique Zawadzki2,3,
Cláudio Henrique Zawadzki2,3, ![]() Marcos Otávio Ribeiro4,
Marcos Otávio Ribeiro4, ![]() Carlos Alexandre Fernandes1,2 and
Carlos Alexandre Fernandes1,2 and ![]() Luciana Andreia Borin-Carvalho1
Luciana Andreia Borin-Carvalho1
PDF: EN XML: EN | Cite this article
Abstract
Hypostomus é um grupo notável dentro dos Siluriformes pelo grande número de espécie, alta variabilidade morfológica e cromossômica, o que dificulta o processo de identificação de suas espécies. Nesse contexto, o objetivo deste estudo foi identificar possíveis marcadores cromossômicos interpopulacionais que poderiam contribuir significativamente para nossa compreensão da diferenciação em H. strigaticeps de tributários distintos. Duas populações de H. strigaticeps da bacia do alto rio Paraná foram analisadas usando métodos citogenéticos. Nossos resultados revelaram variações interpopulacionais em H. strigaticeps, apesar da amostragem limitada da população do Rio Ivaí, particularmente no número de sítios de DNAr 18S e 5S. Além disso, os indivíduos de ambas as populações exibiram a mesma contagem de cromossomos (2n = 72) e número fundamental (FN = 108), bem como morfologia cromossômica idêntica e similaridades na distribuição da heterocromatina constitutiva. Os elementos transponíveis(TEs) Rex1e Rex3foram encontrados espalhados ao longo dos cromossomos, com uma concentração maior de elementos Rex3 em pares carregando DNAr.Nossas descobertas fornecem importantes novos insights sobre um possível envolvimento de TEs na variabilidade dos padrões de distribuição de DNAr, contribuindo para a diversificação cariotípica em H. strigaticeps.
Palavras-chave: Elementos transponíveis, Evolução do cariótipo, Heterocromatina, NORs, Retrotransposons.
Introduction
Hypostomus Lacépède, 1803 is a highly diverse genus of Loricariidae, with 161 valid species (Fricke et al., 2024). This genus presents a remarkable morphological variability, featuring fish that range in size from small to large, popularly known as plecos or “cascudos” (Weber, 2003; Zawadzki et al., 2014; Zanata, Pitanga, 2016). The intraspecific diversity in morphology and color pattern of Hypostomus species makes taxonomic identification challenging, necessitating constant taxonomic reviews (Weber, 2003).
As a result, several cytogenetic studies have used the terms sp., aff., and cf., to describe species within Hypostomus, leading to an estimate of around 40 species with diploid numbers (2n) described (Rubert et al., 2022 in supplementary table). These species exhibit a wide variation in 2n from 64 chromosomes in H. faveolus Zawadzki, Birindelli & Lima, 2008 from Taquaralzinho River, upper Araguaia River basin, H. cochliodon Kner, 1854 from Iguassu River, Paraná River basin (Bueno et al., 2013),and H. soniae Hollanda Carvalho & Weber, 2005from Teles Pires River, Southern Amazon basin (Oliveira et al., 2019) to 84 chromosomes in H. perdido Zawadzki, Tencatt & Froehlich, 2014 from Perdido River, Paraguay River basin (Cereali et al., 2008; Zawadzki et al., 2014). The karyotypic diversity of the genus has demonstrated not only variability in 2n but also intra- and interspecific structural chromosomal diversity. This suggests that chromosomal rearrangements, mainly centric fissions, and pericentric inversions, were the possible mechanisms involved in the diversification of this group (Artoni, Bertollo, 1996; Artoni, Bertollo, 2001; Alves et al., 2005; Bueno et al., 2012).
Among the diverse diploid numbers in Hypostomus, Hypostomus strigaticeps (Regan, 1908) stands out for its conserved 2n, with 72 chromosomes, in various populations (Fernandes et al., 2012; Bueno et al., 2013; Baumgärtner et al., 2014; Rubert et al., 2016). This species, initially thought to be unique in its distribution confined to the Tiete River basin (SP) (Weber, 2003), was described in 1908 (Regan, 1908) from the Piracicaba River. Contrary to initial speculations, Borba et al. (2013) suggested based on their phylogenetic analysis of different populations of H. strigaticeps from the Paraná River basin using ATP synthase sequences (mitochondrial DNA) that they all belong to the same species, rejecting any claim for cryptic species. Additionally, their phylogeographic analysis concluded that the species distribution extends beyond the Tiete River basin to other subbasins of the upper Paraná River (Borba et al., 2013).
In Hypostomus, cytogenetic variations are not restricted to diploid numbers and karyotype composition. Interspecific, intraspecific and interpopulation variation in the number of heterochromatic bands and ribosomal DNA (rDNA) sites [which include major (45S) and minor (5S) rDNA clusters] is a common cytogenetic feature in Hypostomus (Baumgärtner et al., 2014; Bueno et al., 2014). These events within this genus highlight the pressing need for further investigation to gain a more comprehensive understanding of these polymorphisms.
Transposable elements (TEs) are categorized as retrotransposons (Class I) or DNA transposons (Class II) based on their transposition mechanism, which involves either an RNA intermediate or DNA, respectively (Wicker et al., 2007). The teleosts have the highest diversity of TEs in vertebrates (Chalopin et al., 2015; Shao et al., 2019; Carducci et al., 2020), and TEs have been used as chromosomal markers in fish cytogenetics (Ferreira et al., 2011), providing valuable insights into the dispersions of specific DNA fragments along chromosomes. According to Volff et al. (1999, 2000, 2001), the teleost genomes contain abundant retroelements Rex1, Rex3, and Rex6 and others transposable elements (Ferreira et al., 2011; Glugoski et al., 2018). In a recent study, Traldi et al. (2019) revealed a wide dispersion of Rex1 and Rex3 retroelements in both heterochromatic and euchromatic regions in species of Hypostomus.
Given that H. strigaticeps is a resident of the upper Paraná River basin, the present article compared two populations collected from the Pirapó and Ivaí rivers through cytogenetics analysis. The aim was to identify possible interpopulational chromosomal markers that could significantly contribute to our understanding of the differentiation in H. strigaticeps of distinct tributaries.
Material and methods
Study area and sampling. Fourteen individuals of H. strigaticeps were collected from populations of the Paraná River basin:seven males and four females from the Pirapó River (municipality of Maringá, Paraná State; 23º18’15” S 51º53’41”W), and two males and one female from the Ivaí River (municipality of Ivatuba, Paraná State; 23º38’17”S 52º15’22”W).
