![]() Dhiego G. Ferreira1,
Dhiego G. Ferreira1, ![]() Bruno A. Galindo1,
Bruno A. Galindo1, ![]() Tais C. de Souza1,
Tais C. de Souza1, ![]() Leonardo B. Pereira1,
Leonardo B. Pereira1, ![]() Victor A. P. Bernardes1,
Victor A. P. Bernardes1, ![]() Ana J. C. Marques1
Ana J. C. Marques1 ![]() ,
, ![]() Wilson Frantine-Silva1,
Wilson Frantine-Silva1, ![]() Thais Kotelok-Diniz2,
Thais Kotelok-Diniz2, ![]() Carlos E. G. Aggio1,
Carlos E. G. Aggio1, ![]() Caroline Apolinário-Silva2,
Caroline Apolinário-Silva2, ![]() Augusto S. Zanatta1 and
Augusto S. Zanatta1 and ![]() Silvia H. Sofia2
Silvia H. Sofia2
PDF: EN XML: EN | Supplementary: S1 S2 | Cite this article
Abstract
Cnesterodon hypselurus is a small fish that has a restricted distribution in southern Brazil, including headwaters of the Tibagi and Itararé river basins (Upper Paraná River). This study reported C. hypselurus in a headwater of Cinzas River basin, where there were no previous records of this species, and employed microsatellite loci and mitochondrial haplotypes in a population genetic analysis. A total of 57 specimens was analyzed, including 30 from Cinzas River basin, 25 from Itararé River basin and two from Tibagi River basin. Results indicated low genetic diversity levels (HE = 0.334 and h = 0.246) for the sample from Cinzas River, suggesting reflections of a founder effect after the species had dispersed from one watershed to another, possibly by headwater captures. Since different populations were detected between the Cinzas and Itararé rivers (DEST = 0.248,P-value < 0.05) and other occurrence sites are still unknown in the Cinzas River basin, the data herein have great relevance and should be taken into account in future management and conservation actions, as well as in evolutionary studies of C. hypselurus.
Keywords: D-Loop, Guarú, Microsatellite, Paraná River, Population genetics.
Cnesterodon hypselurus é um pequeno peixe que possui distribuição restrita no sul do Brasil, incluindo cabeceiras das bacias dos rios Tibagi e Itararé (alto rio Paraná). Este estudo reportou C. hypselurus na cabeceira da bacia do rio das Cinzas, onde não havia registros prévios desta espécie, e empregou locos microssatélites e haplótipos mitocondriais em uma análise genética de populações. Um total de 57 espécimes foi analisado, incluindo 30 do rio das Cinzas, 25 da bacia do rio Itararé e dois da bacia do rio Tibagi. Os resultados indicaram baixos níveis de diversidade genética (HE = 0,334 e h = 0,246) para a amostra do rio das Cinzas, sugerindo reflexos de um efeito fundador após a espécie ter dispersado de uma bacia para a outra, possivelmente a partir de captura de cabeceiras. Uma vez que diferentes populações foram detectadas entre os rios das Cinzas e Itararé (DEST = 0,248,valor de P < 0,05) e que outros pontos de ocorrência ainda são desconhecidos na bacia do rio das Cinzas, os dados do presente estudo mostram grande relevância e deveriam ser considerados em futuras ações de manejo e conservação, bem como em estudos evolutivos de C. hypselurus.
Palavras-chave: D-Loop, Genética de Populações, Guarú, Microssatélites, rio Paraná.
Introduction
The growing number of endangered species has been one of the main concerns to the planet’s biodiversity (Frankham et al., 2010; Allendorf et al., 2012) and this is no different for ichthyofauna in freshwater systems (Geist, 2011; Ahmed et al., 2022). In Neotropical region, for example, about 18% of all freshwater fish species is already represented by threatened or potentially endangered species (Tagliacollo et al., 2021), including, for the most part, small-sized fish species (Castro, Polaz, 2020). This is mainly due to intense interference from anthropic actions, including overfishing, pollution, aquatic contamination, river fragmentation by hydroelectric plants and the insertion of non-native species (Agostinho et al., 2016; Arya, 2021). Thus, studies that identify the occurrence areas of these species are increasingly urgent and necessary, including analyses of their population distributions and their genetic diversity levels (Geist, 2011; ICMBio, 2018). In fact, genetic analyses have been of great importance for the management and conservation of threatened species (Landweber, Dobson, 2021; Willi et al., 2022), since the levels of genetic diversity of populations are determinant for the evolutionary/adaptive potential of the species (Freeland, 2005; Hartl, Clark, 2007).
Cnesterodon hypselurus Lucinda & Garavello, 2001, popularly known as guarú, is a small-sized fish that has a restricted distribution in the Paraná River basin (Shibatta et al., 2002; Silva et al., 2015), one of the main hydrographic systems of the Neotropical region (Reis et al., 2016). This species was listed as endangered until the end of the last decade, however, it is currently placed in the Least Concern (LC) category (Silva et al., 2015; ICMBio, 2023). This fish belongs to the family Poeciliidae, order Cyprinodontiformes, and has a restricted occurrence, currently limited to the head-waters of some small drainages in the Tibagi and Itararé rivers basins, Upper Paraná River basin (Lucinda, 2005; Silva et al., 2015). However, the present study reports a new record of C. hypselurus in a head-water stream of the Cinzas River basin, a drainage system where this species had not yet been reported. Fish of the Poeciliidae family are characterized by small-sized body, including, so far, 273 valid species (in 27 genera) along Neotropical hydrographic systems (Fricke et al., 2022). These species are in a wide variety of levels in trophic networks, from primary consumer to primary and secondary predators, eating benthic algae, insect larvae, plant material or even plankton, while they are the food of several piscivorous species (Quintans et al., 2009, 2010). Interestingly, some species are also reported in biological control of mosquito larvae (Quintans et al., 2010).
Adults of C. hypselurus, in particular, can reach about 3 cm in standard length, showing internal fertilization and reproduction by viviparity (Lucinda, 2003). They are commonly found living in the clear waters of higher altitude stretches in streams (Shibatta et al., 2002; Silva et al., 2015), where they show a low displacement range throughout life, like other small-sized fish species in Neotropical streams (Castro, 1999). Before the present study, the last record of C. hypselurus was in the Arroio do Gica stream, tributary of the Tibagi River basin, extending the estimated occurrence area of this species to 3,989 km² (Silva et al., 2015), including five confirmed occurrence sites in Tibagi River basin and three in Itararé River basin (Tab. S1). Although the Tibagi and Itararé river basins have several head-water streams geographically close to the C. hypselurus occurrence site recorded in Cinzas River basin, these drainages are not currently connected and population genetic differences would be expected among their head-waters. Furthermore, recent studies report that some Neotropical fish can show genetic structuring even in streams with good connectivity levels, including population differentiation in short geographical distances along drainages and among tributaries of a same watershed (Ferreira et al., 2016; Apolinário-Silva et al., 2021), which is a very important knowledge for conservation strategies that aim to preserve as much of the genetic diversity of species as possible (Frankham et al., 2010).
Small-sized fish species that inhabit head-water streams tend to be highly endemic and to pass their entire life cycles in restricted areas (Castro, 1999; Castro, Polaz, 2020). Thus, identifying small and isolated populations, as well as their genetic diversity levels, is another aspect of great importance for conservation, since theses populations are more likely to suffer genetic diversity losses (Freeland, 2005). In Cinzas River basin, several previous samplings conducted along the main channel and tributaries have not reported C. hypselurus anywhere else, suggesting that populations of this species could be small and restricted to the upper reaches of the drainage system. Thus, knowing the genetic diversity of C. hypselurus in the occurrence site recorded herein is an essential step in the attempt to conserve this species within the basin, also allowing clues about the populational evolutionary history, since ancient geological events could have contributed to headwater captures and fauna exchanges among the Cinzas, Tibagi and Itararé river basins (Ribeiro, 2006; Frota et al., 2020).
