![]() Katiane M. Ferreira1,
Katiane M. Ferreira1, ![]() Alexandre C. Ribeiro1,
Alexandre C. Ribeiro1, ![]() Flávio C. T. Lima2,
Flávio C. T. Lima2, ![]() Hugmar P. da Silva1,
Hugmar P. da Silva1, ![]() Daniela C. Ferreira1 and
Daniela C. Ferreira1 and ![]() Juan Marcos Mirande3
Juan Marcos Mirande3 ![]()
PDF: EN XML: EN | Supplementary: S1 S2 S3 S4 S5 S6 | Cite this article
Abstract
A new species of Inpaichthys is described from a tributary of the rio Canamã, rio Aripuanã basin, Mato Grosso State, Brazil. The new species can be diagnosed from its congeners by the color pattern in life and by morphometric and meristic features. A comprehensive phylogenetic analysis of the Characidae, conducted to assess the generic placement of the new species, revealed that Hasemania nambiquara is also a member of Inpaichthys and thus transferred to this genus. A monophyletic group composed of the three known species of Inpaichthys is hypothesized to be related to Nematobrycon and a clade composed of Carlana, Pseudochalceus, and Rhoadsia, among the taxa herein analyzed. A diagnosis for Inpaichthys and morphometric and meristic data of I. kerri are also presented.
Keywords: Hasemania nambiquara, Inpaichthys kerri, rio Madeira basin, Phylogeny, Taxonomy.
Uma espécie nova de Inpaichthys é descrita de um tributário do rio Canamã, bacia do rio Aripuanã, estado de Mato Grosso, Brasil. A espécie nova pode ser diagnosticada de suas congêneres por seu padrão de colorido em vida e por caracteres morfométricos e merísticos. Uma ampla análise filogenética de Characidae, conduzida para avaliar o posicionamento genérico da nova espécie, demonstrou que Hasemania nambiquara também é um membro do gênero Inpaichthys e esta espécie é aqui transferida a esse gênero. Um grupo monofilético composto pelas três espécies de Inpaichthys é hipotetizado como sendo relacionado a Nematobrycon e o clado composto por Carlana, Pseudochalceus e Rhoadsia, entre os táxons aqui examinados. Uma diagnose para Inpaichthys e os dados morfométricos e merísticos de I. kerri também são apresentados.
Palavras-chave: Bacia do rio Madeira, Filogenia Hasemania nambiquara, Inpaichthys kerri, Taxonomia.
Introduction
Inpaichthys Géry & Junk, 1977 is currently a monotypic genus of Characidae. It was described by Géry, Junk (1977) to include Inpaichthys kerri Géry & Junk, 1977. The generic name honors the Instituto Nacional de Pesquisas da Amazônia (INPA), which had a research station in the Aripuanã city, above the great falls of Dardanelos and Andorinhas, on the upper rio Aripuanã, the type-locality of I. kerri.
Inpaichthys kerri is a small-sized species, not surpassing 28 mm SL, with a unique life color pattern among characids composed of a wide, conspicuous longitudinal black stripe from the jaw to the caudal fin, and the dorsal half of the body is bright blue and glows like “neon tubes” (Géry, Junk, 1977). Despite uncertainties regarding its relationship within Characidae, Géry, Junk (1977) proposed the genus Inpaichthys based on a combination of non-exclusive morphological characters, viz., a subsuperior mouth, tricuspid teeth arranged in two very irregular series on the premaxilla, the inner series generally composed of only two median teeth on each side, maxilla with 7–8 conical to tricuspid teeth, third infraorbital (“suborbital” of Géry, Junk, 1977) large and in contact with the preopercular canal, absence of infraorbitals 5 and 6 (single postorbital of Géry, Junk, 1977), dorsal fin slightly behind the middle of the body, a deep caudal peduncle, and a peculiar coloration consisting of a wide and conspicuous longitudinal stripe extending from the dentary to the caudal fin, curved ventrally (Géry, Junk, 1977). According to a recently published phylogenetic hypothesis based on combined morphological and molecular data, Inpaichthys belongs to the tribe Rhoadsiini, being closely related to the genera Nematobrycon Eigenmann, 1911, Pseudochalceus Kner, 1863, and the clade of Carlana Strand, 1928, and Rhoadsia Fowler, 1911 (Mirande, 2019).
Recent collecting efforts at the rio Canamã, a tributary of the rio Aripuanã (the river basin where Inpaichthys kerri occur) revealed an additional, very distinctive, new Inpaichthys species. During our investigation on this new species, we tested our suspicion based on color pattern and distribution that Hasemania nambiquara Bertaco & Malabarba, 2007, would be actually a species of Inpaichthys, which was corroborated by including morphological data of the species into a phylogenetic analysis including both I. kerri and the new species, thus raising the number of species within this genus from one to three. We also present a hypothesis about the phylogenetic relationships between the species of genus using the framework provided by Mirande (2019), expanded in Terán et al. (2020) and subsequent articles (Lima et al., 2021; Ferreira et al., 2023).
Material and methods
Morphological analysis. Counts and measurements follow Fink, Weitzman (1974) and Menezes, Weitzman (1990), except for counts of the horizontal scale rows below the lateral line, that were counted to the pelvic-fin insertion. In the description, presented counts are followed by the number of specimens presenting a specific count in parentheses. Asterisks indicate values of the holotype. Counts of vertebrae and supraneurals are taken from cleared and stained (c&s) specimens; total vertebrae counts include the four vertebrae of the Weberian apparatus and the terminal centrum. Circuli and radii counts are taken from scale row immediately above the lateral line. Specimens were cleared and stained following the method of Taylor, Van Dyke (1985). Osteological terminology follows Weitzman (1962) with the modifications listed by Vari, Harold (2001). Institutional abbreviations follow Sabaj (2020). In the original description, the morphometrics of Inpaichthys kerri deviate from standard established by Fink, Weitzman (1974) and followed ever since in most of the literature on Characiformes systematics. For this reason, the morphometrics data of I. kerri were also obtained and presented in Tab. 1 for comparison reasons. Data on Hasemania nambiquara was obtained both from the direct examination of two paratypes, and from the original description (Bertaco, Malabarba, 2007).
Extraction of the DNA, amplification, purification, and sequencing. The DNA of seven specimens of the new species of Inpaichthys was extracted using the saline extraction protocol of Aljanabi, Martinez (1997). We amplified two mitochondrial molecular markers, Cytochrome c Oxidase I (COI) and 16S rRNA with primers described by Ward et al. (2005) and Palumbi, 1996, respectively. The Polymerase Chain Reaction (PCR) was prepared in a final volume of 25 µL containing: 12,5 μl GoTaq® DNA Polymerase (Promega), 0.5 μl of the each primer (10 mM), 1 μl of DNA, and ultrapure water to complete the final volume. The reactions were processed in a thermocycler (Applied Byosystems, Veriti), with initial denaturation at 94 ºC for 5 min, followed by 35 cycles of denaturation at 94 °C for 45 seconds, annealing at 53 °C for 45 sec, and extension at 72 ºC for 60 sec, with a final extension at 72 °C for 5 min. The amplicons of the genes COI and 16S rRNA were purified and sequenced by Biotecnologia Pesquisa e Inovação (BPI, https://bpibiotecnologia.com.br). The samples were purified using a magnetic bead kit of the AMpure XP type, and sequenced with a BigDye® Terminator v. 3.1 Cycle Sequencing kit (Applied Biosystems), following the manufacturer’s protocols. The samples were sequenced automatically by capillary electrophoresis in an ABI3730xl Genetic Analyzer (Applied Biosystems). The new sequences were deposited in the Genbank (16S rRNA: OR502673, OR502674, OR502675, OR502676, OR502677, OR502678) and BOLD Systems (COI: INPAI001–23, INPAI002–23, INPAI003–23, INPAI004–23).