Voucher specimens were deposited in the Fish Collection of the Núcleo de Pesquisas em Limnologia, Ictiologia e Aquicultura (NUPELIA), Universidade Estadual de Maringá, municipality of Maringá, Paraná State, Brazil, as Hypostomus strigaticeps from the Pirapó River (NUP 22964), and the Ivaí River (NUP 22970).
Cytogenetic analysis. Metaphase chromosomes were obtained from anterior kidney cells using the air-drying technique (Bertollo et al., 2015). The nucleolus organizer regions (NORs) were detected employing silver nitrate staining (Howell, Black, 1980). Constitutive heterochromatin was determined following the C-banding technique (Sumner, 1972) and stained with propidium iodide (Lui et al., 2012).
At least 30 metaphases were analyzed for each individual, and those with better chromosome morphology were used for the karyotype analysis. The chromosomes were classified as metacentric (m), submetacentric (sm), subtelocentric (st), and acrocentric (a) according to (Levan et al., 1964). The fundamental number (FN) was calculated according to the chromosomal arm numbers (the chromosomes m, sm, and st were considered to contain two arms – and the chromosomes with one arm).
The location of the rDNA sites in the chromosomes was performed by fluorescence in situ hybridization (FISH) with modifications (Margarido, Moreira-Filho, 2008; Pinkel et al., 1986) using probes from the genome of Megaleporinus elongatus (Valenciennes, 1850) (Martins, Galetti Jr., 1999) for 5S rDNA sites and Prochilodus argenteus Spix & Agassiz, 1829 (Hatanaka, Galetti, 2004) for 18S rDNA sites. The probes were labeled through nick translation with digoxigenin-11-dUTP (5S rDNA) and biotin-16-dUTP (18S rDNA) (Roche Diagnostics, Mannheim, Germany). Detection and amplification of the hybridization signal were carried out using avidin-FITC and anti-avidin biotin (Sigma- Aldrich, St. Louis, Missouri, United States) for probes labeled with biotin and anti-digoxigenin rhodamine (Roche Diagnostics, Mannheim, Germany) for probes labeled with digoxigenin. Chromosomes were counterstained with 4’-6-diamin-2-phenylindole (DAPI, 50 ìg ml−1, Sigma- Aldrich, St. Louis, Missouri, United States).
Transposable element probes were produced using the primers Rex-3 [Foward (5’- CGGTGAYAAAGGGCAGCCCTG-3’) and Reverse (5’-TGGCAGACNGGGGTGGTGGT-3’) (Volff, 2006). These nucleotide sequences of the Rex-3 element, obtained from the genome of different Ancistrus populations, revealed a high degree of similarity to the non-LTR Rex-3 retrotransposon, according to sequences deposited in the online BLAST database and confirmed by the CENSOR (Prizon et al., 2018). REX-1 [Foward (5’- TTCTCCAGTGCCTTCAACACC-3’) and Reverse (5’ – TCCCTCAGCAGAAAGAGTCTGCTC-3’) (Volff et al., 2000). Amplification was performed using PCR, and the probes were labeled according to the nick translation method using the Anti-digoxigenin-Rhodamine Kit (Roche Diagnostics, Mannheim, Germany). Chromosomes were counterstained with 4’-6-diamin-2-phenylindole (DAPI, 50 ìg ml−1, Sigma- Aldrich, St. Louis, Missouri, United States).
Admittedly, the obtained PCR fragments were not sequenced. Nonetheless, although the probes obtained by PCR may occasionally include nonspecific fragments, the primers used (Volff et al., 1999, 2000) are recognized for their high specificity in the amplification of Rex1 and Rex3 elements. The identity of the fragments amplified by these primers has been widely confirmed by sequencing, as demonstrated in several studies (Ferreira et al., 2011; Prizon et al., 2018; among others), including in Hypostomus species (Traldi et al., 2019). Additionally, cross-hybridizations with Rex1 and Rex3 probes between different species of Loricariidae showed distribution patterns similar to those observed with species-specific probes, indicating a high nucleotide identity between the species (Ferreira et al., 2011).
Conventional and fluorescence chromosome preparations were analysed under a BX 51 epifluorescence microscope (Olympus America, Inc). The images were captured using the DP controller (Media Cybernetics) software and the image composition with Adobe Photoshop CS6.
Results
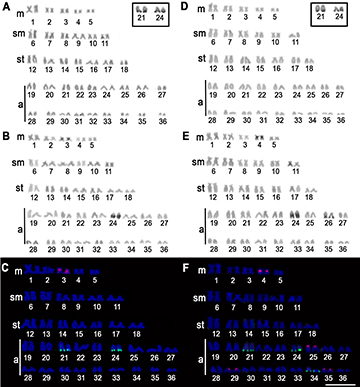
All the 14 individuals of H. strigaticeps from the two collection sites had invariably diploid chromosome number of 72 chromosomes, with the karyotype composed of 10 metacentric, 12 submetacentric, 14 subtelocentric, and 36 acrocentric chromosomes and FN value 108 in both sexes (Figs. 1A, D). Heteromorphic sex chromosomes were not identified.
The constitutive heterochromatin was located in the pericentromeric region of pairs 3 for Pirapó River population (Fig. 1B) and 4 for Ivaí River population (Fig. 2E), both populations showed large heterochromatic blocks located in terminal position in the long arm of the acrocentric pair 24, and in the same position of the acrocentric pairs 28 for Pirapó River population (Fig. 1B) and 26 for Ivaí River population (Fig. 2E).
FIGURE 1| Karyotypes of Hypostomus strigaticeps from Pirapó River (A, B, C) and Ivaí River (D, E, F). Karyotypes (A and D) stained with Giemsa, (B and E) C-banded, and after double FISH (C and F) with 5S rDNA probes (red) and 18S rDNA (green). The AgNOR-bearing chromosomes are boxed. Scale bar = 10 µm.
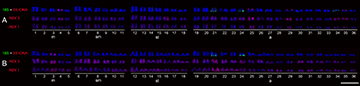
FIGURE 2| Karyotypes of Hypostomus strigaticeps after FISH with rDNA and retroelements Rex1 and Rex3 from Ivaí River (A) and Pirapó River (B). Scale bar = 10 µm.