The present study records the occurrence of C. hypselurus in Cinzas River basin and use microsatellite and mitochondrial (mtDNA D-loop region) markers to investigate the genetic diversity and population structure of this species in the new location and in some other occurrence site from neighboring watersheds.
Material and methods
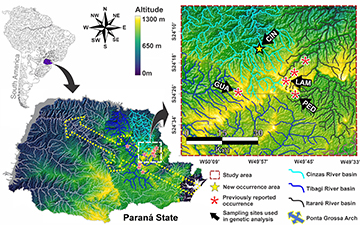
Study area and fish sampling. Cinzas River is one of the main tributaries of the Paranapanema River basin (Upper Paraná River), draining 9,658.8 km2 entirely within Paraná State, Brazil (Britto et al., 2003; Vianna, Nogueira, 2008). Its headwaters are in the Serra de Furnas (24°27’34”S 49°55’49”W), near the area influenced by Ponta Grossa Arch, and its main channel extends 371 km, flowing in a south-north direction to the confluence (23°01’51”S 50°26’43”W) with the Paranapanema River (Maack, 2002) (Fig. 1).
FIGURE 1| Distribution of Cnesterodon hypselurus occurrence locations and the sampling sites used in the genetic study. CIN – new occurrence record in Cinzas River basin (unnamed stream), LAM – Lambari and PED – Pedrinhas streams in Itararé River basin and GUA – Guaricanga stream in Tibagi River basin (Source: modified from Silva et al., 2015; Franco-Magalhaes et al., 2010, and Google Earth, 2018, https://www.google.com.br/maps).
The Ponta Grossa Arch (PGA), in particular, involves a large deformational structure with northwest-southeast trend axis, including two large trending Early Cretaceous magmatic dike swarms, possibly related to the break-up of the southwestern Gondwana. The highest points in PGA are in the center and southeastern (>1,500 m), while the lower regions are in the northwest (around 600 m) (Franco-Magalhaes et al., 2010).
The occurrence site of C. hypselurus recorded herein (CIN – 24°13’45”S 49°56’49”W) is in the lower section of a small head-water stream (unnamed), showing little flow and small pools, rocky and sandy substrate, an average depth of 0.40 m and an average width of approximately 0.90 m. The sampled stretch is crossed by a bridge and rural road, while the upstream and downstream sections cross areas that includes riparian vegetation bordered by cultivation of exotic trees (Pinus sp.).
Genetic analysis was conducted on 57 specimens, including 30 from the new occurrence site in the Cinzas River basin (CIN), 20 from Lambari stream (LAM) and five from Pedrinhas stream (PED) – both from the Itararé River basin, and two from Guaricanga stream (GUA) in Tibagi River basin (Fig. 1; Tab. S1). Considering the sample size sufficient for population studies, only CIN and LAM were included in the microsatellite analysis, while mtDNA analyses were conducted for all specimens.
CIN and LAM samples were obtained in three sampling expeditions, conducted from April 2020 to November 2020 (three per sample site), using sieve and seines during one hour of collection efforts. These expeditions also included sampling sites in other drainages where C. hypselurus was previously reported, however, specimens were not obtained in these locations. Meanwhile, GUA (MZUEL 14641, 2 ex) and PED (MZUEL 15264, 5 ex) were provided by the Zoology Museum of the State University of Londrina (Tab. S1). Tissue samples were placed in 70% alcohol and stored at -20°C. Voucher specimens were deposited in the Museu de Zoologia da Universidade Estadual de Londrina (accession numbers: MZUEL 22693, 6 ex, and 22694, 16 ex) (Tab. S1).
TABLE 1 | Genetic diversity of Cnesterodon hypselurus (from microsatellite loci and mtDNA) in a new occurrence area herein reported in the Cinzas River basin (CIN) and in sites from other watersheds, including Lambari (LAM) and Pedrinhas (PED) streams in Itararé River basin and Guaricanga stream (GUA) in Tibagi River basin. N = number of individuals analyzed, Ne = effective population size, A = total number of alleles found per sample, NP = number of private alleles, RA = allelic richness, NA = mean number of alleles, NE = mean number of effective alleles, HO = observed heterozygosity, HE = expected heterozygosity, FIS = rate of inbreeding, HWE = loci deviated from Hardy–Weinberg proportions, Nh = number of haplotypes found, h = haplotype diversity, π = nucleotide diversity, D = Tajima’s neutrality test (Tajima, 1989), Fs = Fu neutrality test (Fu, 1997). *Significant values – P value < 0.05.
Sample | Microsatellite | ||||||||||||||
N | Ne (CI 95%) | A | NP | RA | NA | NE | HO | HE | FIS | HWE | |||||
CIN | 30 | 16.3 (5.1-118.7) | 22 | 4 | 2.606 | 2.750 | 1.771 | 0.349 | 0.334 | 0.025 | 3 Pvm11, Pvm14, Pvm18 | ||||
LAM | 20 | 81.7 (24-∞) | 52 | 34 | 6.450 | 6.500 | 3.692 | 0.446 | 0.528 | 0.183* | 2 Pvm11, Pvm14 | ||||
General | 50 | – | 56 | – | – | 7.000 | 2.471 | 0.384 | 0.477 | 0.205* | All loci | ||||
D-Loop | |||||||||||||||
N | Nh | h | π | D | Fs | ||||||||||
CIN | 30 | 3 | 0.246 | 0.0005 | -1.022 | -1.206 | |||||||||
LAM | 20 | 3 | 0.358 | 0.0007 | -0.768 | -0.723 | |||||||||
PED | 5 | 2 | 0.800 | 0.0020 | -0.816 | 0.090 | |||||||||
GUA | 2 | 2 | 1.000 | 0.0040 | 0.000 | 0.693 | |||||||||
General | 57 | 7 | 0.398 | 0.0011 | -0.386 | -0.428 | |||||||||
Microsatellite loci and mitochondrial haplotypes (D-loop). Genomic DNA was extracted from muscle tissue or rayed fins and subsequently quantified and diluted following the steps in Apolinário-Silva et al. (2021). Microsatellite loci analysis was conducted using eight pairs of primers (Pvm5, Pvm7, Pvm8, Pvm10, Pvm11, Pvm14, Pvm16 and Pvm18) described for Poecilia vivipara (Tonhatti et al., 2014), another species of the Poeciliidae family, which were evaluated and selected from cross-amplification tests. Polymerase Chain Reactions (PCR) were performed in a final volume of 10μL, using the concentrations and conditions employed by Apolinário-Silva et al. (2018), including the method from Schuelke (2000) for labeling PCR products with fluorescence (FAM, HEX, NED and PET), aiming at further genotyping in an automatic sequencer ABI-PRISM 3500 xL (Applied Biosystems), using GeneScan 600 Liz (Applied Biosystems) as a molecular weight marker.
Part of the D-Loop region of the mitochondrial DNA (mtDNA) was amplified for all samples using the primers L 5’-AGAGCGTCGGTCTTGTAAACC-3’ (Cronin et al., 1993) and H 5’-CCTGAAGTAGGAACCAGATG-3’ (Meyer et al., 1990) and the reagent concentration, as well as PCR conditions employed by Apolinário-Silva et al. (2021). PCR products were purified using EXOSap IT® (Prodimol) and sequenced from the Big Dye Terminator v 3.1 kit (Applied Biosystems), according to the manufacturer’s instructions. Sequence reading was performed on an ABI-PRISM 3500 XL automatic sequencer (Applied Biosystems), followed by editing and alignment in the MEGA 5.0 program (Tamura et al., 2011).
Molecular data analysis. Genetic diversity estimates from mtDNA data were obtained using the software: DnaSP v.5 (Librado, Rozas, 2009) to calculate the number of haplotypes, haplotype diversity (h) and nucleotide diversity (π); Network 4.6.1.1 (Fluxus Technology Ltd – http://www.fluxus-engineering.com) to obtain a haplotype network based on the median-joining algorithm (Bandelt et al., 1999). Aiming to investigate the demographic history, Tajima’s D (Tajima, 1989) and Fu’s Fs (Fu, 1997) statistics were estimated using Arlequin v. 3.5.1.3 (Excoffier, Lischer, 2010), while pairwise mismatch distributions were obtained with DnaSP v.5 (Librado, Rozas, 2009).