Phylogenetic analysis. Phylogenetic relationships of the new species were evaluated through the analysis of a dataset combining morphological and molecular evidence of the Characidae. The dataset included 520 morphological characters and nine blocks of molecular data. The morphological dataset was firstly defined by Mirande (2009, 2010), and had several stages of addition and revision of data (Mirande et al., 2013, 2023; Ohara et al., 2017, 2019; Mirande, 2019; Terán et al., 2020; Lima et al., 2021; Ferreira et al., 2023). The molecular data included mitochondrial ribosomal 12S and 16S, ATP synthase F0 subunit 6 (ATP6), cytochrome c oxidase subunit I (COI), cytochrome b (CYTB) and the nuclear coding markers myosin heavy chain 6 (MYH6), patched domain containing 4 (PTCHD4) [mentioned as PTCHD1 in Thomaz et al. (2015)], and recombination activating 1 and 2 (RAG1 and RAG2). Sequences were aligned with Muscle (Edgar, 2004) through its implementation in Aliview (Larsson, 2014). The latter program was also used to corroborate codon positions, in order to prevent problems with reading frames of protein-coding sequences. Most of the molecular data were extracted from previous articles (e.g., Javonillo et al., 2010; Oliveira et al., 2011; Thomaz et al., 2015; Terán et al., 2020) and curated through the corroboration of vouchers identification in collection databases or personal examination of the specimens, when possible. The list of morphological characters was provided in Mirande (2010, 2019) and a table with the Genbank (www.ncbi.nlm.nih.gov/nuccore) or BOLD (www.boldsystems.org/) accessions herein analyzed was published in Terán et al. (2020). Herein, we added new molecular data available in Genbank since Terán et al. (2020) both for the Characidae and outgroup families (Tab. S1), in addition to the sequences of the new species, listed in the previous section. The complete dataset herein analyzed, with data for 578 species of Characiformes, is provided as supplementary data (S2). Relative to previous articles, we added herein the new species and Hasemania nambiquara.
All analyses were performed with parsimony under extended implied weighting (Goloboff, 1993, 2014) in TNT v. 1.6 for Linux (Goloboff, Morales, 2023). Analyses followed the same procedure than in Ferreira et al. (2023), exploring a broad range of conditions combining five weighting schemes (general ways of weighting characters against their homoplasy) and six weighting strenghts (values of K), resulting then in 30 analyses. In the different weighting schemes, molecular characters were weighted according to (1) BLK: the average homoplasy of entire block of data (marker), (2) COD: the average homoplasy of three contiguous characters (a codon; applicable for coding sequences), (3) GRO: the average homoplasy of groups formed by 30 contiguous characters (10 codons, for coding sequences), (4) SEP: the homoplasy of each separate character, and (5) POS: the average homoplasy of characters from each codon position for each coding marker, with the ribosomal characters weighted according to their own homoplasy. Weighting schemes were combined with six strengths determined for different K-values (Goloboff, 1993, 2014) in which an averagely homoplastic character had 70, 74, 78, 82, 86, and 90% of the weight of a character with no homoplasy. The analyzed K-values (16, 20, 25, 32, 43, and 64) sample regularly decreasing weighting strengths (Mirande, 2009). Each analytical condition is expressed as the abbreviation of the weighting scheme followed by the weighting strength (value of K). For instance, in COD64, three contiguous sites (a codon) are grouped to be collectively weighted (COD scheme) with the homoplasy downweighted according to a K = 64. After searching for the most parsimonious trees on each condition, a metacriterion based on global parsimony was used to select the final hypothesis (Terán et al., 2020; Mirande et al., 2023). According to that criterion, the optimality of each most parsimonious tree is evaluated according to all the explored parameters, selecting the one that is better in the average of conditions. Clade supports were estimated through symmetric resampling (300 replicates and the values are represented as GC-values (difference of frequencies between the obtained clade and its most frequent alternative).
Supplementary files provided include: the analyzed GenBank accessions and their vouchers, when known (Tab. S1); the analyzed dataset combining all the morphological and molecular data (S2); the scripts for TNT software to reproduce searches and trees comparisons herein performed (S3); a collection of all trees obtained under the complete range of explored analytical conditions, formatted for TNT software (S4); the list of autapomorphies and synapomorphies of the obtained nodes of the phylogenetic hypothesis herein proposed (S5); and our final hypothesis, as cladograms in vectorial graphic format, containing node numbers (corresponding to those listed in S5), frequencies, and support values (Fig. S6).
Results
Inpaichthys Géry & Junk, 1977
Type-species. Inpaichthys kerri Géry & Junk, 1977
Diagnosis. We found eight morphological non-exclusive synapomorphies supporting the monophyly of Inpaichthys that, in combination, could be useful to diagnose the genus:nasal septum of mesethmoid formed by two parallel lamellae (9:1), absence of a dorsal expansion of rhinosphenoid projected medial to olfactory nerves (35:0), short supraoccipital spine covering dorsally only anterior axis of neural complex of Weberian apparatus (62:1), articulation between second and third infraorbitals anteroventrally angled (86:1), absence of anguloarticular portion of mandibular laterosensory canal (105:1), frontal and pterotic laterosensory canals forming an angle on pore receiving infraorbital canal (118:1), maxillary teeth extended approximately to half maxillary length (191:0), and presence of 24 or fewer branched anal-fin rays, with a reversal to 21–25 in the new species herein described (420:0). Species of Inpaichthys can also be recognized by the combination of the following characters: two or more maxillary teeth with larger one, tri or pentacuspidate, ten or fewer gill rakers on first hypobranchial and ceratobranchial, interrupted lateral line, lack of parietal branch of supraorbital cephalic laterosensory canal, pore of the supraorbital cephalic laterosensory canal situated just anterior to dilatator fossa oriented dorsomedially (Mirande, 2018: S11, figure 11), circuli absent on posterior field of scales, iii+9 dorsal-fin rays with the anterior one visible only in c&s specimens, lack of secondary-sexual hooks on males, and lack of pseudotympanum.
Inpaichthys nambiquara (Bertaco & Malabarba, 2007), new combination
(Fig. 1)
Hasemania nambiquara Bertaco & Malabarba, 2007:350–53 (Type-locality: “Brazil, Mato Grosso, Comodoro, rio Doze de Outubro on Highway BR-364 between Comodoro and Vilhena, tributary of rio Juruena, upper rio Tapajós drainage, 12º58’39”S 60º00’30”W”; see Remarks below). ―Ohara, Loeb, 2016:3–6 (Brazil, Mato Grosso State, igarapé Mutum, rio Juruena basin; photo in life, abundance).