The silver nitrate impregnation identified the Ag-NOR in a terminal position of the long arm of the acrocentric pairs 21 and 24 in both populations (Fig. 1, Box). FISH with 18S rDNA probes confirmed the Ag-NOR sites (Figs. 1C, F) and detected further inactive major ribosomal clusters in a terminal position of the long arm of the acrocentric pair 34 for Ivaí River population (Fig. 1F). Moreover, the FISH revealed a single 5S rDNA site in the pericentromeric position of pair 3 for the Pirapó River population. Multiple 5S rDNA sites were observed in the pericentromeric position of pair 4 and the terminal position of the short arm of the acrocentric pairs 25, 29, and 35 in the Ivaí River population (Fig. 1F). None of the 5S rDNA sites co-localized with major rDNA (45S) clusters which have been visualized using the probe for 18S rDNA.
In FISH experiments with transposable elements, we found that the Rex1 and Rex3 elements are scattered along the chromosomes, including both euchromatic and heterochromatic regions, in all analyzed populations (Fig. 2). This dispersed pattern of Rex1 and Rex3 elements observed in our study is widely described in the literature (including with highly specific probes obtained by cloning and sequencing). Notably, in H. strigaticeps from the Ivaí River, there was a higher concentration of Rex3 elements in pairs 25, 29, 34, and 35, as well as in one of the homologs of pair 16 (Fig. 2A).
Discussion
Our results revealed interpopulation variations in H. strigaticeps, despite limited sampling for the Ivaí River population, particularly in the number of 18S and 5S rDNA sites. Notably, individuals from both populations share the same 2n = 72, FN = 108, identical chromosome morphology, and similarities in the distribution of constitutive heterochromatin. This diploid number of 72 chromosomes is predominant in most populations of H. strigaticeps, with the exceptions of populations from Corumbataí River (Alves et al., 2012) and the samples from unspecified locality examined by Michele et al. (1977), which exhibit a diploid number of 2n = 74 (Tab. 1). Considering 2n = 54 chromosomes as basal for Loricariidae (Artoni, Bertollo, 2001), centric fission is likely an important mechanism that contributed to the increase in 2n as observed for H. strigaticeps (2n = 72 and 74). Additionally, various populations of H. strigaticeps with 2n = 72 exhibit diversification in their karyotype structures (Tab. 1). This suggests chromosome rearrangements, such as pericentric inversions, addition and/or deletions that can alter the chromosome morphology without changing 2n. The geographical isolation of H. strigaticeps populations might have played a fundamental role in the establishment of this karyotype variation. The restriction of gene flow between them, as a result of their isolation, could have favored the fixation of such chromosome change. The majority of Hypostomus species, including H. strigaticeps, are typically formed by small groups of non-migratory animals (Britski et al., 1988), suggesting a strong effect of genetic drift.
TABLE 1 | Cytogenetic data available in Hypostomus strigaticeps from the upper Paraná River basin. ***Various points in the Paranapanema River basin, PR; 2n = diploid number; m = metacentric; sm = submetacentric; st = subtelocentric; a = acrocentric; NORs = nucleolar organizer regions (number of bearing chromosomes utilizing of Ag-NOR and/or 18S rDNA-FISH); 5S = 5S rDNA cistrons number.
Localities | 2n | Karyotype structure | NORs | 5S | References |
– | 74 | 8m+4sm+62st-a | – | – | Michelle et al. (1977) |
*** | 72 | 10m+16sm+46st-a | 4 | – | Rubert et al. (2011) |
Água Boa stream, MS | 72 | 12m+18sm+20st+22a | 4 | – | Fernandes et al. (2012) |
Corumbataí River, SP | 74 | 10m+14sm+14st+36a | 8 | – | Alves et al. (2012) |
Ivaí River, PR | 72 | 10m+14sm+18st+30a | 8 | – | Endo et al. (2012) |
Piquiri River, PR | 72 | 12m+12sm+18st+30a | 3 | – | Bueno et al. (2013) |
Hortelã stream, SP | 72 | 10m+18sm+18st+26a | 4 | 12 | Pansonato-Alves et al. (2013) |
Atlântico stream, PR Piquiri River, PR | 72 | 12m+12sm+18st+30a |
5 6 |
6 | Baumgärtner et al. (2014) |
Paraná River, PR Iguaçu River, PR | |||||
Mogi-Guaçu River, SP | 72 | 10m+16sm+46st-a | 2 | – | Rubert et al. (2016) |
Pirapó River, PR Ivaí River, PR | 72 | 10m+12sm+14st+36a | 4 6 | 2 8 | Present study |
In our study, the constitutive heterochromatin distribution did not differ significantly between the two populations analyzed. Both populations exhibited pericentromeric markings on a pair of metacentric chromosomes (pair 3 from the Pirapó River and pair 4 from the Ivaí River), coincident with 5S rDNA sites, as well as terminal markings on the long arm of two pairs of acrocentric chromosomes (pairs 24 and 28 from the Pirapó River and pairs 24 and 26 from the Ivaí River). Thus, the C-banding method did not provide any population-specific patterns of heterochromatic segments distribution. Nonetheless, large heterochromatic blocks on the long arm of acrocentric chromosomes identified intra and interpopulation polymorphism in other populations of H. strigaticeps (Baumgärtner et al., 2014). In other Hypostomus species, studied revealed the existence of a conserved pattern of distribution of constitutive heterochromatin. Notably, a pair of metacentric chromosomes exhibits pericentromeric markings (Rubert et al., 2011; Traldi et al., 2012). Additionally, some st-a chromosomes are characterized by conspicuous blocks located in the terminal regions of the long arms, which may account for approximately one-third to one-half of the chromosomes total lengths (Artoni, Bertollo 1999; Kavalco et al., 2004; Cereali et al., 2008; Baumgartner et al., 2014). Both forms described for these chromosomes are similar to those found in H. strigaticeps from the populations analyzed.
Although both populations of H. strigaticeps presented the same karyotypic structure and similarities in the pattern of C-bands, they differed in the number of 18S and 5S rDNA sites. The application of 18S rDNA-FISH confirmed the Ag-NORs sites, and detected further inactive major ribosomal clusters for Ivaí River population. Multiple NORs are common in H. strigaticeps (Tab. 1), except for the individuals of the Mogi-Guaçu River (Rubert et al., 2016). Furthermore, the locations of NORs in acrocentric chromosomes in H. strigaticeps coincide with observations in other Hypostomus species and generally correspond to regions of constitutive heterochromatin (Kavalco et al., 2004; Rubert et al., 2008; Traldi et al., 2013; Baumgartner et al., 2014), such as the pair 24 in both populations in the present study. This association may contribute to the dispersion of NOR sites throughout the genome via transposable elements (TEs), which tend to accumulate in heterochromatic regions and may carry portions of rDNA clusters (Cioffi et al., 2010; Symonová et al., 2013; Sember et al., 2015; Glugoski et al., 2018; Yano et al., 2020). Chromosomal pairs 21, 24, and 34 of H strigaticeps (population from the Ivaí River), carrying Ag-NOR/18S rDNA sites, showed a high concentration of Rex3 elements, distributed throughout the entire length of these chromosomes, corroborating the above hypothesis.