On microsatellite data, Micro-Checker v. 2.2.1 (Van Oosterhout et al., 2004) was employed to analyze missing data and to evaluate the presence of null alleles. Genepop v.1.2 (Raymond, Rousset, 1995) was employed to test deviations from the HWE (Hardy-Weinberg Equilibrium) and linkage disequilibrium (LD), using sequential Bonferroni corrections (Rice, 1989) to adjust the alpha values. The number of alleles (A), mean alleles per locus (NA), number of effective alleles (NE), number of private alleles (NP), and expected (HE) and observed (HO) heterozygosity were obtained using Popgen v. 1.31 (Yeh et al., 1999), while Fstat v. 2.9.3 (Goudet, 2001) was used to evaluate the allelic richness (RA), using a rarefaction approach (corrected for a minimum sample size of 20 diploid individuals, and significant inbreeding (FIS) (P < 0.05). NeEstimator v. 2.0 (Do et al., 2014) was used to estimate the effective population size (Ne) for each population, applying a random mating model and PCrit of 0.02 to allele frequencies to Linkage Disequilibrium (LD) method (Waples, Do, 2008).
Signs of recent population bottlenecks were investigated in Bottleneck v. 1.2.02 program (Piry et al., 1999), using two tests to evaluate deviations from the mutation-drift equilibrium: i – Wilcoxon sign-rank test (Luikart, Cornuet, 1998) to investigate significant excess heterozygosity (bottleneck sign) on the Infinite Alleles Model (IAM), Two-Phase Model (TPM, with 90 % SMM and 10% IAM) and Stepwise Mutation Model (SMM) (P-value < 0.05); ii – Mode-shift test that indicates bottlenecks resulting from alterations in allele frequency distributions (Luikart et al., 1998).
On mtDNA data, Arlequin v. 3.5.1.3 (Excoffier, Lischer, 2010) was used to estimate the genetic structure level using pairwise ΦST, analogous to Wright’s F-statistics, and to run the AMOVA (Analysis of Molecular Variance) among the samples. The analyses were conducted using the Tamura, Nei (1993) model, the best-fit nucleotide sequence evolution model according to ModelTest v. 3.7 (Posada, Crandall, 1998). Population differentiation analyses from microsatellite data were conducted in Structure v. 2.3.3 (Pritchard et al., 2000) program, which was used to perform a Bayesian cluster analysis, running twenty replicates for each K value (number of clusters), from K = 1–5, including a burn-in of 10,000 Markov Chain Monte Carlo (MCMC) iterations, followed by 100,000 MCMC of data collection. The optimal K was estimated using Structure Harvester (Earl, Von- Holdt, 2012), while Clumpp 1.1.2 (Jakobsson, Rosenberg, 2007) summarized the best K runs and Distruct 1.1 (Rosenberg, 2004) plotted the results on a graph. Additionally, pairwise DEST (Jost, 2008) value was calculated in the R statistical environment (R Development Core Team, 2021) with the DEMEtics package (Jueterbock et al., 2012), while FST and RST values were obtained in the Arlequin v. 3.5.1.3 (Excoffier, Lischer, 2010) program, using significant estimates based on 10,000 permutations.
Results
Genetic diversity and demographic history. A 492-bp (base pairs) fragment from mtDNA (D-loop region) was sequenced and analyzed for all C. hypselurus samples, revealing six polymorphic sites and six different haplotypes, H1–H6 (Genbank accession numbers: OP583660– OP583665). Considering just samples with N ≥ 20, the haplotype diversity (h) ranged from 0.246 (CIN) to 0.358 (LAM), while nucleotide diversity (π) ranged from 0.0005 (CIN) to 0.0007 (LAM) (Tab. 1).
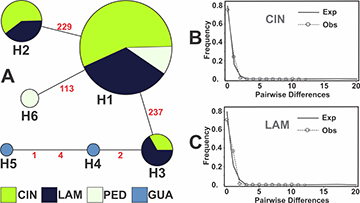
Three haplotypes occurred in CIN and LAM (H1, H2 and H3) and two in GUA and PED (H4 and H5). H1 was the most frequent haplotype, occurring in CIN, LAM and GUA, while three haplotypes (H4, H5, and H6) were singletons (Fig. 2A). No significant values were obtained from Tajima (D) and Fu (Fs) neutrality tests (Tab. 1) and the mismatch distribution graphic showed a unimodal distribution for haplotypes (Figs. 2B–C).
FIGURE 2| Results from mtDNA (D-Loop) of Cnesterodon hypselurus samples obtained in Cinzas River basin (CIN) and some other locations along the occurrence area reported for this species. A. Haplotype network. Circle sizes are proportional to haplotype frequency. Mismatch distributions of mitochondrial haplotypes for CIN and LAM are shown in B and C, respectively.
In the microsatellite analysis, Micro-Checker program showed no presence of scoring error or allele dropout and indicated possible null alleles and excess homozygotes only for two loci (Pvm11, Pvm14) in LAM. After sequential Bonferroni correction (α = 0.05, k = 8) for multiple comparisons, three loci in CIN (Pvm11, Pvm14 and Pvm18) and two (Pvm11 and Pvm14) in LAM showed significant deviations from Hardy–Weinberg proportions. Significant (α = 0.05, k = 28) linkage disequilibrium (LD) values were no detected in any loci combination. A total of 56 different alleles were obtained among the C. hypselurus samples, ranging from 22 (CIN) to 52 (LAM) per sample. In general, all CIN estimates, including Ne (16.3), NA (2.750), NE (1.771), HO (0.349), HE (0.334) and RA (2.606), were smaller than those obtained for LAM (Tab. 1). In the recent bottleneck analysis (microsatellite data), significant excess heterozygosity values (bottleneck sign) were not found in Wilcoxon’s signed-rank test and all samples showed a typical L-shaped distribution (non-bottleneck) in the allele frequency from the mode-shift test (Tab. S2).
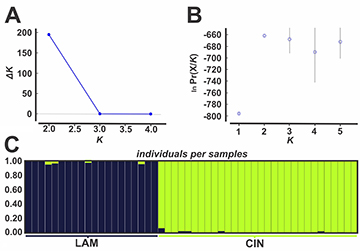
Genetic variation between populations. Considering only CIN and LAM(N ≥ 20), AMOVA results from microsatellite data indicated a large percentage of molecular variation (25.13%) occurred between these populations while mtDNA data showed no variation between samples. The pairwise ΦST, calculated from mtDNA, was not significant (ΦST = -0.027, P-value > 0.05). On the other hand, all population differentiation estimators used on microsatellite data (DEST =0.248, FST = 0.255 and RST = 0.360, P-value < 0.05) showed significant values. Bayesian clustering analysis indicated, from ln Pr(X/K) (Pritchard et al., 2000) and ∆K ad hoc statistics (Evanno et al., 2005), that the most probable K value (cluster number) was K = 2 (Figs. 3A–B). From this result, the graphic representation shows two distinct groups, one formed by CIN and the other by LAM (Fig. 3C).
FIGURE 3| Results of Bayesian analysis (STRUCTURE) for Cnesterodon hypselurus in Cinzas River basin (CIN) and Lambari stream (LAM), Itararé River basin. Estimates of the number of K groups based on mean A. Likelihood Ln(K) and B. ∆K statistic. C. Graphical representation based on K = 2. Each column represents a different individual and the colors represent the probability membership coefficient of that individual for each genetic cluster.
