![]() María Doris Escobar L.1
María Doris Escobar L.1 ![]() ,
, ![]() Izeni P. Farias1 and
Izeni P. Farias1 and ![]() Tomas Hrbek1,2
Tomas Hrbek1,2
PDF: EN XML: EN | Supplementary: S1 S2 S3 S4 S5 S6 S7 S8 S9 S10 S11 S12 S13 S14 S15 S16 | Cite this article
Abstract
Recently, the large migratory fishes of the family Serrasalmidae (Piaractus brachypomus and P. orinoquensis) were described as restricted to the Orinoco and Amazon basins. Both species provide important ecosystem services. They also are an important fisheries resource, which has caused that their populations have decreased in recent years. National fisheries policies still consider both species as one, which leads to inefficiencies in their management and conservation. The aim of this study was to genetically characterize these two species, using microsatellite and mitochondrial markers, and discuss the implication of these results for conservation and management. We found that both species have moderate genetic diversity and varied patterns of genetic distribution in the fluvial landscape. Piaractus brachypomus presented genetic diversity of A=6.5; He=0.72; Ho=0.67; Ĥ=1.000; ᴫ=0.0092, three management units related to the evolutionary process of the Amazon basin and the effective sizes of local populations were smaller compared to P. orinoquensis,which presented genetic diversity of A=6.1; He=0.66; Ho=0.55; Ĥ=0.968; ᴫ=0.010 and comprises only one management unit. These results demonstrate the need to design management policies that focus on species and geographically restricted populations.
Keywords: Cachama blanca, Fishery management, Morocoto, Pacu, Pirapitinga.
Recientemente, los grandes peces migratorios Serrasalmidae (Piaractus brachypomus y P. orinoquensis) fueron diferenciados en las cuencas del Orinoco y Amazonas. Ambas especies brindan importantes servicios ecosistémicos, entre ellos la pesca para la dieta humana, lo que ha provocado que sus poblaciones hayan disminuido en los últimos años. Las políticas pesqueras nacionales aún consideran a ambas especies como una sola, lo que conduce a ineficiencias en su gestión y conservación. El objetivo de este estudio fue caracterizar genéticamente estas dos especies de Piaractus, usando marcadores microsatélites y mitocondriales, y discutir la implicación de estos resultados para la conservación y el manejo. Encontramos que ambas especies tienen una diversidad genética moderada y patrones variados de distribución genética en el paisaje fluvial. Piaractus brachypomus presentó diversidad genética de A=6.5; He=0.72; Ho=0.67; Ĥ=1.000; ᴫ=0.0092, tres unidades de manejo relacionadas con el proceso evolutivo de la cuenca Amazónica y los tamaños efectivos de las poblaciones locales fueron menores en comparación con P. orinoquensis, que presentó diversidad genética de A=6.1; He=0.66; Ho=0.55; Ĥ=0.968; ᴫ=0,010 y comprende una sola unidad de gestión. Estos resultados demuestran la necesidad de diseñar políticas de manejo que se enfoquen en especies y poblaciones geográficamente restringidas.
Palabras clave: Cachama blanca, Manejo pesquero, Morocoto, Pacu, Pirapitinga.
Introduction
The species Piaractus brachypomus (Cuvier, 1818) and Piaractus orinoquensis Escobar L., Ota, Machado-Allison, Andrade-López, Farias & Hrbek, 2019 are the second largest fishes of the order Characiformes with maximum sizes of approximately 88 cm TL, after Colossoma macropomum (Cuvier, 1816) (max. of 108 cm TL) (Ortí et al., 2008; Froese, Pauly, 2022). Both species of Piaractus were considered a single species for more than two centuries. Escobar L et al. (2019) differentiated P. orinoquensis from P. brachypomus, the first from the Orinoco basin and the second from the Amazon basin, where they are widely distributed, occupying lacustrine and fluvial environments of the white and clear waters. Both species migrate with schools moving between 100 to 150km for reproductive, food and dispersal purposes at different seasons of the year (Goulding, 1979; Loubens, Panfili, 2001; Novoa, 2002; Barthem, Fabré, 2004; Barthem, Goulding, 2007; Usma et al., 2009; Van Damme et al., 2011). In each basin the species provide important ecosystem services. They are important to the local and regional economies where they are a fishing and aquaculture resource being caught and reared for human consumption; additionally, due to their trophic omnivore characteristics (with a tendency to frugivory), they play an important role in the ecosystem as seed dispersers (Novoa, 2002; Barthem, Fabré, 2004; Carvajal-Vallejos, Zeballos-Fernández, 2011; Lasso et al., 2011; Lasso, Sanchez-Duarte, 2011). Given their economic importance, these species have suffered from severe overfishing, and also have been negatively impacted by other anthropic activities that degrade their natural environments (Goulding, 1979; CCI, 2008). Although there are national fishery management policies focused on species protection in each basin where they occur (Acuerdos No 008/1997, No 006/1998 INPA y Resolución 1924/2005 AUNAP in Colombia; Resolución 003/2002 INAPESCA in Venezuela; Portaria No 48/2007 IBAMA in Brazil and Resolución Ministerial No 147/2001 Ministerio de Producción in Perú), these policies are ambiguous, ineffective and fail in their conservation aims. Generally, as policy actions are based on the differences in phenotypic characteristics, abundance and mean size indicators associated with the geographic region these species occupy, all policies assume that the species comprise one homogeneous population (Batista et al., 2012). This management strategy is advantageous for the fishing economy in the short-term, given its easy application; however, in the long-term, it can cause the extinction of natural populations because the structure of biological units is ignored (Begg, Waldman, 1999).
Disregarding biological units, combined with high exploitation, can result in the depletion or even extinction of fish stocks (Begg, Waldman, 1999). Thus, the traditional methods of evaluating fishery resources are facing criticism because the applied models have repeatedly failed and are not successful in preventing population reductions (Ruffino et al., 2012). Therefore, it is fundamental to define the Management Units (MUs) within species to help guide management and conservation efforts and to facilitate the application of laws. MUs are biological units or subpopulations that are demographically independent and equivalent to “stocks” in fisheries biology. The MUs are nested within an Evolutionary Significant Unit (ESU), which can have one or several MUs (Allendorf, Luikart, 2007). For the definition of the MUs, the most common criterion is the rejection of panmixia (Palsbøll et al., 2006).
According to Allendorf and Luikart (2007), among important factors for fishery management is the knowledge of genetic diversity, and identification of the structure of population or MU. For that reason, genetic tools can be applied in fishery management as a fast and efficient methods to define stocks (Carvalho, Hauser, 1995). Molecular markers, such as microsatellites (nuclear) and D-loop (mitochondrial), are tools that are often applied in these types of studies. The current publication constitutes the third part of a larger study of the Piaractus species. The first part delimited ESU of Piaractus from northern South America (Escobar L et al., 2015), and the second was the taxonomic description of P. orinoquensis and redescription of P. brachypomus (Escobar L et al., 2019). This final part aims to evaluate panmixia of the two Piaractus species and characterize genetic diversity; to identify management units; and finally, to suggest the applications in the fisheries management policies.
Material and methods
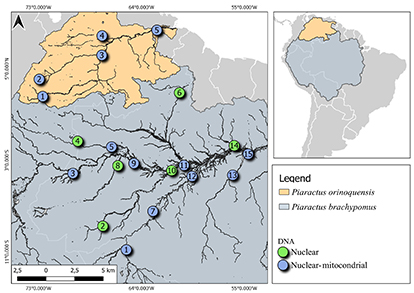
Sampling sites. The Piaractus specimens were collected by artisanal fishermen in the Amazon and Orinoco River basins. The specimens were identified to the species level, and the adipose fin was sampled from each individual according to Machado-Allison (1982). The sample was preserved in 96% ethanol and deposited in the Laboratório de Genética e Evolução Animal (LEGAL) tissue collection of the Universidade Federal do Amazonas, Manaus, Brazil. In the Amazon basin, 15 sampled localities were selected in Brazil and Colombia representing the main channel of the Solimões-Amazonas River and main tributaries. In the Orinoco basin, five localities were sampled, distributed in Colombia and Venezuela, representing the main channel of the Orinoco River to the delta and some of its main tributaries (Fig. 1; Tab. 1). Vouchers specimens are deposited in scientific collections of the Colección de Referencia Ictica de la Estación de Investigaciones Hidrobiológicas de Guayana, San Félix in Venezuela (CRI-EDIHG); Instituto de Ciencias Naturales, Museo de Historia Natural, Facultad de Ciencias, Universidad Nacional de Colombia, Bogotá (ICN-MHN) and Instituto Nacional de Pesquisas da Amazônia, Manaus in Brazil (INPA). Piaractus brachypomus (INPA 39188; 46334; 46335; 57799), and Piaractus orinoquensis (CRI-EDIHG 8195; 8196; ICN-MHN 24051; INPA 57798).
TABLE 1 | Sampling areas of Piaractus analyzed in the Amazon and Orinoco river basins (see Fig. 1).