The two populations of Hypostomus strigaticeps exhibited variability in both the quantity and the spatial distribution of 5S rDNA sequences. These genes, located at the interstitial position of a metacentric chromosome pair, similar to pair 4 (Ivaí River population) and 3 (Pirapó River population) in the present study, have been reported for H. strigaticeps (Pansonato-Alves et al., 2013; Baumgärtner et al., 2014), H. regani (Ihering, 1905) (Mendes-Neto et al., 2011; Ferreira et al., 2019), H. nigromaculatus (Schubart, 1964) (Martins, Galetti Jr., 2001), and H. iheringii (Regan, 1908) (Traldi et al., 2012). In Hypostominae, the 5S rDNA arrays located at the interstitial position of a metacentric chromosome pair also have been reported for Rhinelepis aspera Spix & Agassiz, 1829 (Bueno et al., 2018), Ancistrus populations (Prizon et al., 2017), Hypancistrus cf. debilittera Armbruster, Lujan & Taphorn, 2007, H. zebra Isbrüeker & Nijssen, 1991(Silva et al., 2014), and Baryancistrus xanthellus Rapp Py-Daniel, Zuanon & de Oliveira, 2011 (Medeiros et al., 2016).
In contrast, the additional 5S rDNA clusters identified on three pairs of acrocentric chromosomes were found in the terminal position on the short arm in the present study for H. strigaticeps from the Ivaí River population. A hypothesis that may explain the increased dispersion of 5S rDNA gene copies in this H. strigaticeps population is the presence of retrotransposons near these sequences, which could contribute to this dispersion. This hypothesis is supported by the high concentration of Rex3 elements in pairs carrying 5S rDNA (pairs 25, 29, 35) in this population, which was not observed in any chromosome pairs in the population of Pirapó River. Therefore, Rex3 elements could be one of those responsible for this numerical variation in 5S rDNA sites between the two populations analyzed here. In Hypostomus, Pansonato-Alves et al. (2013) described an association of Rex1 with 5S rDNA clusters in H. ancistroides (Ihering, 1911) and H. nigromaculatus. A co-localization of Rex3 and rDNA 5S was also observed in the karyotype of some Ancistrus populations from the upper Paraná River basin (Prizon et al., 2018). Therefore, these data reinforce the important role of transposable elements in the chromosomal evolutionary dynamics of very diverse fish groups, such as those of the Loricariidae family.
Other fish species also evidenced TEs found in rDNA sequences or in the spacer of the rDNA sequences, such as Erythrinus erythrinus (Bloch & Schneider, 1801) (Cioffi et al., 2010), Coregonus albula (Linnaeus, 1758) (Symonová et al., 2013), Diplodus sargus (Linnaeus, 1758) (Merlo et al., 2013), Gymnotus mamiraua Albert & Crampton, 2001 (Silva et al., 2016), Rineloricaria latirostris (Boulenger, 1900) (Glugoski et al., 2018), Megalamphodus eques (Steindachner, 1882) (Piscor et al., 2020), and Triportheus species (Yano et al., 2020). This association of TEs and rDNA has also been detected in other groups of animals, plants, fungi, protozoans, and bacteria (Garcia et al., 2024). On the other hand, TEs are not the only agents that may contribute to rDNA spreading. Among eukaryotes, chromosomal rearrangements mediated by ectopic recombination and integration of extrachromosomal rDNA circles facilitated the spread of rDNA (Cohen, Segal, 2009; Nguyen et al., 2010; Proux-Wéra et al., 2013; Sproul et al., 2020; Muirhead, Presgraves, 2021; Dalíková et al., 2023).
Multiple 5S rDNA clusters have been observed in other H. strigaticeps populations (Pansonato-Alves et al., 2013; Baumgärtner et al., 2014) and other species of Hypostomus, such as H. affinis (Steindachner, 1877) (Kavalco et al., 2004), H. regani (Mendes-Neto et al., 2011), H. nigromaculatus (Pansonato-Alves et al., 2013), Hypostomus cf. plecostomus (Linnaeus, 1758) (Oliveira et al., 2015), H. commersoni Valenciennes, 1836 (Lorscheider et al., 2018), H. regani (Ferreira et al., 2019), Hypostomus aff. ancistroides, and H. topavae (Godoy, 1969) (Lara-Kamei et al., 2017), which confirms the diversity in the distribution of these sequences in Hypostomus.
The findings of this study have contributed to the cytogenetic characterization of H. strigaticeps, a species found in the upper Paraná River basin, thereby enhancing our understanding of the Loricariidae family. The study has confirmed the presence of rDNA polymorphism in the species, same in the conserved karyotype shared by the two populations. This polymorphism can be attributed to the synteny of retrotransposons with chromosomes carrying rDNA, mainly Rex3 elements. These transposable elements may have played a vital role in the polymorphism of this ribosomal DNA within the genome of this species.
Acknowledgments
We thank Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), through the Dean of the Postgraduate and Research Department of the Universidade Estadual de Maringa (UEM-PPG), for the master’s scholarship granted to LCB.