Discussion
In general, genetic diversity estimates for the microsatellite loci in LAM (HE = 0.528 and NA = 6.5), and mainly in CIN (HE = 0.334 and NA = 2.75), were below the average reported for freshwater fish (HE = 0.54 and NA = 9.1 alleles per locus) (Dewoody, Avise, 2000), as well as below the values previously obtained for other small-sized fish in drainages throughout the Neotropical region (Zaganini et al., 2014; Apolinário-Silva et al., 2018; Ferreira et al., 2021). However, other Neotropical Poeciliidae, such as Poecilia vivipara (HE = 0.507 and NA = 5.3) (Tonhatti et al., 2014) and Poecilia reticulata (HE = 0.660-0.728 and NA = 5.1-11.3) (Becher et al., 2002; Paterson et al., 2005; Suk, Neff, 2009), have already shown estimates close to the values of LAM, while Poeciliidae species in North America (HE = 0.267-0.462, NA = 2- 4.4) (Echelle et al., 2013) and Australia (HE = 0.457, NA = 3.2) (Umbers et al., 2012) had lower estimates, suggesting that lower variation levels could be common among Poeciliidae populations. Indeed, similar to the present study, other analyses of the D-Loop region in Poeciliidae species also report populations with low variation levels in Neotropical region, such as Cnesterodon decemmaculatus (h = 0.153-0.888 and π = 0.0005-0.0037) (Bruno et al., 2016), and in other regions of the planet, such as Gambusia krumholzi, G. clarkhubbsi, G. speciosa (h = 0.000-0.410 and π = 0.0000-0.0024) (Echelle et al., 2013) and Xiphophorus cortezi (h = 0.000-0.530 and π = 0.0000-0.0006) (Gutiérrez-Rodríguez et al., 2007) in Mexico.
Anyway, the genetic diversity levels and Ne of C. hypselurus in the new occurrence site (CIN) were very low compared to those reported in LAM and highlight the need for special attention in its management and conservation, mainly because other populations of this species, as well as their genetic diversity levels, are not yet known from any other location along the Cinzas River basin. Indeed, small-sized fish such as C. hypselurus tend to have a low displacement amplitude along their lifetime (Castro, 1999), often forming small and restricted populations, which are subject to genetic diversity loss due to evolutionary factors such as genetic drift, natural selection and inbreeding (Freeland, 2005; Hartl, Clark, 2007). Since the genetic diversity levels are determinant for the evolutionary/adaptive potential of the species, populations showing low estimates raise major concerns, mainly about how they will respond to different environmental changes (Frankham et al., 2010; Allendorf et al., 2012), including those derived from anthropogenic disturbances (Landweber, Dobson, 2021; Willi et al., 2022). Similar to the occurrence site in Arroio do Gica stream (altitude: 728 m) (Silva et al., 2015), CIN was sampled at an altitude (746 m) below the range (from 861 to 1,136 m) common among previous records of C. hypselurus (Lucinda, 2005; Silva et al., 2015). As already mentioned above, some of the most evident anthropogenic disturbances in this sampling site include the cultivation of exotic trees near the riparian forest and the presence of a bridge and rural road. However, signs of recent population bottlenecks were not detected in CIN and the most of the data seems to suggest that the genetic diversity levels would have greater influences from events that occurred in an older scenario, such as the founder effect during the colonization of a new drainage.
A founder effect occurs when few individuals from an original population establish a new population in an area where the species did not occur. Since the few founders carry only a small portion of all genetic diversity present in the original population, the new population will have its genetic diversity restricted to the alleles brought by the founders (at least as long as mutations do not generate new alleles) (Freeland, 2005; Hartl, Clark, 2007). In the present study, CIN had considerably low h, π, HO and HE estimates and showed RA,NA and NE values, as well as the number of alleles (22), which were less than or close to half of those obtained for LAM. Besides, both CIN and LAM showed h < 0.5 combined with π < 0.5%, which is commonly associated with founder effect involving one or a few mitochondrial DNA lineages (Grant, Bowen, 1998). Still in mtDNA data, some sign of population expansion (e.g., after founder effect) also seems to be suggested by the unimodal mismatch distribution and the haplotype network tending towards a star-shaped (Slatkin, Hudson, 1991; Rogers, Harpending, 1992). However, results of these last two analyses require caution, since factors such as low polymorphism levels (as observed in 492 bp from the D-loop region in the present study), small sample sizes (as for GUA and PED) and the number of analyzed populations can influence the outcome of a study (Grant, 2015). At the same time, the contrasting population differentiation patterns obtained between nuclear (microsatellite) and mitochondrial (D-Loop) DNA and the haplotypes and microsatellite alleles composition also seem to be evidence in favor of an older scenario (founder effect) influencing the low genetic diversity levels in CIN.
In general, the significant population differentiation (DEST, FST, RST and Bayesian clustering) obtain between CIN and LAM from microsatellite data was already expected, since the sites sampled are in the headwaters of different watersheds, without current connections in the study area. However, these samples shared the same haplotypes (H1, H2, and H3) and showed no significant genetic structuring from mtDNA. Indeed, while microsatellite loci are multiallelic, codominant, and highly polymorphic markers exhibiting biparental inheritance (Freeland, 2005), the mtDNA shows haploid uniparental inheritance, absence of genetic recombination (Avise, 2004) and mutation rates lower than those in microsatellite loci (Estoup, Angers, 1998; Santos et al., 2007). Thus, considering that microsatellite loci recover genetic diversity faster than mtDNA after a population bottleneck or founder effect (Freeland, 2005; Allendorf et al., 2012) and that the variation obtained herein for mtDNA is very low, it seems plausible that microsatellite loci already have an allelic composition that allows the detection of population differentiation while mtDNA still does not. In this context, microsatellite alleles coming with founders of CIN, even in smaller numbers than those in the original population, would have already had enough time for the establishment of new variants and particular frequencies, determining populational differences such as that observed in relation to LAM. Meanwhile, the same scenario would not have been possible for mtDNA, mainly due to its inherent characteristics and the very low variation level among the founders, since haplotypes different from H1 are still very few and have a very low frequency.
Since the present study did not cover all occurrence sites previously recorded for C. hypselurus, it is difficult to suggest the origin of the CIN’s founders. In any case, the sharing of the same haplotypes (H1, H2, and H3) between CIN and LAM, as well as the presence of almost all microsatellite alleles (18, except four private alleles) obtained for CIN among the 52 identified for LAM, also seems to support the founder effect in CIN. Indeed, regardless of the origin of the founders, the same haplotypes and part of the same microsatellite alleles were taken to different head-water streams without current connections, suggesting dispersion events that possibly involved headwater captures. As already mentioned, the previously reported occurrence area for C. hypselurus encompassed sites (altitude: from 728 to 1,136 m) in head-water streams of Tibagi and Itararé river basins (Silva et al., 2015). These watersheds, as well as the Cinzas River basin, are tributaries of the Paranapanema River (Maack, 2002; Britto et al., 2003), that is, the main dispersion route of fish species among these watersheds would be from their confluences with the Paranapanema River. However, this dispersal route seems less plausible for C. hypselurus, since this species has only been reported in headwater streams from higher altitudes in the upper sections of the watersheds (Silva et al., 2015). In fact, studies of ichthyofauna along drainages of the Paranapanema River basin (Jarduli et al., 2020), including several collections along the Cinzas River main channel (unpublished data) and its tributaries (Costa et al., 2013; Galindo et al., 2020, 2021; Caetano et al., 2021), did not find records of this species in the lower sections of the watersheds. In addition, in the case of Cinzas River basin, a waterfall of ~15 m high in the medium-high section of the main channel (Vianna, Nogueira, 2008) would also be another very limiting factor for dispersion, especially for a fish showing small body like C. hypselurus (Castro, 1999; Castro, Polaz, 2020). Thus, it seems plausible that C. hypselurus dispersed to CIN from a headwater capture and that this event has been reflected in founder effect signals and low genetic diversity levels. Since this event occurred a long time ago, the population would have already had a very large number of generations, establishing new allele and genotypic frequencies (closer to mutation-drift equilibrium), which show no signs of recent genetic bottlenecks, indicate few loci with deviations from the HWE, and do not suggest significant FIS for the population.