Diagnosis. Five morphological autapomorphies were found for Inpaichthys nambiquara: third infraorbital not reaching horizontal arm of preopercle in its anterior margin (88:1), presence of (a reduced, in this species) fourth infraorbital (90:0), three or fewer maxillary teeth (190:0), five or more cusps on anterior maxillary teeth (193:1), and the possession of four or fewer supraneurals (392:0). Inpaichthys nambiquara can be easily distinguished from both I. kerri and the new taxon herein described by the absence of an adipose fin (vs. adipose fin present in other Inpaichthys species), and by having a conspicuous dark stripe along predorsal area (vs. absence of a conspicuous dark stripe along predorsal area in the remaining Inpaichthys species). Inpaichthys nambiquara can additionally be distinguished from the new species by the following combination of characters: presence of a conspicuous, broad, ventrally curved midlateral stripe extending from snout to caudal peduncle (vs. presence of a narrow, straight dark midlateral stripe along the longitudinal septum of body, extending from second humeral blotch to median caudal-fin rays), absence of six red dotted longitudinal stripes on the body flanks in living specimens (vs. presence), 16–18 branched anal-fin rays (vs. 21–25 branched anal-fin rays), presence of a single humeral blotch (vs. two humeral blotches), and largest dentary teeth with five cusps (vs. largest dentary teeth with three cusps). Inpaichthys nambiquara can additionally be distinguished from I. kerri by the presence of a humeral blotch (vs. absence of any humeral blotch), and 2–3 maxillary teeth, largest one pentacuspid (vs. 6–9 maxillary teeth, largest tricuspid), respectively.
Description. See Bertaco, Malabarba (2007) for a complete description of the species.
Remarks. The type-locality of Inpaichthys nambiquara was originally indicated to be at the rio Doze de Outubro at the road BR-364. The rio Doze de Outubro is a tributary of the rio Camararé, a major tributary of the rio Juruena. Ohara, Loeb (2016) sampled extensively some of the headwater tributaries of the rio Camararé crossing the highlands of the Chapada dos Parecis at the road BR-364, including the rio Doze de Outubro, and only collected the species at the rio Mutum, another tributary of the rio Camararé, which runs parallel to the rio Doze de Outubro, roughly 20 km apart along their lengths (> 100 km) before joining the rio Camararé. Despite their proximity and the fact that they both belong to the same sub-basin, they are apparently quite distinct in the composition of their ichthyofaunas (Ohara, Loeb, 2016). The type-locality of I. nambiquara has been recently recognized as being incorrectly stated in the original description, and to be indeed the rio Mutum (V. A. Bertaco, 2023, pers. comm.). In fact, one of the authors (FCTL) participated in the field expedition where the type-material of I. nambiquara was obtained and photographed specimens sampled at the rio Mutum, including a specimen of I. nambiquara, which lends support to the idea that the locality was incorrectly stated at the original description. We take this opportunity to amend the type-locality of the species to “Brazil, Mato Grosso, Comodoro, rio Mutum, tributary of rio Camararé, at highway BR-364 between Comodoro and Vilhena, rio Juruena basin, 13º05’08”S 59º53’32”W”. In the original description, Bertaco, Malabarba (2007) did not mention the presence of sexual dimorphism in the species. However, H.-G. Evers (2023, pers. comm.), who bred the species in captivity, noticed that males are slightly larger than females, presenting a dark margin on the anal fin, and present a broader midlateral stripe (compare Figs. 1A–B). Inpaichthys kerri apparently possesses a similar sexual dimorphism, with males presenting a broader midlateral stripe and an anal-fin margin straighter than females (see Pastana et al., 2017:1308, figs. 3E–F).
FIGURE 1| Inpaichthys nambiquara,new combination, in life. A. Male; B. Female. Photos by Hans-Georg Evers.
Material examined. Inpaichthys nambiquara.Brazil, Mato Grosso State, Comodoro, upper rio Tapajós basin:MCP 38038, paratypes, 2 of 4, 1 c&s, 21.1–23.5 mm SL.
Inpaichthys parauapiranga, new species
urn:lsid:zoobank.org:act:94422A47-3AAF-47D1-9832-17A364AFE9CB
(Figs. 2–4; Tab. 1)
Holotype. CPUFMT 7908, 25.4 mm SL, Brazil, Mato Grosso State, Aripuanã, tributary of the rio Canamã, rio Aripuanã basin, upper rio Madeira basin, 10º24’40”S 58º54’13”W, 14 Mar 2021, A. C. Ribeiro & H. Pains da Silva.
Paratypes. All from the same locality of the holotype. CI-FML 7946, 4, 14.3–16.0 mm SL; CPUFMT 7891, 51, 10.3–25.4 mm SL (8 c&s, 16.1–24.4 mm SL); ZUEC 17698, 4, 13.7–16.1 mm SL; collected with the holotype; CPUFMT 7892, 10, 12.9–23.6 mm SL, 20 Aug 2021, H. Pains da Silva. CPUFMT 7893, 11, 13.1–24.2 mm SL, 20 Aug 2021, H. Pains da Silva. MUBIO 113, 5, 19.0–26.3 mm SL; ZUEC 17920, 4, 20.9–22.4 mm SL, 25 Oct 2022, F. C. P. Dagosta, T. J. Seren, B. A. Nagamatsu & J. Damasceno.
FIGURE 2| Inpaichthys parauapiranga, holotype, CPUFMT 7908, 25.4 mm SL, Brazil, Mato Grosso State, Aripuanã, rio Canamã, rio Aripuanã basin. A. When transferred to alcohol after several days of fixation; B. Recently fixed in formaldehyde. Photos by Alexandre C. Ribeiro.
Diagnosis. Inpaichthys parauapiranga can be easily distinguished from both I. kerri and I. nambiquara by its life color pattern. Inpaichthys parauapiranga has six red dotted longitudinal stripes on the flanks that are absent in I. kerri and I. nambiquara. Additionally, I. parauapiranga has a narrow dark midlateral stripe along the longitudinal septum of body, extending from second humeral blotch to median caudal-fin rays (vs. a large dark midlateral stripe along longitudinal septum of body extending from snout to caudal peduncle in I. kerri and I. nambiquara). Inpaichthys parauapiranga can also be distinguished from I. kerri by the following combination of characters: presence of two humeral blotches (vs. absence of humeral blotches); five scale rows between dorsal-fin origin and lateral line (vs. six scale rows between dorsal-fin origin and lateral line); and 12–14 circumpeduncular scales (vs. 16 circumpeduncular scales). Inpaichthys parauapiranga is also distinguished from I. nambiquara by the following combination of characters: adipose fin present (vs. adipose fin absent); 21–25 anal-fin branched rays (vs. 16–18 anal-fin branched rays); two humeral blotches (vs. a single humeral blotch), and largest dentary teeth with three cusps (vs. largest dentary teeth with five cusps).We found three morphological autapomorphies for I. parauapiranga: the absence of a bony rhinosphenoid (34:0), the absence of a conspicuous ethmopalatine cartilage (208:0), and the absence of denticles on gill rakers (300:1).