References
Alves AL, Borba RS, Oliveira C, Nirchio M, Granado A, Foresti F. Karyotypic diversity and evolutionary trends in the neotropical catfish genus Hypostomus Lacépède, 1803 (Teleostei, Siluriformes, Loricariidae). Comp Cytogenet. 2012; 6(4):443–52. https://doi.org/10.3897/CompCytogen.v6i4.4028
Alves AL, Oliveira C, Foresti F. Comparative cytogenetic analysis of eleven species of subfamilies Neoplecostominae and Hypostominae (Siluriformes: Loricariidae). Genetica. 2005; 124(2):127–36. https://doi.org/10.1007/s10709-004-7561-4
Artoni RF, Bertollo LAC. Cytogenetic studies on Hypostominae (Pisces, Siluriformes, Loricariidae). Considerations on karyotype evolution in the genus Hypostomus. Caryologia. 1996; 49(1):81–90. https://doi.org/10.1080/00087114.1996.10797353
Artoni RF, Bertollo LAC. Nature and distribution of constitutive heterochromatin in fishes, genus Hypostomus (Loricariidae). Genetica. 1999; 106:209–14. https://doi.org/10.1023/A:1003957719178
Artoni RF, Bertollo LAC. Trends in the karyotype evolution of Loricariidae fish (Siluriformes). Hereditas. 2001; 134(3):201–10. https://doi.org/10.1111/j.1601-5223.2001.00201.x
Baumgärtner L, Paiz LM, Zawadzki CH, Margarido VP, Portela Castro ALDB. Heterochromatin polymorphism and physical mapping of 5S and 18S ribosomal DNA in four populations of Hypostomus strigaticeps (Regan, 1907) from the Paraná River Basin, Brazil: evolutionary and environmental correlation. Zebrafish. 2014; 11(5):479–87. https://doi.org/10.1089/zeb.2014.1028
Bertollo L, Cioffi M, Moreira-Filho O. Direct chromosome preparation from freshwater teleost fishes, fish cytogenetic techniques. In: Ozouf-Costaz C, Pisano E, Foresti F, Toledo L, editors. Fish citogenetic techniques: rayfin fishes and Chondrichthyans. Boca Raton: CRC Press; 2015. p.21–26.
Borba RS, Zawadzki CH, Oliveira C, Perdices A, Parise-Maltempi PP, Alves AL. Phylogeography of Hypostomus strigaticeps (Siluriformes: Loricariidae) inferred by mitochondrial DNA reveals its distribution in the upper Paraná River basin.Neotrop Ichthyol. 2013; 11(1):111–16. https://doi.org/10.1590/s1679-62252013000100013
Britski HA, Sato Y, Rosa ABS. Manual de identificação de peixes da região de Três Marias: com chave de identificação para os peixes da bacia do São Francisco. Brasília: CODEVASF, Divisão de Piscicultura e Pesca; 1988.
Bueno V, Henrique C, Margarido VP. Trends in chromosome evolution in the genus Hypostomus Lacépède, 1803 (Osteichthyes, Loricariidae): a new Lace perspective about the correlation between diploid number and chromosomes types. Rev Fish Biol Fish. 2012; 22(1):241–50. https://doi.org/10.1007/s11160-011-9215-9
Bueno V, Konerat JT, Zawadzki CH, Venere PC, Blanco DR, Margarido VP. Divergent Chromosome evolution in Hypostominae tribes (Siluriformes: Loricariidae): correlation of chromosomal data with morphological and molecular phylogenies. Zebrafish. 2018; 15(5):492–503. https://doi.org/10.1089/zeb.2018.1612
Bueno V, Venere PC, Konerat JT, Zawadzki CH, Vicari MR, Margarido VP. Physical mapping of the 5S and 18S rDNA in ten species of Hypostomus Lacépède 1803 (Siluriformes: Loricariidae): evolutionary tendencies in the genus. Sci World J. 2014; 2014(1):943825. https://doi.org/10.1155/2014/943825
Bueno V, Venere PC, Zawadzki CH, Margarido VP. Karyotypic diversification in Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae): biogeographical and phylogenetic perspectives. Rev Fish Biol Fish. 2013; 23(1):103–12. https://doi.org/10.1007/s11160-012-9280-8
Biscotti MA. Mobile elements in ray-finned fish genomes. Life. 2020; 10(10):221. https://doi.org/10.3390/life10100221
Cereali SS, Pomini E, Rosa R, Zawadzki CH, Froehlich O, Giuliano-Caetano L. Karyotype description of two species of Hypostomus (Siluriformes, Loricariidae) of the Planalto da Bodoquena, Brazil. Genet Mol Res. 2008; 7(3):583–91. https://doi.org/10.4238/vol7-3gmr404
Chalopin D, Naville M, Plard F, Galiana D, Volff JN. Comparative analysis of transposable elements highlights mobilome diversity and evolution in vertebrates. Genome Biol Evol. 2015; 7(2):567–80. https://doi.org/10.1093/gbe/evv005
Cioffi MB, Martins C, Bertollo LA. Chromosome spreading of associated transposable elements and ribosomal DNA in the fish Erythrinus erythrinus. Implications for genome change and karyoevolution in fish. BMC Evol Biol. 2010; 10(1):271. https://doi.org/10.1186/1471-2148-10-271
Cohen S, Segal D. Extrachromosomal circular DNA in eukaryotes: possible involvement in the plasticity of tandem repeats. Cytogenet Genome Res. 2009; 124(3–4):327–38. https://doi.org/10.1159/000218136
Dalíková M, Provazníková I, Provazník J, Grof-Tisza P, Pepi A, Nguyen P. The role of repetitive sequences in repatterning of major ribosomal DNA clusters in Lepidoptera. Genome Biol Evol. 2023; 15(6):evad090. https://doi.org/10.1093/gbe/evad090
Endo KS, Martinez ERM, Zawadzki CH, Paiva LRS, Júlio Júnior HF. Karyotype description of possible new species of the Hypostomus ancistroides complex (Teleostei: Loricariidae) and other Hypostominae. Acta Scient Biol Sci. 2012; 34(2):181–89.