Interestingly, species inventories have also suggested headwater captures enabling exchanges of ichthyofauna between headwaters of Cinzas and Itararé basins (Frota et al., 2020) as well as among headwaters of other rivers throughout the state of Paraná, including the Iguaçu, Piquiri, Ivaí, Ribeira de Iguape and other tributaries of the Paranapanema River basin (Ribeiro, 2006; Frota et al.,2016; Cavalli et al.,2018; Morais-Silva et al.,2018). In all these cases, headwater captures have been associated with intense geological activities and activation of old faults during the Cenozoic, mainly in areas under influences of the Serra da Esperança and the Ponta Grossa Arch (Ribeiro, 2006; Frota et al.,2016). Indeed, although the upper Paraná River basin seems to maintain its main physiography since about 9.0 million years ago (Mya) (Albert, Reis, 2011), several events from the Miocene to the Pleistocene, including marine incursions, climate oscillations (Hubert, Renno, 2006; Carvalho et al., 2016), tectonism (Morais-Silva et al.,2018) and inter- and intra-basin headwater captures (Lundberg et al., 1998; Ribeiro, 2006; Cavalli et al., 2018), had important influences on the diversification of their species and fish populations.
In the case of CIN, the sampling site is in the area influenced by Ponta Grossa Arch, a large deformational structure formed since the break-up of southwestern Gondwana (around 110 Mya) (Lundberg et al.,1998; Franco-Magalhaes et al., 2010). The history of this Arch includes periods of tectonism, such as 1.7 Mya (Morais-Silva et al.,2018), which possibly influenced high altitude regions, including the Serra de Furnas and neighboring mountains where headwater streams of the Cinzas, Tibagi and Itararé river basins are located (Maack, 2002; Morais-Silva et al., 2018). At the same time, it is also important to consider that the headwater floodings during Pleistocene climatic oscillations (2.58 to 0.117 Mya) could also have been a pathway enabling exchanges of ichthyofauna between the different watersheds (Ribeiro, 2006). In this context, even observations of the current relief in the study area also suggest stretches where past headwater connections could have occurred (lowlands between mountains). Genetic data from species in Paraná River basin have already suggested influences of Pleistocene climatic oscillations on the distribution of fish populations (Artoni et al., 2006; Carvalho et al., 2016; Mondin et al., 2018). In the case of the suckermouth catfish Hypostomus ancistroides, for example, alternating cycles of humidity and dryness along the Quaternary seem to have resulted in isolation and subsequent merging of different populations, as well as in the dispersal of this species to the Ribeira de Iguape River basin from a Pleistocene headwater capture event (Carvalho et al., 2016).
In conclusion, present study identified low genetic diversity levels for a rare fish species in a previously unreported occurrence area (Cinzas River basin), suggesting influences from demographic history, possibly involving founder effect after dispersal made possible by a headwater capture, as well as from some biological aspects of C. hypselurus, including small body size, low displacement amplitude and occurrence commonly restricted to clear waters of headwaters (Castro, 1999; Silva et al., 2015). Since the population identified in Cinzas River is genetically different from that analyzed in the Itararé River basin and that other occurrence sites are unknown in the Cinzas River basin, the data herein are quite relevant in future actions to preserve C. hypselurus and their occurrence areas. Considering the low estimate of effective population size, measures including long-term monitoring of genetic diversity and inbreeding could provide a basis for future decision-making. Furthermore, the search for additional undiscovered populations in Cinzas, Tibagi and Itararé River basins, as already highlighted by Silva et al. (2015), and a broad population genetic analysis would be important future steps to improve management and conservation strategies and the understanding of phylogeographical aspects.
Acknowledgments
We are grateful to the Fundação Araucária for financial support (number 04/2017), the Universidade Estadual do Norte do Paraná (UENP) for financial and logistical support, Dra. Lenice Souza-Shibatta for donating some samples of C. hypselurus; Dr. Oscar Akio Shibatta (Universidade Estadual de Londrina) for his help in identifying the species studied and the Instituto Brasileiro de Meio Ambiente e dos Recursos Naturais Renováveis (IBAMA) and Instituto Chico Mendes de Conservação da Biodiversidade (ICMBio) for granting permission to collect samples.
References
Agostinho AA, Gomes LC, Santos NCL, Ortega JCG, Pelicice FM. Fish assemblages in Neotropical reservoirs: Colonization patterns, impacts and management. Fish Res. 2016; 173:26–36. https://doi.org/10.1016/j.fishres.2015.04.006
Ahmed SF, Kumar PS, Kabir M, Zuhara FT, Mehjabin A, Tasannum N et al. Threats, challenges and sustainable conservation strategies for freshwater biodiversity. Environ Res. 214:113808. https://doi.org/10.1016/j.envres.2022.113808
Albert JS, Reis RE. Historical biogeography of Neotropical freshwater fishes. Berkeley: University of California Press; 2011.
Allendorf FW, Luikart GH, Aitken SN. Conservation and the genetics of populations. Oxford: Wiley Blackwell Publishing; 2012.
Apolinário-Silva C, Ferreira DG, Cavenagh AF, Aprígio NGO, Galindo BA, Carlsson J et al. Development and characterization of fifteen polymorphic microsatellite loci in Bryconamericus aff. iheringii (Teleostei: Characidae) and cross-amplification in related Characidae species. Neotrop Ichthyol. 2018; 16(1):e170135. https://doi.org/10.1590/1982-0224-20170135
Apolinário-Silva C, Galindo BA, Nascimento RHC, Frantine-Silva W, Kotelok-Diniz T, Sofia SH et al. Fine-scale genetic structure of suckermouth Hypostomus ancistroides populations: the importance of Neotropical streams for fish conservation. Biol J Linn Soc. 2021; 134(1):198–213. https://doi.org/10.1093/biolinnean/blab039
Artoni RF, Shibatta OA, Gross MC, Schneider CH, Almeida MCD, Vicari MR et al. Astyanax aff. fasciatus Cuvier, 1819 (Teleostei; Characidae): evidences of a species complex in the upper rio Tibagi basin (Paraná, Brazil). Neotrop Ichthyol. 2006; 4(2):197–202. https://doi.org/10.1590/S1679-62252006000200005
Arya S. Freshwater biodiversity and conservation challenges: A review. Int J Biol Inno. 2021;3(1):74–78. https://doi.org/10.46505/IJBI.2021.3106
Avise JC. Molecular markers, natural history, and evolution. 2nd ed. Sunderland: Sinauer Associates; 2004.
Bandelt HJ, Forster P, Röhl A. Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol. 1999; 16(1):37–48. https://doi.org/10.1093/oxfordjournals.molbev.a026036
Becher SA, Russell ST, Magurran AE. Isolation and characterization of polymorphic microsatellites in the Trinidadian guppy (Poecilia reticulata). Mol Ecol Notes. 2002; 2(4):456–58. https://doi.org/10.1046/j.1471-8286.2002.00276.x
Britto SGC, Sirol RN, Vianna NC, Jardim SM, Santos JC, Pelisari E. Peixes do rio Paranapanema. São Paulo: Duke Energy; 2003.