Basin | River | Map
location | Locality | Latitude | Longitude | n (mtDNA) | n (nDNA) |
Orinoco | Guaviare | 1 | San José del Guaviare | 02º35’5.11”N | 72º38’32.34”W | 23 | 20 |
Meta | 2 | Puerto López | 04º05’34.81”N | 72º56’34.04”W | 23 | 20 | |
Middle Orinoco | 3 | Puerto Carreño | 06º11’18.14”N | 67º28’13.33”W | 18 | 20 | |
Apure | 4 | San Fernado de
Apure | 07º53’22.52”N | 67º26’53.01”W | 16 | 20 | |
Lower Orinoco | 5 | San Felix | 08º22’48.98”N | 62º39’20.73”W | 12 | 15 | |
Amazon | Upper Madeira | 1 | Guajará-Mirim | 10º48’35”S | 65º19’34.31”W | 16 | 20 |
Upper Purus | 2 | Boca de Acre | 08º44’28.37”S | 67º22’59.79”W | – | 20 | |
Solimões | 3 | Leticia | 04º7’55.45”S | 69º59’56.68”W | 10 | 20 | |
Caqueta-Japurá | 4 | La Pedrera | 01º19’10.37”S | 69º35’28.36”W | – | 20 | |
Japurá | 5 | Cidade Japurá | 01º44’45.77”S | 65º50’15.07”W | 9 | 20 | |
Branco | 6 | Parimé River | 02º54’44.41”N | 60º40’45.05”W | – | 20 | |
Middle Madeira | 7 | Humaita | 07º26’43.53”S | 63º0’15.27”W | 9 | 20 | |
Juruá | 8 | Cidade Juruá | 03º27’53.42”S | 66º04’07.23”W | – | 20 | |
Solimões | 9 | Tefé | 03º15’35.20”S | 64º40’5.46”W | 3 | 20 | |
Lower Purus | 10 | Beruri | 03º53’8-56”S | 61º22’4.16”W | – | 20 | |
Amazonas | 11 | Janauacá | 03º25’48.56”S | 60º17’10.81”W | 11 | 20 | |
Low Madeira | 12 | Borba | 04º23’26.12”S | 59º35’10.92”W | 14 | 20 | |
Tapajós | 13 | Itaituba | 04º17’21.09”S | 56º1’48.13”W | 2 | 20 | |
Trombetas | 14 | Óbidos | 01º42’43.60”S | 55º53’45.97”W | – | 20 | |
Amazonas | 15 | Santarém | 02º26’43.34”S | 54º40’34.45”W | 10 | 20 |
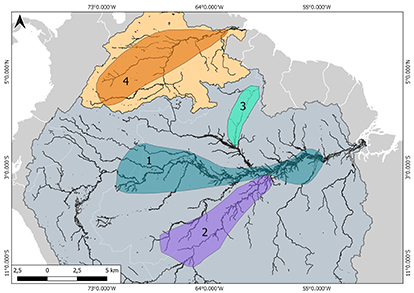
FIGURE 1| Map of the study area for sampling sites for Piaractus brachypomus and P. orinoquensis samples. The number inside the circle indicates the sampling location. The green circle indicates the samples that were analyzed for nDNA and the blue circle the samples analyzed with nDNA and mDNA. Orinoco basin (1. San José del Guaviare, 2. Puerto López, 3. Puerto Carreño, 4. San Fernando de Apure, 5. San Felix). Amazon basin (1. Guajará-Mirim, 2. Boca do Acre, 3. Leticia, 4. La Pedrera, 5. Cidade Japurá, 6. Parimé River, 7. Humaitá, 8. Cidade Juruá, 9. Tefé, 10. Beruri, 11. Janauacá, 12. Borba, 13. Itaituba. 14. Óbidos, 15. Santarém).
DNA extraction was performed using CTAB 2% and phenol chloroform protocols (Doyle, Doyle, 1987; Sambrook et al., 1989). Quality of the DNA was checked in 0.8% agarose gel stained with Gel Red (Biotum), quantified using Nanodrop 2000 (Thermo Scientific, USA) and diluted to final concentration of 50 ng/mL. The amplification of microsatellite markers was done, using markers developed for Piaractus mesopotamicus (Holmberg, 1887) (Calcagnotto et al., 2001) and Colossoma macropomum (Santos et al., 2009). The D-loop mitochondrial region was obtained using the markers developed by Sivasundar et al., (2001) (Tab. S1). The reaction conditions for both types of molecular markers are described in Escobar L et al. (2015). The sequence data were deposited in GenBank under the accession numbers: 870711461, 2400618, 2401711.
Statistical analyses
Nuclear data analyses. A total of 13 microsatellite markers were tested for their polymorphism in the target species; eight loci from Santos et al. (2009) (Cm1A8, Cm1A11, Cm1F4, Cm1F7, Cm1G7, Cm1B8, Cm1C6, Cm2B17) and five loci from Calcagnotto et al. (2001) (Pme4, Pme5, Pme20, Pme21, Pme28).
Microsatellite data were revised for the presence of genotyping errors (null alleles) using the Micro-Checker (Van Oosterhout et al., 2004) and FreeNA (Chapuis, Estoup, 2007) software. Genetic diversity was estimated as allelic richness (A), observed (Ho) and expected (He) heterozygosity, and fixation indices (FIS) were obtained using the Arlequin 3.1 software (Excoffier et al., 2005). The same program was used to test Hardy Weinberg equilibrium (H&W) and linkage disequilibrium between pairs of loci with statistical significance inferred using 10,000 permutations.
We inferred population structure in the Structure software (Pritchard et al., 2000), using the ‘admixture’ and ‘correlated allelic frequencies’ model, testing K=1 to K=9 clusters, repeating each cluster 10 times. The calculation was carried out with 1,000,000 MCMC discarding the first 100,000 permutations as ‘burn-in’. The most likely number of clusters (K) was selected using the model log-likelihood (log (P(X|K)) (Pritchard et al., 2000) implemented in the Pophelper software (Francis, 2017). Additionally, the structure was also estimated through a Discriminant Analysis of Principal Components (DAPC) using the Adegenet package (Jombart, 2008) implemented in R software, version 3.4.2 (R Development Core Team, 2011). To identify groups of individuals we performed two analyses: the Bayesian Information Criterion (BIC) and the optimal number of Principal components (PCs) (Jombart, Collins, 2015). The differentiation of inferred groups was estimated through the FST index in the Arlequin 3.1 software (Excoffier et al., 2005), as well as the association between pair-wise FST values of all sampling localities. The significance was estimated through sequential Bonferroni correction (Rice, 1989).
The correlation between genetic distance and geographic distance was assessed by the Mantel test (Mantel, 1967), which was implemented in the Arlequin 3.1 software (Excoffier et al., 2005), with significance assessed by 10,000 permutations. Significance of pair-wise comparisons was adjusted through sequential Bonferroni correction (Rice, 1989). The geographic distance between collection points was plotted in Google Earth software (http://earth.google.com), following the course of the rivers between each pair of localities.
The connection between the sampled localities was made through the analysis of gene flow between them which was calculated in the Migrate 3.2 software (Beerli, Palczewski, 2010), which we also used to calculated effective population size. The data were converted into effective population size (Ne) and gene flow (Nm) by assuming microsatellite substitution rate of 1.0×10-4 (Chistiakov et al., 2006). The bidirectional model was used to calculate the parameters of each locality, whereas the “panmixia” model was used to find the effective population size.
Analyses of mitochondrial data. The sequences of the control region were compiled, checked, edited, and aligned using Crustal W in the Bioedit 7.2 software (Hall, 1999). Genetic diversity estimates were based on haplotypic diversity (Ĥ) and nucleotide diversity (ᴫ) estimates, which were calculated in the Arlequin 3.1 software (Excoffier et al., 2005) with significance assessed by 10,000 permutations.
A haplotype network was reconstructed using the HapView software (Salzburger et al., 2011) from a maximum likelihood phylogeny inferred in the Treefinder software (Jobb et al., 2004), selecting the HKY85 as the best molecular evolution model for the data obtained in Modeltest software (Posada, Crandall, 1998).
The subpopulation units were identified in the BAPS software (Corander et al., 2008) by using the grouping model for individuals, and by setting the maximum number of clusters to 7 (K=7). We also calculated pair-wise ƟST in Arlequin 3.1 (Excoffier et al., 2005), and assessed significance by 10,000 permutations and corrected by Bonferroni correction (Rice, 1989). Gene flow was carried out in Migrate 3.2 software (Beerli, Palczewski, 2010) using the same parameters described in the nDNA analysis. The mitochondrial substitution rate of 7.0 X 10-9 was assumed for data conversion (Lessios, 2008).
Results
Seven of the 13 tested microsatellite markers were selected for their polymorphism characteristics in the target species (Tab. S1). The analysis in Micro-Checker suggested the excess of homozygotes was possibly caused by the presence of null alleles in loci Cm1B8 and Cm1G7 in P. brachypomus and Pme4, Pme20, Pme21, Pme20 and Cm1B8 in P. orinoquensis. However, the FreeNa analysis suggested non-significant levels of null alleles, indicated by the correlated and non-correlated data, that showed similar Fst values in each basin. In the P. brachypomus, FST=0.052 (CI95%: 0.023-0.104) vs. FST=0.053 (CI95%: 0.022-0.108) did not present significant differences (W-value=2, P > 0.05). In the P. orinoquensis, FST=0.042 (CI95%: 0.015-0.077) vs. FST=0.043 (CI95%: 0.013-0.84) also did not present significant differences (W-value=3, P > 0.05). Based on the verification of the selected markers, mainly FreeNa, they were considered reliable for this study. The linkage disequilibrium in the two species was not significant after Bonferroni correction.