Fernandes CA, Damásio JF, Martins IC. Cytogenetic studies in species of family Loricariidae (Osteichthyes, Siluriformes) from Iguatemi river basin, Brazil. First cytogenetic report in Farlowella amazonum (Günther, 1864). Caryologia. 2012; 65(4):276–80. https://doi.org/10.1080/00087114.2012.752913
Ferreira DC, Porto-Foresti F, Oliveira C, Foresti F. Transposable elements as a potential source for understanding the fish genome. Mob Genet Elements. 2011; 1:112–17. https://doi.org/10.4161/mge.1.2.16731
Ferreira GEB, Barbosa LM, Prizon-Nakajima AC, Paiva S, Vieira MMR, Gallo RB et al. Constitutive heterochromatin heteromorphism in the Neotropical armored catfish Hypostomus regani (Ihering, 1905) (Loricariidae, Hypostominae) from the Paraguay River basin (Mato Grosso do Sul, Brazil). Comp Cytogenet. 2019; 13(1):27–39. https://doi.org/10.3897/COMPCYTOGEN.V13I1.30134
Fricke R, Eschmeyer WN, Van der Laan R. Eschmeyer’s catalog of fishes: genera, species, references [Internet]. San Francisco: California Academy of Sciences; 2024. Available from: https://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp
Garcia S, Kovarik A, Maiwald S, Mann L, Schmidt N, Pascual-Díaz JP et al. The dynamic interplay between ribosomal DNA and transposable elements: a perspective from genomics and cytogenetics. Mol Biol Evol. 2024; 41(3):msae025. https://doi.org/10.1093/molbev/msae025
Glugoski L, Giuliano-Caetano L, Moreira-Filho O, Vicari MR, Nogaroto V. Co-located hAT transposable element and 5S rDNA in an interstitial telomeric sequence suggest the formation of Robertsonian fusion in armored catfish. Gene. 2018; 650:49–54. https://doi.org/10.1016/j.gene.2018.01.099
Griffiths S. The use of clove oil as an anaesthetic and method for sampling intertidal rockpool fishes. J Fish Biol. 2000; 57(6):1453–64. https://doi.org/10.1006/jfbi.2000.1406
Hatanaka T, Galetti PM. Mapping of the 18S and 5S ribosomal RNA genes in the fish Prochilodus argenteus Agassiz, 1829 (Characiformes, Prochilodontidae). Genetica. 2004; 122:239–44. https://doi.org/10.1007/s10709-004-2039-y
Howell WM, Black DA. Controlled silver-staining of nucleolus organizer regions with the protective coloidal developer: a 1-step method. Experientia. 1980; 36(8):1014–15. https://doi.org/10.1007/BF01953855
Kavalco KF, Pazza R, Bertollo LAC, Moreira-Filho O. Heterochromatin characterization of four fish species of the family Loricariidae (Siluriformes). Hereditas. 2004; 141(3):237–42. https://doi.org/10.1111/j.1601-5223.2004.01850.x
Lara-Kamei MCS, Baumgärtner L, Paiva S, Zawadzki CH, Martins-Santos IC, Portela-Castro ALB. Chromosomal diversity of three species of Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae), from the Paraná River basin, Brazil: a species complex in Hypostomus ancistroides reinforced by a ZZ/ZW sex chromosome system. Zebrafish. 2017; 14(4):357–63. https://doi.org/10.1089/zeb.2017.1429
Levan A, Fredga K, Sandberg AA. Nomenclature for centromeric position on chromosomes. Hereditas. 1964; 52(2):201–20. https://doi.org/10.1111/j.1601-5223.1964.tb01953.x
Lorscheider CA, Oliveira JIN, Dulz TA, Nogaroto V, Martins-Santos IC, Vicari MR. Comparative cytogenetics among three sympatric Hypostomus species (Siluriformes: Loricariidae): an evolutionary analysis in a high endemic region. BABT. 2018; 61:1–14. https://doi.org/10.1590/1678-4324-2018180417
Lui RL, Blanco DR, Moreira-Filho O, Margarido VP. Propidium iodide for making heterochromatin more evident in the C-banding technique. Biotech Histochem. 2012; 87(7):433–38. https://doi.org/10.3109/10520295.2012.696700
Margarido VP, Moreira-Filho O. Karyotypic differentiation through chromosome fusion and number reduction in Imparfinis hollandi (Ostariophysi, Heptapteridae ). Genet Mol Biol. 2008; 31(1):235–38. https://doi.org/10.1590/S1415-7572008000200012
Martins C, Galetti Jr. PM. Chromosomal localization of 5S rDNA genes in Leporinus fish (Anostomidae, Characiformes). Chrom Res. 1999; 7(5):363–67. https://doi.org/10.1023/A:1009216030316
Martins C, Galetti Jr. PM. Two 5S rDNA arrays in Neotropical fish species: is it a general rule for fishes? Genetica. 2001; 111:439–46. https://doi.org/10.1023/A:1013799516717
Medeiros LA, Ginani EG, Sousa LM, Py-Daniel LHR, Feldberg E. Cytogenetic analysis of Baryancistrus xanthellus (Siluriformes: Loricariidae: Ancistrini), an ornamental fish endemic to the Xingu River, Brazil. Neotrop Ichthyol. 2016; 14(2):e150108. https://doi.org/10.1590/1982-0224-20150108
Mendes-Neto EO, Vicari MR, Artoni RF, Moreira-Filho O. Description of karyotype in Hypostomus regani (Ihering, 1905) (Teleostei, Loricariidae) from the Piumhi river in Brazil with comments on karyotype variation found in Hypostomus. Comp Cytogenet. 2011; 5(2):133–42. https://doi.org/10.3897/compcytogen.v5i2.964
Merlo MA, Cross I, Manchado M, Cárdenas S, Rebordinos L. The 5S rDNA high dynamism in Diplodus sargus is a transposon-mediated mechanism. Comparison with other multigene families and Sparidae species. J Mol Evol. 2013; 76(3):83–97. https://doi.org/10.1007/s00239-013-9541-8
Michele JL, Takahashi CS, Ferrari I. Karyotypic study of some species of the family Loricariidae (Pisces). Cytologia. 1977; 42(3–4):539–46. https://doi.org/10.1508/cytologia.42.539
Muirhead CA, Presgraves DC. Satellite DNA-mediated diversification of a sex-ratio meiotic drive gene family in Drosophila. Nat Ecol Evol. 2021; 5:1604–12. https://doi.org/10.1038/s41559-021-01543-8
Nguyen P, Sahara K, Yoshido A, Marec F. Evolutionary dynamics of rDNA clusters on chromosomes of moths and butterflies (Lepidoptera). Genetica. 2010; 138:343–54. https://doi.org/10.1007/s10709-009-9424-5
Oliveira LC, Ribeiro MO, Costa GDM, Zawadzki CH, Prizon-Nakajima AC, Borin-Carvalho LA et al. Cytogenetic characterization of Hypostomus soniae Hollanda-Carvalho & Weber, 2004 from the Teles Pires River, southern Amazon basin: evidence of an early stage of an XX / XY sex chromosome system. Comp Cytogenet. 2019; 13(4):411–22. https://doi.org/10.3897/compcytogen.v13i4.36205
Oliveira LC, Ribeiro MO, Dutra ES, Zawadzki CH, Portela-Castro ALB, Martins-Santos IC. Karyotype structure of Hypostomus cf. plecostomus (Linnaeus, 1758) from Tapajos River basin, southern amazon: occurrence of sex chromosomes (ZZ/ZW) and their evolutionary implications. Genet Mol Res. 2015; 14(2):6625–34. https://doi.org/10.4238/2015.June.18.5