Bruno MC, Mapelli FJ, Casciotta JR, Almirón AE, Lizarralde MS. Phylogeography of Cnesterodon decemmaculatus (Cyprinodontiformes: poeciilidae) in Southern Pampas, Argentina: ancient versus recent patterns in freshwater fishes. Environ Biol Fishes. 2016; 99:293–307. https://doi.org/10.1007/s10641-016-0474-0
Caetano DLF, Oliveira EF, Zawadzki CH. Ichthyofauna of tributary streams of the Cinzas River basin, Paranapanema River, Brazil. Oecologia Aust. 2021; 25(1):142–53. https://doi.org/10.4257/oeco.2021.2501.13
Castro RMC, Polaz CNM. Small-sized fish: the largest and most threatened portion of the megadiverse neotropical freshwater fish fauna. Biota Neotrop. 2020; 20(1):e20180683. https://doi.org/10.1590/1676-0611-bn-2018-0683
Castro RMC. Evolução da ictiofauna de riachos sul-americanos: padrões gerais e possíveis causais. In: Caramaschi EP, Mazzoni R, Peres-Neto PR, editors. Ecologia de Peixes de Riachos. Rio de Janeiro: PPGE-UFRJ; 1999. p.139–55. https://doi.org/10.4257/oeco.2021.2502.02
Cavalli D, Frota A, Lira AD, Gubiani EA, Margarido VP, Graça WJD. Update on the ichthyofauna of the Piquiri River basin, Paraná, Brazil: a conservation priority area. Biota Neotrop. 2018; 18(2):e20170350. https://doi.org/10.1590/1676-0611-BN-2017-0350
Carvalho PH, Lima SMQ, Zawadzki CH, Oliveira C, Pinna M. Phylogeographic patterns in suckermouth catfish Hypostomus ancistroides (Loricariidae): dispersion, vicariance and species complexity across a Neotropical biogeographic region. Mitochondrial DNA Part A. 2016; 27(5):3590–96. https://doi.org/10.3109/19401736.2015.1079822
Costa ADA, Ferreira DG, Silva WF, Zanatta AS, Shibatta AO, Galindo BA. Fishes (Osteichthyes: Actinopterygii) from the Penacho stream, upper Paraná River basin, Paraná State, Brazil. Check List. 2013; 9(3):519–23. http://dx.doi.org/10.15560/9.3.519
Cronin MA, Spearman WJ, Wilmot RL, Patton JC, Bickham JW. Mitochondrial DNA variation in chinook (Oncorhynchus tshawytscha) and chum salmon (O. keta) detected by restriction enzyme analysis of polymerase chain reaction (PCR) products. Can J Fish Aquat Sci. 1993; 50(4):708–15. https://doi.org/10.1139/f93-081
DeWoody JA, Avise JC. Microsatellite variation in marine, freshwater and anadromous fishes compared with other animals. J Fish Biol. 2000; 56(3):461–73. https://doi.org/10.1111/j.1095-8649.2000.tb00748.x
Do C, Waples RS, Peel D, Macbeth GM, Tillett BJ, Ovenden JR. NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour. 2014; 14(1):209–14. https://doi.org/10.1111/1755-0998.12157
Earl DA, Von Holdt BM. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour. 2012; 4(2):359–61. http://dx.doi.org/10.1007/s12686-011-9548-7
Echelle AA, Vilano MLL, Baker S, Wilson WD, Echelle AF, Garrett GP et al. Conservation genetics of Gambusia krumholzi (Teleostei: Poeciliidae) with assessment of the species status of G. clarkhubbsi and hybridization with G. speciosa. Copeia. 2013; 2013(1):72–79. https://doi.org/10.1643/CG-11-167
Estoup A, Angers B. Microsatellites and mini satellites for molecular ecology: theoretical and experimental considerations. In: Carvallo GR, editor. Advances in molecular ecology. Amsterdam: NATO Press; 1998. p.55–86.
Evanno G, Regnaut S, Goudet DJ. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol Ecol. 2005; 14(8):2611–20. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Excoffier L, Lischer HEL. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour. 2010; 10(3):564–67. https://doi.org/10.1111/j.17550998.2010.02847.x
Ferreira DG, Galindo BA, Apolinário-Silva C, Nascimento RHC, Frantine-Silva W, Cavenagh AF et al. Influences of small hydroelectric plants on the genetic differentiation of Neotropical freshwater fish populations: a case study. Stud Neotrop Fauna Environ. 2021; 1–13. https://doi.org/10.1080/01650521.2021.1994349
Ferreira DG, Lima SC, Frantine-Silva W, Silva JF, Apolinário-Silva C, Sofia SH et al. Fine-scale genetic structure patterns in two freshwater fish species, Geophagus brasiliensis (Osteichthyes, Cichlidae) and Astyanax altiparanae (Osteichthyes, Characidae) throughout a Neotropical stream. Genet Mol Res. 2016; 15(4):gmr15048124. http://dx.doi.org/10.4238/gmr15048124
Franco-Magalhaes AO, Hackspacher PC, Glasmacher UA, Saad AR. Rift to post-rift evolution of a “passive” continental margin: the Ponta Grossa Arch, SE Brazil. Int J Earth Sci. 2010; 99(7):1599–613. https://doi.org/10.1007/s00531-010-0556-8
Frankham R, Ballou JD, Briscoe DA. Introduction to conservation genetics. 2nd ed. Cambridge: Cambridge University Press; 2010.
Freeland JR. Molecular ecology. Chichester: John Wiley & Sons Ltd; 2005.
Fricke R, Eschmeyer WN, Van der Laan R. Eschmeyer’s catalog of fishes: genera, species, references. San Francisco: California Academy of Science; 2022. Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp
Frota A, Deprá GC, Petenucci LM, Graça WJ. Inventory of the fish fauna from Ivaí River basin, Paraná State, Brazil. Biota Neotrop. 2016; 16(3):e20150151. http://dx.doi.org/10.1590/1676-0611-BN-2015-0151
Frota A, Ota RR, Deprá GC, Ganassin MJM, Graça WJ. A new inventory for fishes of headwater streams from the rio das Cinzas and rio Itararé basins, rio Paranapanema system, Paraná, Brazil. Biota Neotrop. 2020; 20(1):e20190833. https://doi.org/10.1590/1676-0611-BN-2019-0833
Fu Y-X. Statistical test of neutrality of mutations against population growth, hitchhiking and background selection. Genetics. 1997; 147(2):915–25. https://doi.org/10.1093/genetics/147.2.915
Galindo BA, Costa ADA, Ferreira DG, Garcia TD, Frota A, Frantine-Silva W et al. Ichthyofauna of the Água das Araras stream, Paranapanema River basin, upper Paraná River system, Southern Brazil. Stud Neotrop Fauna Environ. 2021;1–15. https://doi.org/10.1080/01650521.2021.1967690
Galindo BA, Ota RR, Garcia TD, Nascimento RHC, Ohara WM, Zanatta AS et al. Inventory of the fish fauna from Laranjinha River, Paranapanema River system, Brazil. Biota Neotrop. 2020; 20(4):e20200962. https://doi.org/10.1590/1676-0611-BN-2020-0962
Geist J. Integrative freshwater ecology and biodiversity conservation. Ecol Indic. 2011; 11(6):1507–16. https://doi.org/10.1016/j.ecolind.2011.04.002
Goudet J. FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). 2001 [cited 2021 Sep 9]. Available from: http://www.unil.ch/izea/softwares/fstat.html
Grant WAS, Bowen BW. Shallow population histories in deep evolutionary lineages of marine fishes: insights from sardines and anchovies and lessons for conservation. J Hered. 1998; 89(5):415–26. https://doi.org/10.1093/jhered/89.5.415
Grant WS. Problems and cautions with sequence mismatch analysis and Bayesian skyline plots to infer historical demography. J Hered. 2015; 106(4):333–46. https://doi.org/10.1093/jhered/esv020
Gutiérrez-Rodríguez C, Morris MR, Dubois NS, Queiroz K. Genetic variation and phylogeography of the swordtail fish Xiphophorus cortezi (Cyprinodontiformes, Poeciliidae). Mol Phylogenet Evol. 2007; 43(1):111–23. https://doi.org/10.1016/j.ympev.2006.10.022
Hartl DL, Clark AG. Principles of population genetics. Sunderland: Sinauer Associates; 2007. https://doi.org/10.1093/jhered/esm035
Hubert N, Renno JF. Historical biogeography of South American freshwater fishes. J Biogeogr. 2006; 33(8):1414–36. https://doi.org/10.1111/j.1365-2699.2006.01518.x
Instituto Chico Mendes de Conservação da Biodiversidade (ICMBio). Livro Vermelho da Fauna Brasileira Ameaçada de Extinção: Volume VI – Peixes. Brasília: ICMBio/MMA; 2018.