Genetic diversity. Piaractus brachypomus: in the Amazon basin a total of 300 individuals, sampled from 15 localities, were analyzed with seven microsatellite loci, obtaining a total of 79 alleles with 13 alleles on average per locus. Genetic diversity calculated as allelic richness varied from A=5.2 (Boca do Acre) to A=7.8 (Japurá) with an average of A=6.5 alleles per locus; the expected heterozygosity varied from He=0.62 (Parimé) to He=0.79 (Itaitubá) with an average of He=0.72, while the observed heterozygosity ranged from Ho=0.55 (Boca do Acre) to Ho=0.80 (Japurá) with an average of Ho=0.67. Significant deviations from H&W after the Bonferroni correction were observed in three loci at six of the 15 sampled localities: Juruá and Itaituba (Cm1G7), Beruri, Humaitá, and Borba (Cm1F7) and Janauacá (Pme21), so these loci from these localities were not included in the analyses of population structure. The inbreeding coefficient estimates were significant in three of the 15 localities: Humaitá (FIS=0.210), Borba (FIS=0.149) and Santarém (FIS=0.153) (Tab. 2).
TABLE 2 | Genetic diversity parameters of Piaractus brachypomus. Bold values are significant after Bonferroni correction (p<0.0033).
Locations | nDNA | mtDNA | |||||||
n | A | Ho | He | FIS | P<0.05 | n | Ĥ | ᴫ | |
Leticia | 20 | 7.0 | 0.63 | 0.77 | 0.133 | 0.088 | 10 | 1 ± 0.045 | 0.0095 ± 0.005 |
La Pedrera | 20 | 5.7 | 0.74 | 0.72 | -0.025 | 0.691 | – | – | – |
Japurá | 20 | 7.8 | 0.80 | 0.78 | -0.027 | 0.737 | 9 | 1 ± 0.052 | 0.0094 ± 0.005 |
Juruá | 20 | 7.7 | 0.76 | 0.76 | -0.002 | 0.530 | – | – | – |
Tefé | 20 | 6.0 | 0.77 | 0.72 | -0.072 | 0.908 | 3 | 1 ± 0.277 | 0.0092 ± 0.007 |
Beruri | 20 | 6.2 | 0.65 | 0.70 | 0.068 | 0.118 | – | – | – |
Boca do Acre | 20 | 5.2 | 0.55 | 0.63 | 0.115 | 0.040 | – | – | – |
Janauacá | 20 | 5.5. | 0.70 | 0.71 | -0.160 | 0.650 | 11 | 1 ± 0.039 | 0.0083 ± 0.005 |
Parimé | 20 | 5.7 | 0.63 | 0.62 | -0.025 | 0.687 | – | – | – |
Óbidos | 20 | 6.0 | 0.69 | 0.75 | 0.076 | 0.071 | – | – | – |
Guajará-Mirim | 20 | 6.2 | 0.65 | 0.71 | 0.090 | 0.064 | 16 | 1 ± 0.022 | 0.0081 ± 0.005 |
Humaitá | 20 | 6.2 | 0.57 | 0.72 | 0.210 | 0.000 | 9 | 1 ± 0.052 | 0.0102 ± 0.006 |
Borba | 20 | 7.3 | 0.63 | 0.74 | 0.149 | 0.003 | 14 | 1 ± 0.052 | 0.0102 ± 0.006 |
Santarém | 20 | 6.7 | 0.60 | 0.71 | 0.153 | 0.001 | 10 | 1 ± 0.045 | 0.0070 ± 0.004 |
Itaituba | 20 | 7.5 | 0.70 | 0.79 | 0.117 | 0.024 | 2 | 1 ± 0.500 | 0.0113 ± 0.012 |
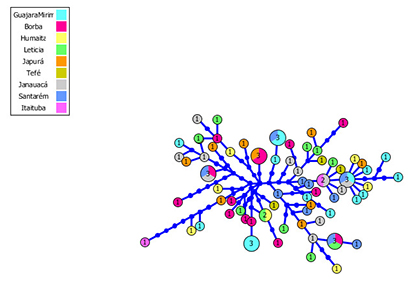
Mitochondrial data of 84 individuals from nine sampled localities of the Amazon basin were analyzed. The sequences were 797 base pairs in size, revealing 70 haplotypes: 8 (11.4%) between two or more localities. Haplotypic diversity in all the localities was Ĥ=1.000, while the nucleotide diversity ranged from ᴫ=0.0070 (Santarém) to ᴫ=0.0113 (Itaituba) with an average of 0.0092 (Tab. 2). The haplotype network showed little clustering of individuals from the same localities and regions, implying a high degree of gene sharing (Fig. 2).
FIGURE 2| Haplotype genealogies of Piaractus brachypomus. The circle size is proportional to the haplotype frequency. Each line represents a single mutation. Colors correspond to localities.
Piaractus orinoquensis: in the Orinoco basin, a total of 95 individuals, sampled from five localities, were analyzed with seven microsatellites markers, obtaining 62 alleles with an average of six alleles per locus. Allelic diversity varied between A=5.57 (Puerto Carreño) and A=6.86 (Guaviare) with an average of A=6.11. The expected heterozygosity ranged from He=0.61 (San Felix) to He=0.70 (Apure) with an average of He=0.66 and observed heterozygosity from Ho=0.42 (San Felix) to Ho=0.64 (Apure) with an average of He=0.55. Significant deviation from H&W was found in some loci in four localities: Meta and Puerto Carreño (Cm1B8), Guaviare (Pme20, Pme21) and San Felix (Pme4, Pme21, Cm1F7) after Bonferroni correction. Inbreeding coefficient was significant in four of the five localities: Meta (FIS=0.19), Puerto Carreño (FIS=0.12), Guaviare (FIS=0.14), and San Felix (FIS=0.32) indicating heterozygote deficiency (Tab. 3).
TABLE 3 | Genetic diversity parameters of Piaractus orinoquensis. Bold values are significant after Bonferroni correction (p<0.01).
Locations | nDNA | mtDNA | |||||||
n | A | Ho | He | FIS | P<0.05 | n | Ĥ | ᴫ | |
Meta | 20 | 5.86 | 0.51 | 0.63 | 0.19 | 0.00 | 23 | 0.971 ± 0.004 | 0.009 ± 0.005 |
Puerto Carreño | 20 | 5.57 | 0.61 | 0.69 | 0.12 | 0.01 | 18 | 0.955 ± 0.003 | 0.012 ± 0.006 |
Apure | 20 | 6.14 | 0.64 | 0.70 | 0.07 | 0.09 | 16 | 1.000 ± 0.022 | 0.011 ± 0.006 |
Guaviare | 20 | 6.86 | 0.59 | 0.69 | 0.14 | 0.00 | 23 | 0.978 ± 0.006 | 0.009 ± 0.005 |
San Felix | 15 | 6.14 | 0.42 | 0.61 | 0.32 | 0.00 | 12 | 0.936 ± 0.008 | 0.010 ± 0.005 |
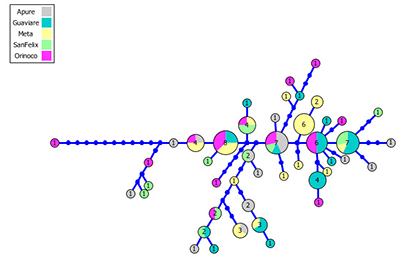
The mitochondrial DNA of 92 individuals from five localities was sequenced and analyzed. The sequences were 509 base pairs; 42 haplotypes were identified of which 11 were shared (26.2%). Haplotypic diversity showed high values of 0.012 and 0.011 (Puerto Carreño and Apure, respectively) and values of 0.009 (Meta and Guaviare) (Tab. 3). The haplotype network also showed low clustering of individuals from different localities although it appeared more extensive than in P. brachypomus (Fig. 3).
FIGURE 3| Haplotype genealogies of Piaractus orinoquensis. The circle size is proportional to the haplotype frequency. Each line represents a single mutation. Colors correspond to localities.
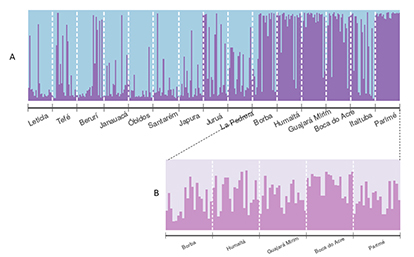
Population structure. In P. brachypomus, the tests applied to determine the degree of “panmixia” showed different values for microsatellite and mtDNA data. The microsatellite analysis detected a moderate population structure supported by a significant value of FST=0.053 (p<0.005). The Bayesian analysis performed in the Structure software showed two subpopulations as the most likely grouping (K=2) (Fig. 4A; Fig. S2). The first subpopulation (blue) was distributed west/east linking the following locations: Leticia, Tefé, Beruri, Janauacá, Óbidos, Santarém, Juruá, Japurá, and La Pedrera – this unit was called “MUs of the sedimentary basin”. The second subpopulation (purple) was distributed north/south and included Borba, Humaitá, Guajará-Mirim, Boca do Acre, and Parimé River; it was called “MUs of the shields”. Itaituba was a transition of the two subpopulations by presenting individuals with similar genotypic proportions between the MUs of the sedimentary basin and the shields (56.6% and 43.4%, respectively), making its assignment to an MU ambiguous, and making the grouping difficult.