Description. Morphometric and meristic data for holotype and paratypes in Tab. 1. Largest specimen examined 26.3 mm SL. Body laterally compressed. Greatest body depth slightly anterior to vertical through dorsal-fin origin. Dorsal profile of head convex from tip of upper jaw to vertical through anterior nostril; slightly convex from that point to tip of supraoccipital spine. Dorsal body profile slightly convex from tip of supraoccipital spine to dorsal-fin origin; straight along dorsal-fin base; straight from terminus of dorsal-fin base to adipose-fin insertion, and slightly concave posteriorly from that point to anteriormost procurrent caudal-fin ray. Ventral profile of head and body convex from tip of lower jaw to pelvic-fin insertion, straight from that point to anal-fin origin, straight along anal-fin base and slightly concave along caudal peduncle.
TABLE 1 | Morphometric data of Inpaichthys parauapiranga (N = 20) and I. kerri (N = 10; CPUFMT 4869; CPUFMT 4988; CPUFMT 5066). Range of I. parauapiranga includes the holotype. SD = standard deviation.
| Inpaichthys parauapiranga | Inpaichthys kerri | |||||
Holotype | Paratypes |
| |||||
| Range | Mean | SD | Range | Mean | SD | |
Standard length | 25.4 | 17.2–25.4 | 21.5 | – | 19.7–26.6 | 22.4 | – |
Percentages of standard length | |||||||
Body depth | 32.5 | 32.1–36.4 | 34.7 | 1.1 | 30.6–36.3 | 32.7 | 1.8 |
Snout to dorsal-fin origin | 54.7 | 53.1–57.6 | 55.1 | 1.1 | 51.6–55.9 | 54.1 | 1.3 |
Snout to pectoral-fin origin | 25.5 | 25.5–28.3 | 26.7 | 0.9 | 24.9–27.3 | 25.7 | 1.1 |
Snout to pelvic-fin origin | 43.6 | 43.6–47.6 | 46.0 | 1.1 | 42.9–47.1 | 44.6 | 1.4 |
Snout to anal-fin origin | 56.9 | 53.3–57.6 | 56.1 | 1.3 | 56.2–60.5 | 59.1 | 1.2 |
Pelvic-fin insertion to anal-fin origin | 13.9 | 11.8–15.3 | 13.7 | 1.0 | 11.3–15.8 | 13.6 | 1.3 |
Caudal–peduncle depth | 13.9 | 11.6–14.0 | 12.6 | 0.7 | 10.0–11.7 | 11.2 | 0.5 |
Caudal-peduncle length | 9.3 | 9.3–11.4 | 10.4 | 0.8 | 8.9–12.9 | 11.8 | 1.3 |
Dorsal-fin origin to caudal-fin base | 51.4 | 46.8–51.5 | 49.3 | 1.2 | 46.4–49.4 | 48.3 | 1.0 |
Dorsal-fin base length | 12.5 | 10.9-13.7 | 12.5 | 0.6 | 10.7–13.2 | 12.0 | 0.7 |
Anal-fin base length | 34.9 | 31.3–37.1 | 34.3 | 1.4 | 29.4–32.7 | 31.3 | 1.2 |
Pectoral-fin length | 20.6 | 18.2–23.9 | 21.3 | 1.6 | 18.3–22.6 | 20.2 | 1.3 |
Pelvic-fin length | 14.3 | 12.6–15.9 | 14.7 | 0.9 | 14.0–16.8 | 15.1 | 0.8 |
Dorsal-fin length | 29.6 | 26.4–34.6 | 29.9 | 1.8 | 26.1–30.6 | 28.5 | 1.8 |
Anal-fin length | 18.5 | 17.1–23.9 | 20.5 | 1.8 | 17.4–21.3 | 19.1 | 1.4 |
Bony head length | 25.7 | 25.6–29.5 | 27.5 | 1.3 | 24.3–28.0 | 25.8 | 1.1 |
Head depth | 24.8 | 23.5–28.1 | 26.1 | 1.3 | 23.1–25.8 | 24.3 | 1.1 |
Percentages of head length | |||||||
Horizontal eye diameter | 37.9 | 37.0–40.4 | 38.9 | 1.0 | 38.7–41.9 | 40,4 | 1.2 |
Snout length | 20.9 | 19.7–24.5 | 22.2 | 1.3 | 20.1–23.6 | 21.9 | 1.2 |
Least interorbital distance | 38.7 | 33.3–38.7 | 35.4 | 1.5 | 35.2–39.9 | 37.1 | 1.4 |
Upper jaw length | 41.3 | 39.5–43.5 | 41.5 | 1.1 | 38.6–41.8 | 40.1 | 1.0 |
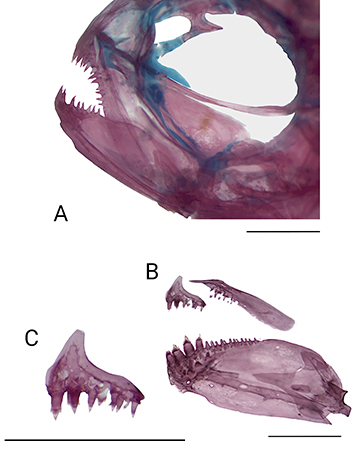
Mouth slightly sub-superior, with dentary protruding slightly over anterior tip of premaxilla. Posterior terminus of maxilla surpassing vertical through anterior margin of eye. Premaxillary teeth in single misaligned row (Fig. 3), with 7(8), 8*(6) or 9(6) tricuspid teeth, with median cusps slightly more developed and the third tooth usually positioned more externally. Maxilla with 6(3), 7(8), 8(4), 9*(1), or 10(2) teeth, the first tooth tricuspid, remaining teeth conical. Dentary with four tricuspid teeth, followed by eight to twelve smaller conical teeth.
FIGURE 3| Inpaichthys parauapiranga, paratype, CPUFMT 7891, 24.4 mm SL. A. Anterior region of cranium; B. Upper and lower jaws; C. Premaxillary; right side, lateral view. Scale bars = 1 mm.
Scales cycloid. Three to five radii, and inconspicuous circuli anteriorly. Lateral line incomplete, with 6(7) or 7*(14) perforated scales. Longitudinal scale series including lateral-line scales 31*(9), 32(1), 33(7), 34(2), or 35(1). Five* (20) scale rows between dorsal-fin origin and lateral line. Four* (20) scale rows between lateral line and pelvic fin insertion. Predorsal scales 11(6), 12(9), 13(4), or 14*(1). Circumpeduncular scales 12(3), 13(12), or 14*(2). Single row of 11–16 scales covering base of anal-fin rays. Presence of two rows of scales extending over caudal-fin lobes; squamation covering evenly both caudal lobes.