Pansonato-Alves JC, Serrano EA, Utsunomia R, Scacchetti PC, Oliveira C, Foresti F. Mapping five repetitive DNA classes in sympatric species of Hypostomus (Teleostei: Siluriformes: Loricariidae): analysis of chromosomal variability. Rev Fish Biol Fish. 2013; 23(4):477–89. https://doi.org/10.1007/s11160-013-9303-0
Pinkel D, Straume T, Gray JW. Cytogenetic analysis using quantitative, high-sensitivity, fluorescence hybridization. PNAS. 1986; 83(9):2934–38. https://doi.org/10.1073/pnas.83.9.2934
Piscor D, Paiz LM, Baumgärtner L, Cerqueira FJ, Fernandes CA, Lui RL et al. Chromosomal mapping of repetitive sequences in Hyphessobrycon eques (Characiformes, Characidae): a special case of the spreading of 5S rDNA clusters in a genome. Genetica. 2020; 148(1):25–32. https://doi.org/10.1007/s10709-020-00086-3
Prizon AC, Bruschi DP, Borin-Carvalho LA, Cius A, Barbosa LM, Ruiz HB et al. Hidden diversity in the populations of the armored catfish Ancistrus Kner, 1854 (Loricariidae, Hypostominae) from the Paraná River basin revealed by molecular and cytogenetic data. Front Genet. 2017; 8:185. https://doi.org/10.3389/fgene.2017.00185
Prizon AC, Bruschi DP, Gazolla CB, Borin-Carvalho LA, Portela-Castro ALB. Chromosome spreading of the retrotransposable Rex-3 element and microsatellite repeats in karyotypes of the Ancistrus populations. Zebrafish. 2018; 15(5):504–14. https://doi.org/10.1089/zeb.2018.1620
Proux-Wéra E, Byrne KP, Wolfe KH. Evolutionary mobility of the ribosomal DNA array in yeasts. Genome Biol Evol. 2013; 5(3):525–31. https://doi.org/10.1093/gbe/evt022
Regan OT. Descriptions of new loricariid fishes from South America. Proc Zool Soc Lond. 1907; 77(4):795–800. https://doi.org/10.1111/j.1469-7998.1907.tb06957.x
Rubert M, Rosa R, Jerep FC, Bertollo LAC, Giuliano-Caetano L. Cytogenetic characterization of four species of the genus Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae) with comments on its chromosomal diversity. Comp Cytogenet. 2011; 5(5):397–410.https://doi.org/10.3897/compcytogen.v5i5.1589
Rubert M, Rosa R, Zawadzki CH, Mariotto S, Moreira-Filho O, Giuliano-Caetano L. Chromosome mapping of 18S ribosomal RNA genes in eleven Hypostomus species (Siluriformes, Loricariidae): diversity analysis of the sites. Zebrafish. 2016; 13(4):360–68. https://doi.org/10.1089/zeb.2016.1279
Rubert M, Takagui FH, Santos KF, Santana Pompeo LR, Rosa R, Zawadzki CH et al. Topotype-based chromosomal diversity among five species of freshwater armored catfishes in the Hypostomus auroguttatus supergroup (Actinopterygii: Siluriformes). Zoolog Sci. 2022; 39(5):446–52. https://doi.org/10.2108/zs210103
Rubert M, Zawadzki CH, Giuliano-Caetano L. Cytogenetic characterization of Hypostomus nigromaculatus (Siluriformes: Loricariidae). Neotrop Ichthyol. 2008; 6(1):93–100. https://doi.org/10.1590/S1679-62252008000100011
Sember A, Bohlen J, Šlechtová V, Altmanová M, Symonová R, Ráb P. Karyotype differentiation in 19 species of river loach fishes (Nemacheilidae, Teleostei): extensive variability associated with rDNA and heterochromatin distribution and its phylogenetic and ecological interpretation. BMC Evol Biol. 2015; 15(1):251. https://doi.org/10.1186/s12862-015-0532-9
Shao F, Han M, Peng Z. Evolution and diversity of transposable elements in fish genomes. Sci Rep. 2019; 9:15399. https://doi.org/10.1038/s41598-019-51888-1
Silva M, Barbosa P, Artoni RF, Feldberg E. Evolutionary dynamics of 5S rDNA and recurrent association of transposable elements in electric fish of the family Gymnotidae (Gymnotiformes): the case of Gymnotus mamiraua. Cytogenet Genome Res. 2016; 149(4):297–303. https://doi.org/10.1159/000449431
Silva M, Ribeiro ED, Matoso DA, Sousa LM, Hrbek T, Py-Daniel LR et al. Chromosomal polymorphism in two species of Hypancistrus (Siluriformes: Loricariidae): an integrative approach for understanding their biodiversity. Genetica. 2014; 142(2):127–39. https://doi.org/10.1007/s10709-014-9760-y
Sproul JS, Barton LM, Maddison DR. Repetitive DNA profiles reveal evidence of rapid genome evolution and reflect species boundaries in ground beetles. Syst Biol. 2020; 69(6):1137–48. https://doi.org/10.1093/sysbio/syaa030
Sumner AT. A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res. 1972; 75(1):304–06. https://doi.org/10.1016/0014-4827(72)90558-7
Symonová R, Majtánová Z, Sember A, Staaks GB, Bohlen J, Freyhof J et al. Genome differentiation in a species pair of coregonine fishes: an extremely rapid speciation driven by stress-activated retrotransposons mediating extensive ribosomal DNA multiplications. BMC Evol Biol. 2013; 13(1):42. https://doi.org/10.1186/1471-2148-13-42
Traldi JB, Blanco DR, Vicari MR, Martinez JF, Lui RL, Barros AV et al. Chromosomal diversity in Hypostomus (Siluriformes, Loricariidae) with emphasis on physical mapping of 18S and 5S rDNA sites. Genet Mol Res. 2013; 12(1):463–71. https://doi.org/10.4238/2013.february.8.11
Traldi JB, Lui RL, Martinez JF, Vicari MR, Nogaroto V, Filho OM et al. Chromosomal distribution of the retroelements Rex1, Rex3 and Rex6 in species of the genus Harttia and Hypostomus (Siluriformes: Loricariidae). Neotrop Ichthyol. 2019; 17(2):e190010. https://doi.org/10.1590/1982-0224-20190010
Traldi JB, Vicari MR, Blanco DR, Martinez JF, Artoni RF, Moreira-Filho O. First karyotype description of Hypostomus iheringii (Regan, 1908): a case of heterochromatic polymorphism. Comp Cytogenet. 2012; 6(2):115–25. https://doi.org/10.3897/CompCytogen.v6i2.2595
Volff J-N. Turning junk into gold: domestication of transposable elements and the creation of new genes in eukaryotes. BioEssays. 2006; 28(9):913–22. https://doi.org/10.1002/bies.20452
Volff J-N, Körting C, Froschauer A, Sweeney K, Schartl M. NonLTR retrotransposons encoding a restriction enzyme-like endonuclease in vertebrates. J Mol Evol. 2001; 52(4):351–60. https://doi.org/10.1007/s002390010165
Volff J-N, Körting C, Schartl M. Multiple lineages of the non-LTR retrotransposon Rex1 with varying success in invading fish genomes. Mol Biol Evol. 2000; 17(11):1673–84. https://doi.org/10.1093/oxfordjournals.molbev.a026266
Volff J-N, Körting C, Sweeney K, Schartl M. The non-LTR retrotransposon Rex3 from the fish Xiphophorus is widespread among teleosts. Mol Biol Evol. 1999; 16(11):1427–38. https://doi.org/10.1093/oxfordjournals.molbev.a026055
Weber C. Subfamily Hypostominae (Armored catfishes). In: Reis RE, Kullander SO, Ferraris CJ, Jr., editors. Check list of the freshwater fishes of South and Central America. Porto Alegre: Edipucrs; 2003. p.351–72.