FIGURE 4| A. Barplots created from Structure representing assignments of genotypes to each local population for Piaractus brachypomus, the blue color cluster represents individuals related to the Amazon sedimentary basin and the purple color cluster represents individuals related to the crystalline shields of the Amazon basin. B. Barplots hierarchical from crystalline shields of the Amazon basin.
Based on the same microsatellite data, the DAPC also revealed structuring. This procedure uses 56 retained PC and 7 discriminant functions. The result of the BIC showed seven cluster as the most likely (Fig. S3) with few differences between them, while the result of the optimal number of Principal components characterized the overlapping ellipses, with a tendency to overlap in the upper left axis the localities that Structure identified as MUs of the shields, except Parimé had a tendency in the direction of the lower left axis. There was also another overlap towards the lower right axis of the localities that Structure identified as MUs of the sedimentary basin. Of all the analysis of the DAPC, Parimé was the most different conglomerate (Fig. 5). Due to this, a hierarchical analysis only with the locations of the shields was performed using Structure software with the aim of seeing if the north shield was also separated from the south. Bayesian analysis indicated two very similar values of population units (K=1 and K=2), but K=1 was selected as the most likely according to log-likelihood model (Fig. 4B;Fig. S4). Based on these and taking into account the complexity for specific delineation of the MUs of the shields, this was divided into two: one of them “southern” was formed by the locations of Borba, Humaitá, Guajará-Mirim and Boca do Acre and the other “northern” was formed by the Parimé River (Fig. 6). This association of localities was also supported by the significant pairwise FST values (Tab. S5).
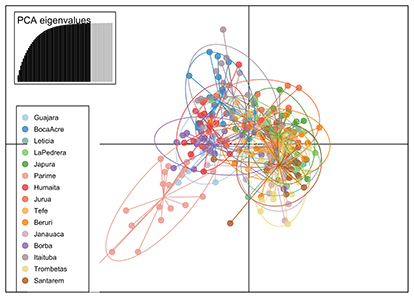
FIGURE 5| Discriminant analysis of principal components (DAPC) based in seven microsatellite loci of 300 individuals of Piaractus brachypomus.
FIGURE 6| Management units of Piaractus. In Piaractus brachypomus the color blue corresponds to the MUs of the sedimentary basin (1), the color violet represents the south shield MUs (2) and the sky-blue color represents the northern shield MUs (3). Piaractus orinoquensis is represented by a single MU indicated in orange (4).
The isolation-by-distance analysis showed as positive and significant correlation between genetic distances and geographic distances (r=0.308, p<0.005). Analyses of the mtDNA data revealed low population structuring, which was expressed by low but significant θST=0.002 (p<0.05) and no difference in pairwise θST values between the localities after Bonferroni correction for multiple comparisons. This difference may be due to the fact that the mitochondrial data did not have the representation of all the localities, and the present ones had a variable sampling number. Despite this, the Bayesian analysis performed with BAPS showed the formation of two biological groups similar to that evidenced by Structure analysis of microsatellite data (Fig. S6;Tab. S7).
Gene flow estimated from microsatellite markers was much lower than for mtDNA data. For microsatellite markers, gene flow values ranged from Nm=0.85 effective migrants (between Santarém and Boca do Acre) to Nm=4.75 (between Beruri and Japurá). In contrast, mtDNA gene flow values ranged from Nm=15.2 effective migrants (between Leticia and Tefé) to Nm=279 (between Guajará-Mirim and Santarém) (Tabs. S8 andS9).
The differences between the markers were also reflected in the effective population size estimates. Estimates based on microsatellite data varied from Ne=641 effective individuals (in Boca do Acre and Janauacá) to Ne=1,218 (in Japurá), and those based on mtDNA data, ranged between Ne=2,127,685 (in Tefé) and Ne=9,340,278 (in Humaitá) (Tabs. S8 and S9).
In P. orinoquensis, the “panmixia” was rejected because the population revealed a discrete structure. Based on nDNA data, AMOVA analysis estimated FST=0.044 (P<0.005), which categorizes the population with low structuring. The pairwise FST values showed a significant difference between San Felix with Meta (FST=0.097), Apure (FST=0.063) and Guaviare (FST=0.087); and between Puerto Carreño with Guaviare (FST=0.058) after Bonferroni correction (Tab. S10). The Bayesian analysis detected a single population unit (K=1) (Fig. S11), but DAPC uses 40 retained PC and 3 discriminant functions and estimated three clusters identified with low support (Fig. S12) while the characterization showed grouping of all localities (Fig. 7). The Mantel test showed lack of significant association between genetic distances and geographic distances (r=0.365, p=0.1961).
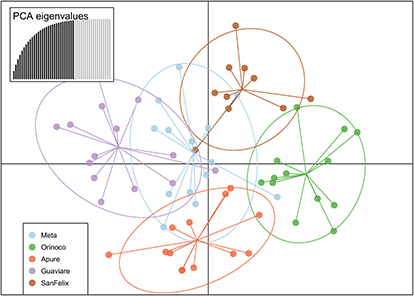
FIGURE 7| Discriminant analysis of principal components (DAPC) based in six microsatellite loci of 95 individuals of Piraractus orinoquensis.
Estimates with mtDNA data also suggested a low level of population structuring θST=0.047 (p<0.005). However, as in nDNA, the pairwise θST values showed significant differences between Meta with Puerto Carreño (θST=0.056), Apure (θST=0.068) and Guaviare (θST=0.083); and between San Felix with Meta (θST=0.107), Puerto Carreño (θST=0.049) and Guaviare (θST=0.11). Analysis in BAPS, however, indicated the existence of a single populational unit (Tab. S13 andFig. S14).
Gene flow estimates differed between the two molecular markers. Gene flow inferred from microsatellite data ranged from Nm=2.033 (from Apure to San Felix) to Nm=7.215 (from Puerto Carreño to Apure), while gene flow inferred from mtDNA ranged between Nm=10.8 (from Meta to Guaviare) and Nm=31.8 (between San Felix and Puerto Carreño) (Tab. S15 and S16). In the same way, effective population size calculated from microsatellite data varied from Ne=1,208 (in Meta) to Ne=2,500 (in Guaviare), and from mtDNA data, the effective population sizes varied between Ne=1,143,667 (Puerto Carreño) and Ne=2,625,222 (Guaviare) (Tabs. S15 and S16).
Discussion
The genetic parameters inferred for the two species from the mtDNA and microsatellite data showed highly contrasting values. These distinctions are related to particular characteristics of each marker such as the maternal inheritance in mtDNA compared to the biparental origin microsatellite loci, and a lack of recombination and low mutation rate in mtDNA (Pesole et al., 1999), as well as the power of temporal resolution, on the historical and contemporary dynamics that each marker provides respectively (Waples, 1998). Therefore, interpretation of the results was done for each marker separately, which allowed us to better understand the characteristics and particularities of each species.
The genetic profile of each Piaractus species was different. Genetic diversity of P. brachypomus was greater based on three of the four estimated parameters (A=6.5; He=0.72; Ĥ=1.000; ᴫ=0.0092) when compared to P. orinoquensis (A=6.1; He=0.66; Ĥ=0.968; ᴫ=0.010). Considering that He is the least biased estimate of diversity, both species had values higher than those recorded for the third species P. mesopotamicus (He=0.60) occurring in the Paraná-Paraguay basin (Calcagnotto, DeSalle, 2009). The diversity of Piaractus is considered moderate taking into account that heterozygosity may be greater than 0.95 in microsatellite markers (Helyar et al., 2011), but within the range of other Amazonian fishes with similar life histories such Colossoma macropomum He=0.78 (Santos et al., 2018), Semaprochilodus insignis (Jardine, 1841)He=0.87 (Passos et al., 2010), Brycon amazonicus (Agassiz, 1829) He=0.83 (de Oliveira et al., 2018), Brachyplatystoma vaillantii (Valenciennes, 1840) He=0.75 (Rodrigues et al., 2009), B. rousseauxii (Castelnau, 1855) He=0.61 (Batista et al., 2010), and B. platynemum Boulenger, 1898 He=0.50 (Ochoa et al., 2015). However, all heterozygosity values of these Amazonian fish were higher than values normally observed for freshwater fish (He=0.46) (DeWoody, Avise, 2000). Such difference can be due to the fact that DeWoody, Avise (2000) analyzed information for temperate zone freshwater fish species that have extensive literature, especially population data, and coexist in an ecological, geographic and evolutionary environment very different from that of Amazonian (Neotropical) fishes. Therefore, it is necessary to have additional data on other Neotropical species to provide information on the patterns of genetic diversity in their river systems for their conservation and management.