Instituto Chico Mendes de Conservação da Biodiversidade (ICMBio). Sistema de Avaliação do Risco de Extinção da Biodiversidade – SALVE. 2023. Available from: https://salve.icmbio.gov.br/
Jakobsson M, Rosenberg NA. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics. 2007; 23(14):1801–06. https://doi.org/10.1093/bioinformatics/btm233
Jarduli LR, Garcia DAZ, Vidotto-Magnoni AP, Casimiro ACR, Vianna NC, Almeida FS et al. Fish fauna from the Paranapanema River basin, Brazil. Biota Neotrop. 2020; 20(1):e20180707. https://doi.org/10.1590/1676-0611-BN-2018-0707
Jost LOU. GST and its relatives do not measure differentiation. Mol Ecol. 2008; 17(18):4015–26. https://doi.org/10.1111/j.1365-294X.2008.03887.x
Jueterbock A, Kraemer P, Gerlach G, Deppermann, Jueterbock JMA. Package ‘DEMEtics.’ Mol Ecol. 2012; 19:3845–52.
Landweber L, Dobson A. Genetics and the extinction of species: DNA and the conservation of biodiversity. Princeton: Princeton University Press; 2021.
Librado P, Rozas J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics. 2009; 25(11):1451–52.
Lucinda PHF. Family Poeciliidae. In: Reis RE, Kullander SO, Ferraris CJ, Jr., editors. Check list of the freshwater fishes of South and Central America. Porto Alegre: Edipucrs; 2003. p.558–84.
Lucinda PHF. Systematics of the genus Cnesterodon Garman, 1895 (Cyprinodontiformes: Poeciliidae: Poeciliinae). Neotrop Ichthyol. 2005; 3(2):259–70. https://doi.org/10.1590/S1679-62252005000200003
Lucinda PHF, Garavello JC. Two new species of Cnesterodon Garman, 1895 (Cyprinodontiformes: Poeciliidae) from the upper rio Paraná drainage. Comun Mus Ciênc Tecnol PUCRS, Sér Zool. 2001; 13(2):119–38.
Lucinda PHF, Reis RE. Systematics of the subfamily Poeciliinae Bonaparte (Cyprinodontiformes: Poeciliidae), with an emphasis on the tribe Cnesterodontini Hubbs. Neotrop Ichthyol. 2005; 3(1):1–60. https://doi.org/10.1590/S1679-62252005000100001
Luikart G, Allendorf FW, Cornuet JM, Sherwin WB. Distortion of allele frequency distributions provides a test for recent population bottlenecks. J Hered. 1998; 89(3):238–47. https://doi.org/10.1093/jhered/89.3.238
Luikart G, Cornuet JM. Empirical evaluation of a test for identifying recently bottlenecked populations from allele frequency data. Conserv Biol. 1998; 12(1):228–37. https://doi.org/10.1111/j.15231739.1998.96388.x
Lundberg JG, Marshall LG, Guerrero J, Horton B, Malabarba MCSL, Wesselingh FP. The stage for Neotropical fish diversification: a history of tropical South American rivers. In: Malabarba LR, Reis RE, Vari RP, Lucena ZMS, Lucena CAS, editors. Phylogeny and classification of Neotropical fishes. Porto Alegre: Edipucrs; 1998. p.13–48.
Maack V. Geografia física do Estado do Paraná. Curitiba: Imprensa Oficial do Paraná; 2002.
Meyer A, Kocher TD, Basasibwaki P, Wilson AC. Monophyletic origin of Lake Victoria cichlid fishes suggested by mitochondrial DNA sequences. Nature. 1990; 347:550–53. https://dx.doi.org/10.1038/347550a0
Mondin LAC, Machado CB, Resende EK, Marques DKS, Galetti Jr PM. Genetic pattern and demographic history of Salminus brasiliensis: Population expansion in the Pantanal Region during the Pleistocene. Front Genet. 2018; 9:1–08. https://doi.org/10.3389/fgene.2018.00001
Morais-Silva JP, Oliveira AV, Fabrin TMC, Diamante NA, Prioli SMAP, Frota A et al. Geomorphology influencing the diversification of fish in small-order rivers of neighboring basins. Zebrafish. 2018; 15(4):389–97. https://doi.org/10.1089/zeb.2017.1551
Paterson IG, Crispo E, Kinnison MT, Hendry AP, Bentzen P. Characterization of tetranucleotide microsatellite markers in guppy (Poecilia reticulata). Mol Ecol Notes. 2005; 5(2):269–71. https://doi.org/10.1111/j.1471-8286.2005.00895.x
Piry S, Luikart G, Cornuet JM. BOTTLENECK: A computer program for detecting recent reductions in the effective population size using allele frequency data. J Hered. 1999; 90(4):502–03. https://doi.org/10.1093/jhered/90.4.502
Posada D, Crandall KA. ModelTest: testing the model of DNA substitution. Bioinformatics. 1998; 14(9):817–18. https://doi.org/10.1093/bioinformatics/14.9.817
Pritchard JK, Stephens M, Donnelly P. Inference of population structure using multi locus genotype data. Genetics. 2000; 155(2):945–59. https://doi.org/10.1093/genetics/155.2.945
Quintans F, Scasso F, Defeo O. Unsuitability of Cnesterodon decemmaculatus (Jenyns, 1842) for mosquito control in Uruguay: evidence from food-preference experiments. J Vector Ecol. 2010; 35(2):333–38. https://doi.org/10.1111/j.1948-7134.2010.00091.x
Quintans F, Scasso F, Loureiro M, Yafe A. Diet of Cnesterodon decemmaculatus (Poeciliidae) and Jenynsia multidentata (Anablepidae) in a hypertrophic shallow lake of Uruguay. Iheringia, Sér Zool. 2009; 99(1):99–105. https://doi.org/10.1590/S0073-47212009000100014
Raymond M, Rousset F. GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered. 1995; 86(3):248–49.
Reis RE, Albert JS, Di Dario F, Mincarone MM, Petry P, Rocha LA. Fish biodiversity and conservation in South America. J Fish Biol. 2016; 89(1):12–47. https://doi.org/10.1111/jfb.13016
Ribeiro AC. Tectonic history and the biogeography of the freshwater fishes from the coastal drainages of eastern Brazil: an example of faunal evolution associated with a divergent continental margin. Neotrop Ichthyol. 2006; 4(2):225–46. https://doi.org/10.1590/S167962252006000200009
Rice WR. Analyzing tables of statistical tests. Evolution. 1989; 43(1):223–25. http://dx.doi.org/10.2307/2409177
Rogers AR, Harpending H. Population growth makes waves in the distribution of pairwise genetic differences. Mol Biol Evol. 1992; 9(3):552–69. https://doi.org/10.1093/oxfordjournals.molbev.a040727
Rosenberg NA. DISTRUCT: A program for the graphical display of population structure. Mol Ecol Notes. 2004; 4(1):137–38. https://doi.org/10.1046/j.1471-8286.2003.00566.x
Santos MCF, Ruffino ML, Farias IP. High levels of genetic variability and panmixia of the tambaqui Colossoma macropomum (Cuvier, 1816) in the main channel of the Amazon River. J Fish Biol. 2007; 71:33–44. https://doi.org/10.1111/j.1095-8649.2007.01514.x
Schuelke M. An economic method for the fluorescent labelling of PCR fragments. Nat Biotechnol. 2000; 18(2):233–34. http://dx.doi.org/10.1038/72708
Silva J, Jerep F, Bennemann S. New record and distribution extension of the endangered freshwater fish Cnesterodon hypselurus (Cyprinodontiformes: Poeciliidae) in the upper Paraná River basin, Brazil. Check List. 2015; 11(6):1811. http://dx.doi.org/10.15560/11.6.1811
Shibatta OA, Orsi ML, Bennemann ST, Silva-Souza AT. Diversidade e distribuição de peixes na bacia do rio Tibagi. In: Medri ME, Bianchini E, Shibatta OA, Pimenta JA, editors. A bacia do rio Tibagi. Londrina: Medri ME; 2002. p.403–23.