Supraneurals 7(7) or 8(1). Dorsal-fin rays iii, 9*(20), first unbranched dorsal-fin ray visible only in c&s specimens. Dorsal-fin origin at midpoint of standard length. First dorsal-fin pterygiophore located after neural spine of 11th vertebra (8). First unbranched ray about half the length of second ray. Adipose fin present. Pectoral-fin rays i,10(20), its tip typically falling short before reaching vertical through pelvic-fin insertion, reaching that point in few (5) specimens. Pelvic-fin rays i,6(3) or i,7*(17). Tip of adpressed pelvic fin not reaching origin of anal fin. Anal-fin rays iv(6), 21(1), 22(5), 23(5), 24*(9), or 25(1). First anal-fin pterygiophore inserted behind haemal spine of 15th vertebra (8). Caudal fin forked, lobes similar in size; principal caudal-fin rays i,9/8,i (20). First gill arch with 10(15) gill rakers on hypobranchial and ceratobranchial, 6(2) or 7(9) on epibranchial, and one on cartilage between ceratobranchial and epibranchial. Four (1) branchiostegals, 3(1) on anterior ceratohyal, and 1(1) on posterior ceratohyal. Vertebrae 31(1), 32(3), or 33(4).
Coloration in alcohol. Overall basic color pale yellow (Fig. 2A). Dark chromatophores concentrated on the dorsal surface of the head, from tip of the snout to end of supraoccipital spine, and extend posteriorly over the predorsal scales. Small, scattered, dark chromatophores at infraorbitals, preopercle and opercle bones. Two diffuse humeral spots separated by a clearer area. Anterior humeral blotch located at level of second and third perforated scales of lateral line and extends to about one or two scales rows above lateral line. Second humeral blotch located at sixth and/or seventh perforated scales of lateral line and extending to approximately one row of scales above lateral line. Four brown diffuse lateral stripes present in the flanks, from dorsal profile to midline region. Body sides above midline with chromatophores in middle part of scales, chromatophores absent on posterior edges of scales. Scattered chromatophores below median stripe, with chromatophores concentrated at the edges of myosepta of posterior half of lower body above anal fin. All fins without pigmentation. Immediately after preservation (Fig. 2B), two conspicuous rectangular-shaped humeral blotches present, being the anteriormost blotch more diffuse when compared to the second humeral blotch, and still retaining five to six red dotted longitudinal stripes along body sides. Dorsal, adipose, pectoral, pelvic and anal fins yellow-orange at distal region. Caudal yellowish at the proximal region. Dorsal, caudal, and anal fins with scattered dark chromatophores along interradial membranes.
Coloration in life. Description based on photographs taken in the field of two specimens (Fig. 4), and of a specimen photographed by Fernando C. P. Dagosta (MUBIO 113, SL undetermined). Overall body coloration silvery, grayish dorsally, with a reddish tint in the upper third of the facial bones, occupying the upper half of the operculum and infraorbital 3 to infraorbital 5 (apparently imparted by the transparency of these bones, exposing red pigmentation of the gills), upper margin of eye also red. Blue iridescence present on the flanks, longitudinally to opercle to caudal peduncle. Five red dotted longitudinal stripes along the body sides and a narrow iridescent midlateral stripe along longitudinal septum of body, extending from first humeral blotch to median caudal-fin rays. Dorsal, anal, adipose, pectoral, and pelvic fins yellow-orange (being orange specially concentrated at posterior half of pectoral and anal-fin rays in some specimens) (Fig. 4B), caudal fin yellowish. Tip of anal fin white, forming a stripe along anal-fin margin.
FIGURE 4| Inpaichthys parauapiranga. A. CPUFMT 7891, paratype, 23.1 mm SL. B. CPUFMT 7908, holotype, 25.4 mm SL, immediately after capture. Photos by Alexandre C. Ribeiro.
Sexual dimorphism. Contrary to what is observed for I. kerri and I. nambiquara (see Remarks of I. nambiquara), nosecondary sexual characters were detected on examined specimens of I. parauapiranga.
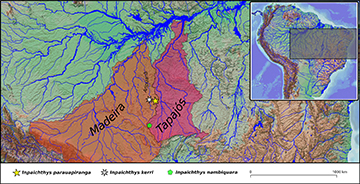
Geographical distribution. Inpaichthys parauapiranga is known from a tributary of the rio Canamã, a tributary of rio Aripuanã, itself a tributary of the rio Madeira, Amazon basin, Mato Grosso State, Brazil (Fig. 5).
FIGURE 5| Map of South America with a detailed view of the southwestern portion of the Amazon basin, showing the distribution of the three Inpaichthys species (I. kerri, I. nambiquara, and I. parauapiranga) in the rio Madeira basin and rio Tapajós basin.
Ecological notes. The type-locality is a narrow and shallow stream (approximately 2 m wide and 60 cm deep), with clear water, slow to moderately-flowing current, and a sandy bottom with abundant aquatic vegetation, where specimens were typically captured (Fig. 6). Stomach contents of four paratypes revealed mainly remnants of insect larvae.
FIGURE 6| Type-locality of Inpaichthys parauapiranga, tributary of the rio Canamã, rio Aripuanã basin, Mato Grosso State, Brazil.
Etymology. The specific epithet parauapiranga derives from Tupi language, paraua, meaning blotch and piranga, meaning red, and it is in allusion to its color pattern in living specimens. A noun in apposition.
Conservation status. Inpaichthys parauapiranga was collected at only one sampling site. Several specimens were collected at the type-locality, indicating that the species is relatively common at this site, which is currently moderately well preserved. However, the species is already being targeted by fish collectors operating for the aquarium hobby and unsustainable levels of extraction of wild specimens may pose a threat to the species in the future. Thus, we suggest that pending more efforts to establish the distributional range, population, and impacts from capture for the aquarium hobby, Inpaichthys parauapiranga should be classified as Data Deficient (DD), according to the International Union for Conservation of Nature (IUCN) categories and criteria (IUCN Standards and Petitions Subcommittee, 2022).
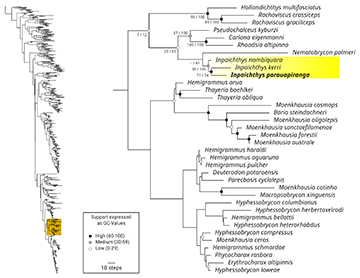
Remarks. The phylogenetic hypothesis herein presented was obtained in SEP64 (i.e., each character weighted according to its own homoplasy and using a K-value of 64, the mildest explored). In these conditions we found three most parsimonious trees of 70439 steps (5600 of them morphological) and fit = 508.27677 (Fig. 7). The monophyly of Inpaichthys, including I. nambiquara as the sister group a clade composed of I. kerri and I. parauapiranga, was obtained in all performed analyses. In SEP64 the monophyly of the genus is moderately well supported (GC =36), while the relationships of I. kerri and I. parauapiranga have better support (GC =71). In the final hypothesis, Nematobrycon palmeri Eigenmann, 1911was obtained as the sister group of Inpaichthys, but such relationship is weakly supported (negative GC value) and unstable among the most parsimonious trees of different analytical conditions (Freq = 41 and GC = -16 in the optimal trees of the explored conditions). Also with low support, we found as successive sister groups of Inpaichthys and Nematobrycon the clade composed of Carlana, Pseudochalceus, and Rhoadsia, and the group composed of Hollandichthys Eigenmann, 1909 and Rachoviscus Myers, 1926 (Fig. 7). The condition in which the number of morphological steps was lowest (5574) is SEP32. In the most parsimonious trees under SEP32 (70685 steps and fit = 738.45437), the relationships between Inpaichthys and Nematobrycon as the sister group of the clade of Carlana, Pseudochalceus, and Rhoadsia are the same than in SEP64. However, the sister group of this clade is composed of the genus Jupiaba Zanata, 1997 (J. ajuricaba (Marinho, Lima, 2009), J. anterior (Eigenmann, 1908), J. anteroides (Géry, 1965), J. polylepis (Günther, 1864), and J. poranga Zanata, 1997). In SEP64, we found eight morphological synapomorphies supporting the monophyly of Inpaichthys: nasal septum of mesethmoid formed by two parallel lamellae (9:1), absence of a dorsal expansion of rhinosphenoid projected medial to olfactory nerves (35:0), short supraoccipital spine covering dorsally only anterior axis of neural complex of Weberian apparatus (62:1), articulation between second and third infraorbitals anteroventrally angled (86:1), absence of anguloarticular portion of mandibular laterosensory canal (105:1), frontal and pterotic laterosensory canals forming an angle on pore receiving infraorbital canal (118:1), maxillary teeth extended approximately to half maxillary length (191:0), and presence of 24 or fewer branched anal-fin rays (420:0). The sister-group relationship of I. kerri and I. parauapiranga is supported by three morphological synapomorphies: three cusps on anterior dentary teeth (201:0), metapterygoid foramen as a simple rounded opening (228:0), and first anal-fin pterygiophore located ventral to last dorsal-fin pterygiophores (416:1).