Piaractus brachypomus. The population structuring of P. brachypomus is a result of a complex interplay of extrinsic and intrinsic forces that act on the species, such as the geomorphological evolution of the watershed and the natural history of the species. We identified three management units that are associated with different geological formations of the Amazon basin, and this evidence is a clear example of the heterogeneous distribution of genetic diversity at the spatial level (Fig. 6).
The first subpopulation (MUs of the sedimentary basin) includes samples from localities of the main Amazon River channel (Leticia, Tefé, Janauaca, and Santarém) and the lower part of some tributaries, such as Japurá River (La Pedrera, Japurá), Juruá River (Juruá), the lower Purus River (Beruri) and the lower Trombetas River (Óbidos). The geographical area of this subpopulation is characterized by the deposits of Andean and cratonic sediments from the Guiana and Brazilian shields, which form a great flood plain of approximately 100,000km2, and which is closely linked to its tributaries through flood pulse (WCS, 2019). This pattern of genetic continuity was also observed in other Amazonian fish species such C. macropomum (Santos et al., 2018) and B. vaillantii (Formiga et al., 2021), which were also analyzed at the same geographic scale as P. brachypomus.
The second subpopulation (southern shield MU) is found in the rivers that flow across to the Brazilian crystalline shield located to the south of the main channel of the Amazon River, although some of them have Andean origin. This subpopulation comprises locations on the Madeira River (Borba, Humaitá, and Guajará-Mirim) and upper Purus River (Boca do Acre). The middle Madeira River (Humaitá) and upper Purus River (Boca de Acre) were much less differentiated from each other (FST=0.036) than locations on the upper and lower Purus River (Boca do Acre and Beruri, respectively) (FST=0.108). The hypothesis that best explains the similarity derives from the evolutionary history of Amazonia during the rise of the Fitzcarraldo Arch. This geomorphological event divided and changed the drainage orientation in these high Amazonian lands, as well as modifying the associated biota (Espurt et al., 2010; Leite, Rogers, 2013). This gene similarity was also evident in Prochilodus nigricans Spix & Agassiz, 1829 (Machado et al., 2017) and C. macropomum (Santos et al., 2018) two species of Amazonian Characiform fishes.
The Tapajós River, which flows through the Brazilian Shield, showed admixed characteristics between the southern shield and sedimentary basin subpopulations, with a tendency to be similar to the latter. In hydrological terms, the lower Tapajós fraction is strongly influenced by the waters of the Amazon River, in times of flooding (Goulding et al., 2003); this feature allows fishes, in the main channel of the Amazon River, to enter the tributaries whose lower portions act as ecological transition zones. However, in geological terms, the Tapajós River is also connected to the Madeira River tributaries, given the geomorphological complexity of this zone. Although little is known about the fish fauna shared by the Madeira and Tapajós basins, mainly in their middle and low sections, there has been extensive geodispersal due to tributary headwater capture between the two river basins (Dagosta, de Pinna, 2019), similar to what was observed in the current study. Piaractus brachypomus was observed just downstream of the São Luiz rapids of the Tapajós River (ca. 50km upstream of Itaituba, and ca. 200km upstream of the mouth of the Tapajós). The species was not found or reported by other researchers or fishermen upstream; therefore, we considered its distribution in the lower Tapajós to the São Luiz rapids. The study carried out by Barthem et al. (2016), in this same river, mentions that P. brachypomus is an important fishery resource in Itaituba, but it is not recorded in fishery landings from Jacareacanga (upper Tapajós); this finding corroborates the hypothesis in the present study. However, the species distribution list by Dagosta, de Pinna (2019) reports the occurrence of this species in Teles Pires, which is one of the main tributaries of the Tapajós River, a site that we did not investigate. It is also interesting, that Isaac et al. (2015) report the species in the upper reaches of the Xingu and Tocantins rivers and, therefore, it would be interesting to investigate the presence of Piaractus in the headwaters of the Brazilian Shield rivers, to have a better understanding of the genetic patterns in this geographic distribution.
The third subpopulation (northern shield MU) was associated with the Guiana Shield, north of the main Amazon River channel. This subpopulation comprised individuals from the Parimé River and probably individuals from the Branco River, where it discharges, although the Branco River was not sampled in this study. It is the subpopulation with the highest genetic purity among all populations, with genetic proportion of Q=0.949. The separation could be attributed to the chemical barrier produced by the type of water, since the Negro River (black water) separates the Branco-Parimé River system (clear water) from the main channel of the Amazon River (white water), these being the last two natural environments of the species.
In addition to this structure, we found a differentiation in the genetic distribution within each subpopulation. Theoretical studies have proposed models that seek to explain the distribution of the pattern of intraspecific diversity at the level of fluvial landscapes, postulating proposals based on gene flow and effective population sizes (Paz-Vinas et al., 2015). Six models have been postulated: 1) the gene flow model (proposes that genetic diversity increases with increasing downstream flow, and assumes equal Ne in all demes and the migration is asymmetric, being greater downstream than upstream); 2) habitat availability model (proposes that genetic diversity increases with habitat availability, assumes that the greatest genetic diversity is concentrated at the mouth of the river due to the increase in the area downstream, and therefore more available habitat. The model is based on the assumption that migration is equal or symmetric, and effective population sizes are greater downstream); 3) colonization model (postulates the decrease in genetic diversity along colonization routes, assumes that founding populations are found downstream, and assumes equal sizes and migration) and the combination of the three models: 4) gene flow/habitat, 5) gene flow/colonization, habitat/colonization, and 6) gene flow/habitat/colonization (Paz-Vinas et al., 2015).
In this study, the distribution pattern of the intraspecific genetic diversity was not completely similar to the proposed models. The MUs of the sedimentary basin had lower genetic diversity (He) in the lower area of the fluvial system, and higher in locations upstream of the main channel (Santarém He=0.71 downstream vs. Itaituba He=0.79 upstream; Janauacá He=0.71 downstream and Japurá He=0.78 and Leticia He=0.77 upstream) (Fig. 1). This spatial pattern of intraspecific genetic diversity can be compared to the pattern of interspecific diversity proposed by Oberdorff et al. (2019) which indicates that the greatest richness of fish species along the main channel of the Amazon basin is greater in the upstream (western) than in the downstream (eastern) reaches of the basin. Such distribution can be attributed to the evolutionary history of the basin during the Miocene, when the basin was divided by the Purus Arch, causing the waters in the west to flow to the marginal basin (Western Paleo-Amazon) and in the east the waters flow to the Atlantic (Paleo-Eastern Amazon) (Galvis et al., 2006).
However, the apparent trend of genetic distribution may also be strongly determined by the natural history of the species. Studies with different patterns of mobility of Amazonian fish show this, such as pirarucu Arapaima gigas (Schinz, 1822), a resident species, that presents a pattern of population structure from west to east consistent with the geological evidence of the basin, with greater genetic diversity towards the west (Hrbek et al., 2006); while fish with great migrations such as dorado (B. vaillantii) and tambaqui (C. macropomum) did not show a differential genetic distribution pattern along the main channel (Santos et al., 2018; Formiga et al., 2021).
In contrast, the southern shield MU, were closer to the gene flow/habitat model, which has lower genetic diversity in the upstream localities and greater diversity in the downstream localities (Humaitá He=0.72; Guajará-Mirim He=0.71 upstream vs. Borba He=0.74 downstream), with variable estimation of gene flow and effective population sizes.
Finally, the lowest values of gene flow and effective population size were in locations farther from the main channel of the basin and, therefore, with the lowest diversity values of the entire population: Boca do Acre and Parimé (Ne=833.3 – He=0.63 and Ne=958.3 – He=0.62, respectively). This pattern of genetic diversity distribution was more related to the gene flow/colonization model, as large distances limit gene flow, with these locations being the most peripheral of the sampled basin, more ecological and evolutionary factors also influenced the observed pattern. In the case of Parimé River, which is a tributary of the Branco River, both characterized as clear water (Goulding et al., 2003), its connection with the Amazon River is through the Negro River, a black water river, which is a type of water not usual for P. brachypomus, and thereby can act as an ecological filter to the connection with the Amazon, increasing the isolation of this subpopulation. In the case of Boca do Acre, its relationship is closer with the localities of the Madeira River (specifically Humaitá) than with the lower Purus River, due to the biogeographic causes of the basin explained above.
In addition, to the distribution pattern found in P. brachypomus, we show that the possible anthropic actions were also reflected in this population study. Although the size of the population is directly related to genetic diversity, these parameters may also be favored by the proximity to conservation areas, as is the case of Japurá and Itaituba. The individuals of these locations had the largest sizes for the species and benefit the local conservation of genetic diversity and its individuals. In comparison, one of the lowest values was estimated at Janauacá, located near Manaus, Amazonas State, where the commercialization of the fishery resource is greater, which can affect local populations living nearby due to high exploitation.