Slatkin M, Hudson RR. Pairwise comparisons of mitochondrial DNA sequences in stable and exponentially growing populations. Genetics. 1991; 129(2):555–62. https://doi.org/10.1093/genetics/129.2.555
Suk HY, Neff BD. Microsatellite genetic differentiation among populations of the Trinidadian guppy. Heredity. 2009; 102:425–34. https://doi.org/10.1038/hdy.2009.7
Tagliacollo VA, Dagosta FCP, Pinna M, Reis RE, Albert JS. Assessing extinction risk from geographic distribution data in Neotropical freshwater fishes. Neotrop Ichthyol. 2021; 19(3):e210079. https://doi.org/10.1590/1982-0224-2021-0079
Tajima F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics. 1989; 123(3):585–95. https://doi.org/10.1093/genetics/123.3.585
Tamura K, Nei M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol. 1993; 10(3):512–26. https://doi.org/10.1093/oxfordjournals.molbev.a040023
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol. 2011; 28(10):2731–39. https://doi.org/10.1093/molbev/msr121
Tonhatti CH, Louvise J, Souza AP, Reis SF, Laborda PR. Development of microsatellite loci for the fish Poecilia vivipara (Cyprinodontiformes: Poeciliidae). Conserv Genet Resour. 2014; 6(2):383–85. https://doi.org/10.1007/s12686-013-0098-z
Umbers KD, Jennions MD, Gardner MG, Keogh JS. Twenty-five new polymorphic microsatellites for the eastern mosquitofish, Gambusia holbrooki (Actinopterygii: Poeciliidae), an invasive species in Australia. Aust J Zool. 2012; 60(4):235–37. http://dx.doi.org/10.1071/ZO12095
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P. MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes. 2004; 4(3):535–38. http://dx.doi.org/10.1111/j.1471-8286.2004.00684.x
Vianna NC, Nogueira MG. Ichthyoplankton and limnological factors in the Cinzas River – an alternative spawning site for fishes in the middle Paranapanema River basin, Brazil. Acta Limnol Bras. 2008; 20(2):139–51.
Waples RS, Do C. ldne: a program for estimating effective population size from data on linkage disequilibrium. Mol Ecol Resour. 2008; 8(4):753–56. https://doi.org/10.1111/j.1755-0998.2007.02061.x
Willi Y, Kristensen TN, Sgrò CM, Weeks AR, Ørsted M, Hoffmann AA. Conservation genetics as a management tool: The five best-supported paradigms to assist the management of threatened species. Proc Natl Acad Sci. 2022; 119(1):e2105076119. https://doi.org/10.1073/pnas.2105076119
Yeh FC, Yang RC, Boyle TBJ. POPGENE (version 1.32): Microsoft Windows-based freeware for population genetic analysis. Edmonton: University of Alberta; 1999.
Zaganini RL, Hashimoto DT, Pereira LHG, Oliveira C, Mendonça FF, Foresti F et al. Isolation and characterization of microsatellite loci in the Neotropical fish Astyanax altiparanae (Teleostei: Characiformes) and cross-species amplification. J Genet. 2014; 93(1):24–27. http://dx.doi.org/10.1007/s12041-012-0143-9
Authors
![]() Dhiego G. Ferreira1,
Dhiego G. Ferreira1, ![]() Bruno A. Galindo1,
Bruno A. Galindo1, ![]() Tais C. de Souza1,
Tais C. de Souza1, ![]() Leonardo B. Pereira1,
Leonardo B. Pereira1, ![]() Victor A. P. Bernardes1,
Victor A. P. Bernardes1, ![]() Ana J. C. Marques1
Ana J. C. Marques1 ![]() ,
, ![]() Wilson Frantine-Silva1,
Wilson Frantine-Silva1, ![]() Thais Kotelok-Diniz2,
Thais Kotelok-Diniz2, ![]() Carlos E. G. Aggio1,
Carlos E. G. Aggio1, ![]() Caroline Apolinário-Silva2,
Caroline Apolinário-Silva2, ![]() Augusto S. Zanatta1 and
Augusto S. Zanatta1 and ![]() Silvia H. Sofia2
Silvia H. Sofia2
[1] Grupo de Pesquisas em Ecologia, Recursos Naturais e Limnologia (GERCOL), Universidade Estadual do Norte do Paraná,Campus Cornélio Procópio, Rodovia PR-160, km 0, 86300-000 Cornélio Procópio, PR, Brazil. (DGF) dhiego@uenp.edu.br, (TCS) taicristina.souza.1@gmail.com, (LBP) leonardo.berto21@outlook.com, (VAPB) victorangelop.vp@gmail.com, (AJCM)anajuliacardoso2507@gmail.com (corresponding author), (WF) wilsonfrantine@gmail.com, (CEGA) aggiocarlos@uenp.edu.br, (BAG) bruno@uenp.edu.br, (ASZ) zanatta@uenp.edu.br.
[2] Laboratório de Genética e Ecologia Animal (LAGEA), Departamento de Biologia Geral, Universidade Estadual de Londrina,Rodovia Celso Garcia Cid, km 380, 86051-970 Londrina, PR, Brazil. (CA) carolapolinario07@gmail.com, (TK) thaiskotelok@gmail.com, (SHS) shsofiabelh@gmail.com.
Authors’ Contribution 

Dhiego G. Ferreira: Conceptualization, Funding acquisition, Investigation, Methodology, Project administration, Software, Writing-original draft, Writing-review and editing.
Bruno A. Galindo: Conceptualization, Funding acquisition, Methodology, Software, Writing-original draft.
Tais C. de Souza: Investigation, Methodology, Software, Writing-original draft.
Leonardo B. Pereira: Investigation, Methodology, Software, Writing-original draft.
Victor A. P. Bernardes: Data curation, Investigation, Methodology, Software, Writing-original draft.
Ana J. C. Marques: Investigation, Methodology, Software, Writing-original draft.
Wilson Frantine-Silva: Investigation, Methodology, Software, Supervision, Writing-original draft.
Thais Kotelok-Diniz: Data curation, Investigation, Software, Validation, Writing-original draft.
Carlos E. G. Aggio: Data curation, Investigation, Methodology, Supervision, Validation, Writing-original draft.
Caroline Apolinário-Silva: Data curation, Investigation, Methodology, Software, Supervision, Writing-original draft.
Augusto S. Zanatta: Data curation, Funding acquisition, Investigation, Software, Supervision, Writing-original draft.
Silvia H. Sofia: Funding acquisition, Investigation, Project administration, Writing-original draft.
Ethical Statement
Sampling was carried out under environmental authorization number 40811-5 SISBIO/MMA and approval from the Ethics Committee of the Universidade Estadual do Norte do Paraná, CEUA: nº 07/2020.
Competing Interests
The author declares no competing interests.
How to cite this article
Ferreira DG, Galindo BA, Souza TC, Pereira LB, Bernardes VAP, Marques AJC, Frantine-Silva W, Kotelok-Diniz T, Aggio CEG, Apolinário-Silva C, Zanatta AS, Sofia SH. Genetic diversity of the species Cnesterodon hypselurus (Cyprinodontiformes: Poeciliidae) in Cinzas River basin: new record and headwater capture evidences. Neotrop Ichthyol. 2023; 21(1):e230007. https://doi.org/10.1590/1982-0224-2023-0007
Copyright
This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Distributed under
Creative Commons CC-BY 4.0

© 2023 The Authors.
Diversity and Distributions Published by SBI
![]() Accepted February 1, 2023 by Claudio Oliveira
Accepted February 1, 2023 by Claudio Oliveira
![]() Submitted October 15, 2022
Submitted October 15, 2022
![]() Epub March 20, 2023
Epub March 20, 2023