FIGURE 7| Phylogenetic relationships of Inpaichthys in the tribe Stethaprionini, as obtained from the phylogenetic analysis under parsimony and extended implied weighting (SEP; K = 64; fit = 508.27677; length = 70435 steps). Support expressed as GC values (differences of frequencies “group present/contradicted”). Numeric values of support are shown only for the clade including Inpaichthys. Branches length represent the number of synapomorphies.
Discussion
The phylogenetic position of the, until now, monotypic genus Inpaichthys was a subject of debate ever since its original description. Géry, Junk (1977:418) considered the genus “difficult to position”. These authors considered Inpaichthys kerri as similar and possibly related to Coelurichthys tenuis Nichols, 1913 [a synonym of Mimagoniates lateralis (Nichols, 1913)], a species currently included in the subfamily Stevardiinae. They also surmised putative relationships with the genera Nematobrycon (based mostly on color pattern), Rachoviscus, and Bryconella Géry, 1965 (the latter based on a similar arrangement of the premaxillary teeth).
The first study to include I. kerri within a phylogenetic framework was Calcagnotto et al. (2005) who, based on molecular data, recovered the species in a group with representatives of the genera Astyanax Baird & Girard, 1854; Astyanacinus Eigenmann, 1907; Moenkhausia Eigenmann, 1903; Hemigrammus Gill, 1858; and Hyphessobrycon Durbin, 1908. Mirande (2010) proposed the first phylogenetic hypothesis based only on morphological data that includes I. kerri, in which the species was considered to be a member of the subfamily Aphyocharacinae, as the sister group of the remaining members of the subfamily (i.e., the genera Aphyocharax Günther, 1868; Leptagoniates Boulenger, 1887; Paragoniates Steindachner, 1876; Prionobrama Fowler, 1913; Phenagoniates Eigenmann & Wilson, 1914; Rachoviscus; and Xenagoniates Myers, 1942). However, subsequent molecular analyses (Javonillo et al., 2010; Thomaz et al., 2010; Oliveira et al., 2011) did not confirm I. kerri as a member of the Aphyocharacinae, but instead, proposed it to be more related to taxa classified in Astyanacinus; Bario Myers, 1940; Ctenobrycon Eigenmann, 1908; Hyphessobrycon; and Moenkhausia. Particularly, Oliveira et al. (2011) did not recover a subfamily Aphyocharacinae with the composition proposed by Mirande (2010), but rather they obtained Inpaichthys as the sister group of Bario, Rachoviscus, and Hollandichthys. Tagliacollo et al. (2012) corroborated that Inpaichthys [as well as Rachoviscus and Protocheirodon pi (Vari, 1978)] did not belong to the subfamily Aphyocharacinae, but instead, to a clade comprised of Astyanacinus, Carlana, and Nematobrycon. Finally, Mirande (2019) published a comprehensive phylogenetic analysis of Characidae combining morphological and molecular data, in which I. kerri was recovered as belonging to the tribe Rhoadsiini within the subfamily Stethaprioninae, as the sister group of Nematobrycon palmeri, Pseudochalceus kyburzi Schultz, 1966, Rhoadsia altipinna Fowler, 1911, and Carlana eigenmanni Meek, 1912. Mirande et al. (2023) found a sister-group relationship of Inpaichthys kerri and the clade of Hollandichthys and Rachoviscus but, in that article, Carlana, Nematobrycon, Pseudochalceus, and Rhoadsia were not included. Our hypothesis, thus, corroborates the findings of Oliveira et al. (2011), Mirande (2019), and Mirande et al. (2023), in a more comprehensive analysis including, among other species, the three currently known species of the genus.
Weitzman, Vari (1988) circumscribed the term “miniature fish” to species that do not exceed 26 mm SL and that reach sexual maturity before 20 mm SL. Additionally, miniature fishes have in common numerous apparently paedomorphic morphological reductions, including the lower degree of development of the laterosensory canal system of the head and the body, reduction in the number of the fin rays, body scales, and loss of cranial bones which contribute to an overall simplification of the skeleton (Weitzman, Vari, 1988:445). Although Inpaichthys species are slightly larger than the arbitrary cut-off standard length (I. kerri: 28 mm SL; I. nambiquara: 27.0 mm SL; I. parauapiranga: 26.3 mm SL) proposed by Weitzman, Vari (1988) and still in use in the literature (e.g., Toledo-Piza et al., 2014), they exhibit some reductive features, such as the absence of the fourth and sixth infraorbitals (the fourth is present but much reduced in I. nambiquara), and an interrupted lateral line.
Despite Inpaichthys being composed of species with restricted distributional ranges, the discovery of I. parauapiranga and the new generic assignment of I. nambiquara imply that the genus Inpaichthys as a whole, now exhibits a relatively broad distribution across the northwestern portion of the Brazilian Shield, in both the upper rio Madeira (rio Aripuanã basin) and the adjacent upper rio Tapajós basin (rio Juruena basin). The sister-group relationship between I. parauapiranga and I. kerri is congruent with the fact that both species occur relatively close to each other (ca. 60 km in a straight line), within the rio Aripuanã basin. On the other hand, I. nambiquara occurs at the westernmost portion of the rio Juruena basin, roughly 300 km southwest from the type-locality of I. parauapiranga (Fig. 5). There are additional undescribed species of Inpaichthys occurring in the same general area (F. C. P. Dagosta, 2023, pers. comm.), which, once described, will add more information to this increasingly complex and interesting biogeographic scenario.