We also observed that rapids and waterfalls on the Madeira River, mainly the two largest ones (Teotônio and Jirau), as well as chorro Córdoba on the Japurá River (Caquetá, Colombia), and Bem Querer waterfall on the Branco River, were not physical barriers to the movement of P. brachypomus. It was evident that gene flow between locations upstream and downstream of all rapids and waterfalls is enough to keep populations connected, with upstream localities receiving fewer migrants than downstream localities. This dynamic was also detected in C. macropomum in the Madeira River (Farias et al., 2010). The authors found bidirectional migration and concluded that rapids and waterfalls on the Madeira River are not an obstacle for gene interchange in the species. This finding confirms the report by Goulding (1979) of catfish and characid migratory patterns in the Madeira River.
Piaractus orinoquensis. Based on the analyses of both molecular markers, we consider it a single population with discrete structure. This can probably be better understood with a greater number of loci. However, based on the information we propose a single management unit(Fig. 6).
Supported by nDNA data, we found that in P. orinoquensis, the intraspecific genetic distribution did not show any relationship with the proposed theoretical models. In this case, the localities from in the middle section of the basin had the highest values of genetic diversity (Apure He=0.70, Puerto Carreño He=0.69) and not in the lower areas of the basin, as theoretically proposed. This pattern of genetic dispersal occurs in the same direction as in the Amazon with the MUs of the sedimentary basin, in which the genetic diversity was greater in the western section (upstream) of the basin than in the eastern section (downstream). It is possible that this distribution may also be associated with the evolutionary processes of the river system, during the Miocene, when the Orinoco River flowed in a northwesterly direction (Díaz de Gamero, 1996).
However, and surprisingly, Guaviare had the largest effective population size of the entire basin and therefore the highest genetic diversity (Ne=2,500 – He=0.69), although it is located far from the main channel of the basin, to the southwest, it shows a different pattern from all models. This river is one of the largest tributaries of the basin, flowing in the Orinoco-Amazon transition zone, where the Guyana shield extends, and at its confluence unites with the Orinoco River from the east to form the upper Orinoco. Although the evolutionary history of the Orinoco basin is poorly known, it is possible that this pattern of genetic distribution is also closely associated with its formation, since during the Miocene, the Centronics rivers, such as the upper Orinoco, may have been flowing into the great humid system through the Guaviare River (Galvis et al., 2006). In contrast to Guaviare, Meta, also located to the west and far from the main channel, has the smallest effective population size and lowest genetic diversity (Ne=1,208 – He=0.63). This is despite its distance from the main channel being less than Guaviare’s (980km vs. 1,450km, respectively).
We find here another possible explanation for this phenomenon. In this case two factors could have occurred. The first is, that the Meta is a headwater river that flows towards the middle basin with the floodplain and where there is greater genetic diversity, the gene flow decreases as it approaches the upper part of the tributary. In this case the gene flow/habitat model would apply. The second factor that can occur is anthropic actions, in which a difference in Ne is a clear example of the conservation condition of the population (Dudgeon et al., 2012). For example, for more than half a century, fishing at Guaviare was indirectly controlled by illegal armed forces that occupied forest areas in Colombia, acting as an indirect protection for the resource of the resource and that allowed the increase of its populations and maintenance of genetic diversity. Conversely, the population is highly exploited in Meta. Since it is located close to big cities such as Villavicencio and Bogotá, which it supplies with fish (Lasso, Sanchez-Duarte, 2011; Olaya Rodríguez et al., 2017) it caused decrease in genetic diversity. A final possibility is that it is a combination of all the factors explained above.
We also observed that Atures and Maipures waterfalls, in middle Orinoco, do structure P. orinoquensis. Gene flow between locations was bidirectional. The two molecular markers showed greater gene exchange between the local populations of the middle Orinoco (Puerto Carreño and Apure); and the mitochondrial data also showed a great exchange between Puerto Carreño (middle Orinoco) and San Felix (lower Orinoco). This result is to be expected, since the alluvial plain of the Orinoco basin is formed in this area, which increases the connection areas between these localities (Hamilton, Lewis, 1990).
Comparison between species and final considerations. The two species of Piaractus show that the intraspecific genetic distribution patterns cannot be adapted to one a priori model, but that a variety of patterns can occur within the same species. Piaractus brachypomus has higher genetic diversity and is formed by three management units, with a smaller number of effective migrants and smaller effective population sizes by locality in relation to its sister species. Piaractus orinoquensis exhibited moderate gene diversity, it is formed by a single population of discrete structure, with a larger number of effective migrants and a bigger population size by locality. Therefore, it is suggested that such management units be considered the equivalent of fish stock as they are useful for short-term management goals, such as delineation of fishing areas, minimum catch sizes, monitoring and populations status.
In this way, the genetic diversity, and the effective populations sizes of sampled localities in both species indicated that there are some that are highly susceptible to environmental changes and anthropic pressures, such as: Boca do Acre, Parimé River and Janauacá, in the Amazon River basin, and the Meta River drainage, in the Orinoco basin, therefore, appropriate local management is necessary to prevent their collapse. These findings indicate the need for specific management in these locations, aimed at the protection of the species. On the other hand, Japurá and Itaitubá in the Amazon basin, and Guaviare and Apure in the Orinoco basin, can be considered as the local populations with the best genetic attributes and therefore the most recommended to be used in conservation and aquaculture programs of the species, which also serve to guarantee their conservation.
Finally, this study provides new information for better fishery management of these species and provides a preliminary basis for future studies on the importance of protection zones and their relationships with fishing zones, as well as the possible effects of restocking with natural populations. Our recommendation is that management policies be updated based on new scientific information and be coordinated among the countries to establish a trans-national management system for these migratory species.
Acknowledgments
We are grateful to Adam León, Carolina Doria, Damaris Gómez, Edwin Agudelo, Henry Elias Yucuna Cubeo, Maria C. Freitas, Michel Catarino, Nicole Dutra, Rosseval Lette for their support in data collection in the Amazon basin. We also thank Nirson Gonzales, Aniello Barbarino, Rosa E. Ajiaco M., Luz M. Rodrigues and Luis F. Cubillos, for their collaboration with data collections in the Orinoco Basin. Vinicius Verona for making of maps and Donald C. Taphorn and reviewers of this manuscript for suggestions. Conselho Nacional de Desenvolvimento Científico e Técnológico (CNPq 490682/2010–3 to TH; CNPq/PPG7 5570090/2005–9 and CNPq/CT–Amazon 575603/2008–9 to IPF), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES 53923790287) PhD fellowship to MDEL, and CNPq research fellowship to IPF (306804/2013–3) and TH (303646/2010–1).
References
Allendorf FW, Luikart G. Conservation and genetics of populations. J Hered. 2007; 97(6):612–13. https://doi.org/10.1093/jhered/esl039
Barthem RB, Fabré NN. Biologia e diversidade dos recursos pesqueiros da Amazônia. In: Ruffino ML, editor. A pesca e os recursos pesqueiros na Amazônia brasileira, Manaus, Brazil: Ibama/ProVárzea; 2004. p.11–55.
Barthem RB, Ferreira E, Goulding M. As migrações do jaraqui e do tambaqui no rio Tapajós e suas relações com as usinas hidrelétricas. Ocekadi: hidrelétricas, conflitos socioambientais e resistência na bacia do Tapajós. 2016; 479–93.
Barthem RB, Goulding M. An unexpected ecosystem: The Amazon as revealed by fisheries. St. Louis, MO, USA: Amazon Conservation Association and Missouri Botanical Garden Press; 2007.
Batista JS, Farias IP, Formiga-Aquino K, Sousa ACB, Alves-Gomes JA. DNA microsatellite markers for “dourada” (Brachyplatystoma rousseauxii, Siluriformes: Pimelodidae), a migratory catfish of utmost importance for fisheries in the Amazon: development, characterization and inter-specific amplification. Conserv Genet Resour. 2010; 2:5–10. https://doi.org/10.1007/s12686-009-9117-5
Batista VS, Isaac VJ, Fabré N. A produção desembarcada por espécie e sua variação por macroregião Amazônica. In: Batista VS, Issac VJ, editors. Peixes e pesca no Solimões-Amazonas: uma avaliação integrada. Brasília: Ibama/ProVárzea; 2012.
Beerli P, Palczewski M. Unified framework to evaluate panmixia and migration direction among multiple sampling locations. Genetics. 2010; 185(1):313–26. https://doi.org/10.1534/genetics.109.112532
Begg GA, Waldman JR. An holistic approach to fish stock identification. Fish Res. 1999; 43(1–2):35–44. https://doi.org/10.1016/S0165-7836(99)00065-X
Calcagnotto D, DeSalle R. Population genetic structuring in pacu (Piaractus mesopotamicus) across the Paraná-Paraguay basin: evidence from microsatellites. Neotrop Ichthyol. 2009; 7(4):606–16. https://doi.org/10.1590/S1679-62252009000400008
Calcagnotto D, Russello M, DeSalle R. Isolation and characterization of microsatellite loci from Piaractus mesopotamicus and their applicability in other Serrasalminae fish. Mol Ecol Notes. 2001; 1(4):245–47. https://doi.org/10.1046/j.1471-8278.2001.00091.x
Carvajal-Vallejos FM, Zeballos-Fernández AJ. Diversidade y distribución de los peces de la Amazonía boliviana. In: Van Damme PA, Carvajal-Vallejos FM, Molina Carpio J, editors. Los peces y delfines la Amazonía Boliviana: hábitats, potencialidades y amenazas. Cochabamba: Editorial INIA; 2011. p.101–47.