Wicker T, Sabot F, Hua-Van A, Bennerzen JL, Capy P, Chalhoub B et al. A unified classification system for eukaryotic transposable elements. Nat Rev Genet. 2007; 8:973–82. https://doi.org/10.1038/nrg2165
Yano CF, Merlo MA, Portela-Bens S, Cioffi MDB, Bertollo LAC, Santos-Júnior CD et al. Evolutionary dynamics of multigene families in Triportheus (Characiformes, Triportheidae): a transposon mediated mechanism? Front Mar Sci. 2020; 7:6. https://doi.org/10.3389/fmars.2020.00006
Zanata AM, Pitanga BR. A new species of Hypostomus Lacépède, 1803 (Siluriformes: Loricariidae) from rio Itapicuru basin, Bahia State, Brazil. Zootaxa. 2016; 4137:223–32. https://doi.org/10.11646/zootaxa.4137.2.4
Zawadzki CH, Tencatt LFC, Froehlich O. A new unicuspid-toothed species of Hypostomus Lacépède, 1803 (Siluriformes: Loricariidae) from the rio Paraguai basin. Neotrop Ichthyol. 2014; 12(1):97–104. https://doi.org/10.1590/S1679-62252014000100010
Authors
![]() Lígia Carla Balini1
Lígia Carla Balini1 ![]() ,
, ![]() Rafael Fernando de Melo1,
Rafael Fernando de Melo1, ![]() Ana Luiza de Brito Portela-Castro1,2,
Ana Luiza de Brito Portela-Castro1,2, ![]() Cláudio Henrique Zawadzki2,3,
Cláudio Henrique Zawadzki2,3, ![]() Marcos Otávio Ribeiro4,
Marcos Otávio Ribeiro4, ![]() Carlos Alexandre Fernandes1,2 and
Carlos Alexandre Fernandes1,2 and ![]() Luciana Andreia Borin-Carvalho1
Luciana Andreia Borin-Carvalho1
[1] Universidade Estadual de Maringá, Departamento de Biotecnologia, Genética e Biologia Celular, Bloco H67, sala 09, Avenida Colombo, 5790, 87020-900 Maringá, PR, Brazil. (LCB) ligia_balini@hotmail.com (corresponding author), (RFM) rafael. leafarmelo@gmail.com, (ALBPC) albpcastro@nupelia.uem.br, (CAF) cafernandes@uem.br, (LABC) labcarvalho@uem.br.
[2] Núcleo de Pesquisas em Limnologia, Ictiologia e Aquicultura, Universidade Estadual de Maringá, 87020-900 Maringá, PR, Brazil.
[3] Universidade Estadual de Maringá, Departamento de Biologia, 87020-900 Maringá, PR, Brazil. (CHZ) chzawadzki@uem.br.
[4] Universidade do Paraná, Departamento de Biologia, campus União da Vitória, 84600-185 União da Vitória, PR, Brazil. (MOR) marcos.ribeiro@unespar.edu.br.
Authors’ Contribution 

Lígia Carla Balini: Conceptualization, Data curation, Investigation, Methodology, Resources, Writing-original draft, Writing-review and editing.
Rafael Fernando de Melo: Methodology, Writing-review and editing.
Ana Luiza de Brito Portela-Castro: Writing-original draft, Writing-review and editing.
Cláudio Henrique Zawadzki: Methodology, Writing-original draft, Writing-review and editing.
Marcos Otávio Ribeiro: Writing-original draft, Writing-review and editing.
Carlos Alexandre Fernandes: Supervision, Writing-original draft, Writing-review and editing.
Luciana Andreia Borin-Carvalho: Methodology, Project administration, Writing-original draft, Writing-review and editing.
Ethical Statement
Animals were captured with the permission of the Instituto Chico Mendes de Conservação da Biodiversidade (ICMBio; number 60070–3). This study was carried out in strict accordance with the recommendations of the Guide for the Care and Use of Laboratory Animals, approved by the Committee on Ethics of Animal Experiments of the Universidade Estadual de Maringá (License number: 1968010719– CEUA/UEM). The experiments followed ethical conduct, and before euthanasia, the fish were anesthetized by an overdose of clove oil (Griffiths, 2000).
Competing Interests
The author declares no competing interests.
How to cite this article
Balini LC, Melo RF, Portela-Castro ALB, Zawadzki CH, Ribeiro MO, Fernandes CA, Borin-Carvalho LA. Polymorphism of 5S and 18S ribosomal DNA in a conserved karyotype of the two Hypostomus strigaticeps (Siluriformes: Loricariidae) populations from the upper Paraná River basin. Neotrop Ichthyol. 2025; 23(2):e240061. https://doi.org/10.1590/1982-0224-2024-0061
Copyright
This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Distributed under
Creative Commons CC-BY 4.0

© 2025 The Authors.
Diversity and Distributions Published by SBI
![]() Accepted March 31, 2025
Accepted March 31, 2025
![]() Submitted July 3, 2024
Submitted July 3, 2024
![]() Epub June 30, 2025
Epub June 30, 2025