Comparative material examined. Inpaichthys kerri: All from Brazil, Mato Grosso State, rio Aripuanã basin: CPUFMT 4869, 5, 22.1–23.9 mm SL (2 c&s, 21.0–25.5 mm SL); CPUFMT 4897, 2, 18.9–20.29 mm SL; CPUFMT 4933, 4, 16.3–19.6 mm SL; CPUFMT 4988, 3, 22.7–27.0 mm SL; CPUFMT 5001, 1, 12.3 mm SL; CPUFMT 5020, 2, 23.2–24.5 mm SL; CPUFMT 5038, 4, 17.2–19.4 mm SL; CPUFMT 5066, 5, 20.5–20.7 mm SL; CPUFMT 5083, 2, 18.5–20.1 mm SL; ZUEC 9775, 62, 9.5–27.9 mm SL (11 c&s, 16.2–21.8 mm SL); ZUEC 13433, 1, 14.5 mm SL. Specimens of other characiform genera, examined and coded for the phylogenetic analysis, are listed in Terán et al. (2020), with additions in Lima et al. (2021) and Ferreira et al. (2023).
Acknowledgments
KMF, DCF and ACR received financial support from Universidade Federal de Mato Grosso – Edital de Apoio à Pesquisa (2021). JMM is funded by FONCyT (PICT–2020–02141), CONICET (PIP–11220200101214), and Fundación Miguel Lillo (Z–0085). We are grateful to Carlos A. S. Lucena (MCP) for loaning material of Inpaichthys nambiquara, to Hans-Georg- Evers for providing the Fig. 1 and comments on the sexual dimorphism of I. nambiquara, and to Fernando C. P. Dagosta (MUBIO) and Vinicius A. Bertaco (MCN) for discussions that helped shaping some of the points herein studied.
References
Aljanabi SM, Martinez I. Universal and rapid salt-extraction of high quality genomic DNA for PCR-based techniques. Nucleic Acids Res. 1997; 25(22):4692–93. https://doi.org/10.1093/nar/25.22.4692
Bertaco VA, Malabarba LR. A new species of Hasemania from the upper rio Tapajós drainage, Brazil (Teleostei: Characiformes: Characidae). Copeia. 2007; 2007(2):350–54. https://doi.org/10.1643/0045-8511(2007)7[350:ANSOHF]2.0.CO;2
Calcagnotto D, Schaefer SA, DeSalle R. Relationships among characiform fishes inferred from analysis of nuclear and mitochondrial gene sequences. Mol Phyl Evol. 2005; 36(1):135–53. https://doi.org/10.1016/j.ympev.2005.01.004
Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004; 32:1792–97. https://doi.org/10.1093/nar/gkh340
Ferreira KM, Lima FCT, Ribeiro AC, Flausino Jr. N, Machado FA, Mirande JM. A new species of Astyanax (Characiformes, Characidae) from the rio Apiacás, rio Teles Pires basin, with a discussion on its phylogenetic position. Can J Zool. 2023; 101(7):522–29. https://doi.org/10.1139/cjz-2022-0153
Fink WL, Weitzman SH. The so-called cheirodontin fishes of Central America with descriptions of two new species (Pisces: Characidae). Smithson Contrib Zool. 1974; 172:1–46. https://doi.org/10.5479/si.00810282.172
Géry J, Junk WJ. Inpaichthys kerri n. g. n. sp., um novo peixe caracídeo do alto rio Aripuanã, Mato Grosso, Brasil. Acta Amazon. 1977; 7(3):417–22.
Goloboff PA. Estimating character weights during tree searches. Cladistics. 1993; 9(1):83–91. https://doi.org/10.1111/j.1096-0031.1993.tb00209.x
Goloboff PA. Extended implied weighting. Cladistics. 2016; 30(3):260–272. https://doi.org/10.1111/cla.12047
Goloboff PA, Morales ME. TNT version 1.6, with a graphical interface for MacOS and Linux, including new routines in parallel. Cladistics. 2023, 39(2):144–53. https://doi.org/10.1111/cla.12524
International Union for Conservation of Nature (IUCN). Standards and petitions committee. Guidelines for using the IUCN Red List categories and criteria. Version 15 [Internet]. Gland; 2022. Available from: http://www.iucnredlist.org/documents/RedListGuidelines.pdf
Javonillo R, Malabarba LR, Weitzman SH, Burns JR. Relationships among major lineages of characid fishes (Teleostei: Ostariophysi: Characiformes), based on molecular sequence data. Mol Phyl Evol. 2010; 54(2):498–511. https://doi.org/10.1016/j.ympev.2009.08.026
Larsson A. AliView: a fast and lightweight alignment viewer and editor for large datasets. Bioinformatics. 2014; 30(22):3276–78. https://doi.org/10.1093/bioinformatics/btu531
Lima FCT, Caires RA, Conde-Saldaña CC, Mirande JM, Carvalho FR. A new miniature Pristella (Actinopterygii: Characiformes: Characidae) with reversed sexual dimorphism from the rio Tocantins and rio São Francisco basins, Brazil. Can J Zool. 2021; 99(5):339–48. https://doi.org/10.1139/cjz-2020-0241
Menezes NA, Weitzman SH. Two new species of Mimagoniates (Teleostei: Characidae: Glandulocaudinae), their phylogeny and biogeography and a key to the glandulocaudin fishes of Brazil and Paraguay. Proc Biol Soc Wash. 1990; 103:380–426.
Mirande JM. Weighted parsimony phylogeny of the family Characidae (Teleostei: Characiformes). Cladistics. 2009; 25(6):574–613. https://doi.org/10.1111/j.1096-0031.2009.00262.x
Mirande JM. Phylogeny of the family Characidae (Teleostei: Characiformes): from characters to taxonomy. Neotrop Ichthyol. 2010; 8(3):385–568. https://doi.org/10.1590/S1679-62252010000300001
Mirande JM. Morphology, molecules and the phylogeny of Characidae (Teleostei, Characiformes). Cladistics. 2019; 35(3):282–300. https://doi.org/10.1111/cla.12345
Mirande JM, Jerep FC, Vanegas-Ríos JA. Phylogenetic relationships of the enigmatic Carlastyanax aurocaudatus (Eigenmann) with remarks on the phylogeny of the Stevardiinae (Teleostei: Characidae). Neotrop Ichthyol. 2013; 11(4):747–66. https://doi.org/10.1590/S1679-62252013000400003
Mirande JM, Baicere-Silva CM, Santana JCO, Quagio-Grassiotto I. Sperm phylogeny of Characidae (Teleostei, Characiformes). Zool Scripta. 2023; 52(2):117–35. https://doi.org/10.1111/zsc.12577
Ohara WM, Loeb MV. Ichthyofauna of the upper Juruena river on Chapada dos Parecis, Mato Grosso, Brazil. Biota Neotrop. 2016; 16(4):e20160224. https://doi.org/10.1590/1676-0611-BN-2016-0224
Ohara WM, Mirande JM, Lima FCT. Phycocharax rasbora, a new genus and species of Brazilian tetra (Characiformes: Characidae) from Serra do Cachimbo, rio Tapajós basin. PLoS ONE. 2017; 12(2):e0170648. https://doi.org/10.1371/journal.pone.0170648
Ohara WM, Teixeira TF, Albornoz-Garzón JG, Mirande JM, Lima FCT. Hyphessobrycon rheophilus, a new species from rapids of the Amazon and Orinoco river basins (Characiformes: Characidae: Stethaprioninae). Zootaxa. 2019; 4712(4):561–75. https://doi.org/10.11646/zootaxa.4712.4.5
Oliveira C, Avelino GS, Abe KT, Mariguela TC, Benine RC, Ortí G, Vari RP, Castro RMC. Phylogenetic relationships within the speciose family Characidae (Teleostei: Ostariophysi: Characiformes) based on multilocus analysis and extensive ingroup sampling. BMC Evol Biol. 2011; 11:275. https://doi.org/10.1186/1471-2148-11-275
Palumbi SR. Nucleic acids II: the polymerase chain reaction. In: Hillis D, Moritz C, Mable B, editors. Molecular systematics. Massachusetts: Sinauer Associates Inc.; 1996. p.205–47.