Carvalho GR, Hauser L. Molecular genetics and the stock concept in fisheries. In: Carvalho GR, Pitcher TJ, editors. Molecular Genetics in Fisheries. Dordrecht, Springer; 1995. https://doi.org/10.1007/978-94-011-1218-5_3
Corporación Colombia Internacional (CCI). Pesca y Acuicultura de Colombia 2008. Informe Técnico Regional Litoral Caribe y Pacífico. 2008. p. 1–91. Available from: http://bibliotecadigital.agronet.gov.co/bitstream/11348/4995/2/InformeLitoralCaribePacifico.pdf
Chapuis M-P, Estoup A. Microsatellite null alleles and estimation of population differentiation. Mol Biol Evol. 2007; 24(3):621–31. https://doi.org/10.1093/molbev/msl191
Chistiakov DA, Hellemans B, Volckaert FAM. Microsatellites and their genomic distribution, evolution, function and applications: A review with special reference to fish genetics. Aquaculture. 2006; 255(1–4):1–29. https://doi.org/10.1016/j.aquaculture.2005.11.031
Corander J, Sirén J, Arjas E. Bayesian spatial modeling of genetic population structure. Comput Stat. 2008; 23:111–29. https://doi.org/10.1007/s00180-007-0072-x
Dagosta FCP, de Pinna M. The fishes of the Amazon: Distribution and biogegraphical pattherns, with a comprensive list of species. Bull Am Mus Nat Hist. 2019(431):1–163. https://doi.org/10.1206/0003-0090.431.1.1
Van Damme PA, Carvajal-Vallejos FM, Camacho J, Muñoz H, Coronel J. Peces migratorios de la Amazonía boliviana. In: Van Damme PA, Carvajal-Vallejos FM, Molina Carpio J, editors. Los peces y delfines de la Amazonía Bolivian: hábitats, potencialidades y amenazas. Cochabamba: Editorial INIA; 2011. p.149–202.
DeWoody JA, Avise JC. Microsatellite variation in marine, freshwater and anadromous fishes compared with other animals. J Fish Biol. 2000; 56(3):461–73. https://doi.org/10.1006/jfbi.1999.1210
Díaz de Gamero ML. The changing course of the Orinoco River during the Neogene: a review. Palaeogeogr Palaeoclimatol Palaeoecol. 1996; 123(1–4):385–402. https://doi.org/10.1016/0031-0182(96)00115-0
Doyle JJ, Doyle JL. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull. 1987; 19(1):11–15.
Dudgeon CL, Blower DC, Broderick D, Giles JL, Holmes BJ, Kashiwagi T et al. A review of the application of molecular genetics for fisheries management and conservation of sharks and rays. J Fish Biol. 2012; 80(5):1789–843. https://doi.org/10.1111/j.1095-8649.2012.03265.x
Escobar LMD, Andrade-López J, Farias IP, Hrbek T. Delimiting evolutionarily significant units of the fish, Piaractus brachypomus (Characiformes: Serrasalmidae), from the Orinoco and Amazon river basins with insight on routes of historical connectivity. J. Hered. 2015; 106:428–38. https://doi.org/10.1093/jhered/esv047
Escobar LMD, Ota RP, Machado-Allison A, Andrade-López J, Farias IP, Hrbek T. A new species of Piaractus (Characiformes: Serrasalmidae) from the Orinoco basin with a redescription of Piaractus brachypomus. J Fish Biol. 2019; 95(2):411–27. https://doi.org/10.1111/jfb.13990
Espurt N, Baby P, Brusset S, Roddaz M, Hermoza W, Barbarand J. The Nazca Ridge and uplift of the Fitzcarrald Arch: implications for regional geology in northern South America. In: Hoorn C, Wesselingh FP, editors. Amazonia, Landscape and Species Evolution: A look into the past. Wiley-Blackwell; 2010.
Excoffier L, Laval G, Schneider S. Arlequin ver. 3.0: An integrated software package for population genetics data analysis. Evol Bioinform. 2005; 1:47–50. https://doi.org/10.1177%2F117693430500100003
Farias IP, Torrico JP, García-Dávila C, Santos MCF, Hrbek T, Renno J-F. Are rapids a barrier for floodplain fishes of the Amazon basin? A demographic study of the keystone floodplain species Colossoma macropomum (Teleostei: Characiformes). Mol Phylogenet Evol. 2010; 56(3):1129–35. https://doi.org/10.1016/j.ympev.2010.03.028
Formiga KM, Batista JS, Alves-Gomes JA. The most important fishery resource in the amazon, the migratory catfish Brachyplatystoma vaillantii (Siluriformes: Pimelodidae), is composed by an unique and genetically diverse population in the solimões-amazonas river system. Neotrop Ichthyol. 2021; 19(1):e200082. https://doi.org/10.1590/1982-0224-2020-0082
Francis RM. pophelper: an R package and web app to analyse and visualize population structure. Mol Ecol Resour. 2017; 17(1):27–32. https://doi.org/10.1111/1755-0998.12509
Froese R, Pauly D. Colossoma macropomum (Cuvier, 1816). Cachama. FishBase. 2022. Available from: www.fishbase.org.
Galvis G, Mojica JI, Duque SR, Castellanos C, Sánchez-Duarte P, Mariangeles A et al. Peces del medio Amazonas región de Leticia. Serie de guías tropicales de campo. Bogotá, D.C.: Editorial Panamericana, Formas e Impresos; 2006.
Goulding M. Ecologia de Pesca do Rio Madeira. Manaus: INPA; 1979.
Goulding M, Barthem RB, Ferreira EJG. The Smithsonian atlas of the Amazon. Washington, DC: Smithsonian Institution Press; 2003.
Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser. 1999; 41(1):95–98.
Hamilton SK, Lewis WM. Basin morphology in relation to chemical and ecological characteristics of lakes on the Orinoco River floodplain, Venezuela. Arch Hydrobiol. 1990; 119(4):393–425.
Helyar SJ, Hemmer-Hansen J, Bekkevold D, Taylor MI, Ogden R, Limborg MT et al. Application of SNPs for population genetics of nonmodel organisms: New opportunities and challenges. Mol Ecol Resour. 2011; 11:123–36. https://doi.org/10.1111/j.1755-0998.2010.02943.x
Isaac VJ, Almeida MC, Cruz REA, Nunes LG. Artisanal fisheries of the Xingu River basin in Brazilian Amazon. Brazilian J Biol. 2015; 75(3):125–37. https://doi.org/10.1590/1519-6984.00314BM
Jobb G, von Haeseler A, Strimmer K. TREEFINDER: a powerful graphical analysis environment for molecular phylogenetics. BMC Evol Biol. 2004; 4(1):e18. https://doi.org/10.1186/1471-2148-4-18
Jombart T. adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics. 2008; 24(11):1403–05. https://doi.org/10.1093/bioinformatics/btn129
Jombart T, Collins C. A tutorial for Discriminant Analysis of Principal Components (DAPC) using adegenet 2.0.0. 2015:1–43. Available from: https://adegenet.r-forge.r-project.org/files/tutorial-dapc.pdf
Lasso CA, Agudelo E, Jiménez L, Ramirez-Gil H, Morales-Betancourt M, Ajiaco-Martínez R et al. Catalogo de los recursos pesqueros continentales de Colombia. Bogotá, Instituto de Investigación de Recursos Biológicos Alexander von Humboldt: 2011. Available from: http://hdl.handle.net/20.500.11761/32542
Lasso CA, Sanchez-Duarte P. Los Peces del delta del Orinoco. Diversidad, bioecología, uso y conservación. Caracas, Venezuela: Fundación La Salle de Ciencias Naturales y Chevron C. A.; 2011. Available from: https://www.iucnssg.org/uploads/5/4/1/2/54120303/los_peces_del_delta_del_orinoco._diversi.pdf
Leite RN, Rogers DS. Revisiting Amazonian phylogeography: Insights into diversification hypotheses and novel perspectives. Org Divers Evol. 2013; 13:639–64. https://doi.org/10.1007/s13127-013-0140-8
Lessios HA. The great American schism: Divergence of marine organisms after the rise of the Central American Isthmus. Annu Rev Ecol Evol Syst. 2008; 39(1):63–91. https://doi.org/10.1146/annurev.ecolsys.38.091206.095815
Loubens G, Panfili J. Biologie de Piaractus brachypomus (Teleostei: Serrasalmidae) dans le bassin du Mamoré (Amazonie bolivienne). Ichthyol Explor Freshw. 2001; 12(1):51–64.
Machado-Allison A. Estudios sobre la subfamilia Serrasalminae (Teleostei, Characidae). Parte 1. Estudio comparado de los juveniles de las “cachamas” de Venezuela (Generos: Colossoma y Piaractus). Acta Biol Venez. 1982; 11(3):1–101.