Pastana MNL, Dagosta FCP, Esguícero ALH. A new sexually dichromatic miniature Hyphessobrycon (Teleostei: Characiformes: Characidae) from the rio Formiga, upper rio Juruena basin, Mato Grosso, Brazil, with a review of sexual dichromatism in Characiformes. J Fish Biol. 2017; 91(5):1301–18. https://doi.org/10.1111/jfb.13449
Sabaj MH. Codes for natural history collections in ichthyology and herpetology. Copeia. 2020; 108(3):593–669. https://doi.org/10.1643/ASIHCODONS2020
Tagliacollo VA, Souza-Lima R, Benine RC, Oliveira C. Molecular phylogeny of Aphyocharacinae (Characiformes, Characidae) with morphological diagnoses for the subfamily and recognized genera. Mol Phyl Evol. 2012; 64(2):297–307. https://doi.org/10.1016/j.ympev.2012.04.007
Taylor WR, Van Dyke GC. Revised procedures for clearing and staining small fishes and other vertebrates for bone and cartilage study. Cybium. 1985; 9(2):107–19.
Terán GE, Benitez MF, Mirande JM. Opening the trojan horse: phylogeny of Astyanax, two new genera and resurrection of Psalidodon (Teleostei: Characidae). Zool J Linn Soc. 2020; 190(4):1217–34. https://doi.org/10.1093/zoolinnean/zlaa019
Thomaz AT, Arcila D, Ortí G, Malabarba LR. Molecular phylogeny of the subfamily Stevardiinae Gill, 1858 (Characiformes: Characidae): classification and the evolution of reproductive traits. BMC Evol Biol. 2015; 15:146. https://doi.org/10.1186/s12862-015-0403-4
Thomaz AT, Malabarba LR, Bonatto SL. The phylogenetic placement of Hollandichthys Eigenmann 1909 (Teleostei: Characidae) and related genera. Mol Phyl Evol. 2010; 57(3):1347–52. https://doi.org/10.1016/j.ympev.2010.10.006
Toledo-Piza M, Mattox GMT, Britz R. Priocharax nanus, a new miniature characid from the rio Negro, Amazon basin (Ostariophysi: Characiformes), with an updated list of miniature Neotropical freshwater fishes. Neotrop Ichthyol. 2014; 12(2):229–46. https://doi.org/10.1590/1982-0224-20130171
Vari RP, Harold AS. Neotropical fishes of the genus Creagrutus (Teleostei: Ostariophysi: Characiformes): a phylogenetic study and a revision of the species east of the Andes. Smithson Contrib Zool. 2001; 612:1–239. https://doi.org/10.5479/si.00810282.613
Ward RD, Zemlak TS, Innes BH, Last PR, Hebert PDN. DNA barcoding Australia’s fish species. Philos Trans R Soc Lond B. 2005; 360(1462):1847–57. https://doi.org/10.1098/rstb.2005.1716
Weitzman SH. The osteology of Brycon meeki, a generalized characid fish, with an osteological definition of the family. Stanford Ichthyol Bull. 1962; 8(1):1–77.
Weitzman SH, Vari RP. Miniaturization in South American freshwater fishes; an overview and discussion. Proc Biol Soc Wash. 1988; 101:444–65.
Authors
![]() Katiane M. Ferreira1,
Katiane M. Ferreira1, ![]() Alexandre C. Ribeiro1,
Alexandre C. Ribeiro1, ![]() Flávio C. T. Lima2,
Flávio C. T. Lima2, ![]() Hugmar P. da Silva1,
Hugmar P. da Silva1, ![]() Daniela C. Ferreira1 and
Daniela C. Ferreira1 and ![]() Juan Marcos Mirande3
Juan Marcos Mirande3 ![]()
[1] Instituto de Biociências, Departamento de Biologia e Zoologia, Universidade Federal de Mato Grosso, Av. Fernando Corrêa, 2367, Boa Esperança, 78060-900 Cuiabá, MT, Brazil. (KMF) kmferreira@gmail.com, (ACR) alexandrecunharibeiro@gmail.com, (HPS) painsbio@gmail.com, (DCF) ferreiradc@gmail.com.
[2] Museu de Diversidade Biológica, Instituto de Biologia, Universidade Estadual de Campinas, Caixa Postal 6109, 13083-863 Campinas, SP, Brazil. (FCTL) fctlima@gmail.com.
[3] Fundación Miguel Lillo, Unidad Ejecutora Lillo (CONICET-FML), Miguel Lillo 251, San Miguel de Tucumán, 4000, Argentina. (JMM) jmmirande@lillo.org.ar (corresponding author).
Authors’ Contribution 

Katiane M. Ferreira: Conceptualization, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing-original draft, Writing-review and editing.
Alexandre C. Ribeiro: Conceptualization, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing-original draft, Writing-review and editing.
Flávio C. T. Lima: Conceptualization, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing-original draft, Writing-review and editing.
Hugmar P. da Silva: Data curation, Formal analysis, Investigation, Methodology, Resources, Software, Writing-original draft.
Daniela C. Ferreira: Conceptualization, Data curation, Formal analysis, Methodology, Resources, Software, Writing-original draft.
Juan Marcos Mirande: Conceptualization, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Software, Supervision, Validation, Visualization, Writing-original draft, Writing-review and editing.
Ethical Statement
The specimens were collected under a collection permit authorized by Sistema de Autorização e Informação em Biodiversidade (SISBIO processes #18454–2 and 71172–2).
Competing Interests
The author declares no competing interests.
How to cite this article
Ferreira KM, Ribeiro AC, Lima FCT, Silva HP, Ferreira DC, Mirande JM. A new species of Inpaichthys from the rio Canamã, rio Aripuanã basin, Mato Grosso State, Brazil, with a redefinition of the genus (Characidae: Stethaprioninae). Neotrop Ichthyol. 2024; 22(1):e230113. https://doi.org/10.1590/1982-0224-2023-0113
Copyright
This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Distributed under
Creative Commons CC-BY 4.0

© 2024 The Authors.
Diversity and Distributions Published by SBI
![]() Accepted November 14, 2023 by Fernando Carvalho
Accepted November 14, 2023 by Fernando Carvalho
![]() Submitted October 16, 2023
Submitted October 16, 2023
![]() Epub February 19, 2024
Epub February 19, 2024