Machado VN, Willis SC, Teixeira AS, Hrbek T, Farias IP. Population genetic structure of the Amazonian black flannelmouth characin (Characiformes, Prochilodontidae: Prochilodus nigricans Spix & Agassiz, 1829): contemporary and historical gene flow of a migratory and abundant fishery species. Environ Biol Fish. 2017; 100(1):1–16. https://doi.org/10.1007/s10641-016-0547-0
Mantel N. The detection of disease clustering and a generalized regression approach. Cancer Res. 1967; 27(2):209–20.
Novoa D. Los recursos pesqueros del eje fluvial Orinoco – Apure: presente y futuro. Caracas, Venezuela: INAPESCA; 2002.
Oberdorff T, Dias MS, Jézéquel C, Albert JS, Arantes CC, Bigorne R et al. Unexpected fish diversity gradients in the Amazon basin. Sci Adv. 2019; 5(9):eaav8681. https://doi.org/10.1126/sciadv.aav8681
Ochoa LE, Pereira LHG, Costa-Silva GJ, Roxo FF, Batista JS, Formiga K et al. Genetic structure and historical diversification of catfish Brachyplatystoma platynemum (Siluriformes: Pimelodidae) in the Amazon basin with implications for its conservation. Ecol Evol. 2015; 5(10):2005–20. https://doi.org/10.1002/ece3.1486
Olaya Rodríguez MH, Escobar Lizarazo MD, Cusva A, Lasso Alcalá CA, Londoño Murcia MC. Mapeo del servicio ecosistémico de alimento asociado a la pesca en los humedales interiores de Colombia. Ecol Austral. 2017; 27(1):123–33. https://doi.org/10.25260/EA.17.27.1.1.261
de Oliveira RC, Santos MCF, Bernardino G, Hrbek T, Farias IP. From river to farm: an evaluation of genetic diversity in wild and aquaculture stocks of Brycon amazonicus (Spix & Agassiz, 1829), Characidae, Bryconinae. Hydrobiologia. 2018; 805:75–88. https://doi.org/10.1007/s10750-017-3278-0
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P. MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes. 2004; 4(3):535–38. https://doi.org/10.1111/j.1471-8286.2004.00684.x
Ortí G, Sivasundar A, Dietz K, Jégu M. Phylogeny of the Serrasalmidae (Characiformes) based on mitochondrial DNA sequences. Genet Mol Biol. 2008; 31(1):343–51. https://doi.org/10.1590/S1415-47572008000200030
Palsbøll PJ, Bérubé M, Allendorf FW. Identification of management using population genetic data. Trends Ecol Evol. 2006; 22(1):11–16. https://doi.org/10.1016/j.tree.2006.09.003
Passos KB, Leão ASA, Oliveira DP, Farias IP, Hrbek T. Polymorphic microsatellite markers for the overexploited Amazonian fish, Semaprochilodus insignis (Jardine and Schomburgk 1841). Conserv Genet Resour. 2010; 2:231–34. https://doi.org/10.1007/s12686-010-9245-y
Paz-Vinas I, Loot G, Stevens VM, Blanchet S. Evolutionary processes driving spatial patterns of intraspecific genetic diversity in river ecosystems. Mol Ecol. 2015; 24(18):4586–604. https://doi.org/10.1111/mec.13345
Pesole G, Gissi C, De Chirico A, Saccone C. Nucleotide substitution rate of mammalian mitochondrial genomes. J Mol Evol. 1999; 48(4):427–34. https://doi.org/10.1007/PL00006487
Posada D, Crandall KA. MODELTEST: Testing the model of DNA substitution. Bioinformatics. 1998; 14(9):817–18. https://doi.org/10.1093/bioinformatics/14.9.817
Pritchard JK, Stephens M, Donnelly P. Inference of population structure using multilocus genotype data. Genetics. 2000; 155(2):945–59. https://doi.org/10.1093/genetics/155.2.945
R Development Core Team. R: The R project for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2011. Available from: https://www.r-project.org/
Rice WR. Analyzing tables of statistical tests. Evolution (NY). 1989; 43(1):223–25. https://doi.org/10.2307/2409177
Rodrigues FC, Farias IP, Batista JS, Alves-Gomes JA. Isolation and characterization of microsatellites loci for piramutaba (Brachyplatystoma vaillantii, Siluriformes: Pimelodidae), one of the commercially most important migratory catfishes in the Amazon basin. Conserv Genet Resour. 2009; 1(1):365–68. https://doi.org/10.1007/s12686-009-9084-x
Ruffino ML, Batista V da S, Isaac VJ, Fabré NN, Almeida OT. Gestão da pesca na Amazônia e seu monitoramento. In: Batista V, Isaac VJ, editors. Peixes e pesca no Solimões-Amazonas uma avaliação integrada. Brasília: MMA/IBAMA; 2012. p.251–76.
Salzburger W, Ewing GB, Von Haeseler A. The performance of phylogenetic algorithms in estimating haplotype genealogies with migration. Mol Ecol. 2011; 20(9):1952–63. https://doi.org/10.1111/j.1365-294x.2011.05066.x
Sambrook J, Fritsch EF, Maniatis T. Molecular cloning: A laboratory manual. vol. 1. 2nd ed. Cold Springs Harbor, NY: Cold Springs Harbor Laboratory Press; 1989.
Santos MCF, Hrbek T, Farias IP. A multilocus approach to understanding historical and contemporary demography of the keystone floodplain species colossoma macropomum (Teleostei: Characiformes). Front Genet. 2018; 9(263):1–17. https://doi.org/10.3389/fgene.2018.00263
Santos MCF, Hrbek T, Farias IP. Microsatellite markers for the tambaqui (Colossoma macropomum, Serrasalmidae, Characiformes), an economically important keystone species of the Amazon River floodplain. Mol Ecol Resour. 2009; 9(3):874–76. https://doi.org/10.1111/j.1755-0998.2008.02331.x
Sivasundar A, Bermingham E, Orti G. Population structure and biogeography of migratory freshwater fishes (Prochilodus: Characiformes) in major South American rivers. Mol Ecol. 2001; 10(2):407–17. https://doi.org/10.1046/j.1365-294x.2001.01194.x
Usma JS, Valderrama M, Escobar MD, Ajiaco-Martínez RE, Villa-Navarro F, Castro F et al. Peces dulceacuícolas migratórios en Colombia. In: Germán Naranjo L, Amaya Espinel JD, editors. Plan Nacional de las Especies Migratorias. Bogotá, Colombia: Ministro de Ambiente, Vivienda y Desarrollo Territorial and World Wildlife Fund Colombia; 2009. p.103–31.
Waples RS. Separating the wheat from the chaff: Patterns of genetic differentiation in high gene flow species. J Hered. 1998; 89(5):438–50. https://doi.org/10.1093/jhered/89.5.438
Wildlife Conservation Society (WCS). Aguas Amazónicas. O maior sistema de água doce do mundo. 2019. Available from: http://pt.aguasamazonicas.org/
Authors
![]() María Doris Escobar L.1
María Doris Escobar L.1 ![]() ,
, ![]() Izeni P. Farias1 and
Izeni P. Farias1 and ![]() Tomas Hrbek1,2
Tomas Hrbek1,2
[1] Laboratório de Evolução e Genética Animal, Departamento de Biologia, Instituto de Ciências Biológicas, Universidade Federal do Amazonas, Av. General Rodrigo Octávio Jordão Ramos, 300, 69077-000 Manaus, AM, Brazil. (MDEL) ptigrinum@yahoo.com (corresponding author), (IPF) izeni@evoamazon.net, (TH) tomas@evoamazon.net.
[2] Biology Department, Trinity University, One Trinity Place, San Antonio, TX, 78212, USA
Authors’ Contribution 

María Doris Escobar L.: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Writing-original draft.
Izeni P. Farias: Formal analysis, Funding acquisition, Methodology, Resources, Validation, Writing-review and editing.
Tomas Hrbek: Funding acquisition, Project administration, Resources, Supervision, Validation, Writing-review and editing.
Ethical Statement
All specimens were collected in accordance with Brazilian laws, using the collection license approved by the Instituto Brasileiro do Meio Ambiente e Recursos Naturais Renováveis (IBAMA #49641–2 and #13048–1). The fish samples were obtained from artisanal fishers, so the research does not need to be approved by the Ethical Committee for animal Use.
Competing Interests
The authors declare no competing interests.
How to cite this article
Escobar L MD, Farias IP, Hrbek T. Genetic comparison of populations of Piaractus brachypomus and P. orinoquensis (Characiformes: Serrasalmidae) of the Amazon and Orinoco basins. Neotrop Ichthyol. 2022; 20(3):e220056. https://doi.org/10.1590/1982-0224-2022-0056
Copyright
This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Distributed under
Creative Commons CC-BY 4.0

© 2022 The Authors.
Diversity and Distributions Published by SBI
![]() Accepted August 22, 2022 by Matt Kolmann
Accepted August 22, 2022 by Matt Kolmann
![]() Submitted June 13, 2022
Submitted June 13, 2022
![]() Epub October 21, 2022
Epub October 21, 2022