![]() Guilherme Moreira Dutra1
Guilherme Moreira Dutra1 ![]() ,
, ![]() George Vita1,
George Vita1, ![]() Péricles Vinícius Gentile1,
Péricles Vinícius Gentile1, ![]() Luz Eneida Ochoa2 and
Luz Eneida Ochoa2 and ![]() Andre Luiz Netto-Ferreira3
Andre Luiz Netto-Ferreira3
PDF: EN XML: EN | Supplementary: S1 | Cite this article
Abstract
A new species of Cyphocharax is described from the Upper Paraíba do Sul River basin, São Paulo, Brazil based on integrated morphological and molecular delimitation criteria. It is morphologically distinguished from its congeners by the presence of a round, dark blotch at the midlength of the caudal peduncle not extending to the proximal portions of the median caudal-fin rays, 19–20 circumpeduncular scales, 34–41 perforated lateral-line scales, 6–7 longitudinal scale rows above and below the lateral line, greatest body depth corresponding to 34.7–39.9% of standard length (SL), and the caudal peduncle depth corresponding to 13.3–15.2% of SL. The lowest genetic distances between the new species and other congeners are: 2.5% from C. gilbert, followed by 3.0% from C. santacatarinae, and 3.2% from C. aff. gilbert. All species delimitation criteria employed herein corroborated the recognition of the new species. In addition, comments on its conservation status are provided.
Keywords: Cyphocharax gilbert clade, Genetic distance, Molecular species delimitation, Taxonomy.
Uma nova espécie de Cyphocharax é descrita do alto rio Paraíba do Sul, São Paulo, Brasil, baseada em dados morfológicos e moleculares integrados. A espécie se distingue morfologicamente de seus congêneres por: apresentar uma mancha escura arredondada na porção média do pedúnculo caudal, não se estendendo à porção proximal dos raios medianos da nadadeira caudal, 19–20 escamas circunpedunculares, 34–41 escamas perfuradas na linha lateral, 6–7 escamas acima e abaixo da linha lateral, altura do corpo correspondendo à 34,7–39,9% do comprimento padrão (CP), e a altura do pedúnculo caudal correspondendo a 13,3–15,2% do CP. As menores distâncias genéticas entre a espécie nova e outras congêneres são: 2,5% de C gilbert, seguida de 3,0% de C. santacatarinae, e 3,2% de C. aff. gilbert. Todos os critérios de delimitação de espécies aqui empregados corroboraram o reconhecimento da nova espécie. Adicionalmente, comentários sobre seu estado de conservação são fornecidos.
Palavras-chave: CladoCyphocharax gilbert, Delimitação molecular de espécie, Distância genética, Taxonomia.
Introduction
The family Curimatidae includes 117 valid species, 14 of which have been described in the past decade (Fricke et al., 2022a). Among the eight curimatid genera, Cyphocharax Fowler, 1906 is the most species-rich, with currently 44 valid species (Fricke et al., 2020b), representing approximately 40% of the family, most of which originate from the Amazon River basin (i.e.,Vari et al., 2012; Wosiacki, Miranda, 2013; Melo, Vari, 2014; Melo, 2017; Bortolo et al., 2018; Bortolo, Lima, 2020). In addition to being the most speciose genus in the family, the genus also has the broadest distribution range within Curimatidae. Most representatives occur throughout South America, including species with cis- and trans-Andeans distribution (Vari, 1992; Vari et al., 2010). Cyphocharax magdalenae (Steindachner, 1878), the species with the northernmost distribution, reaches Costa Rica (Vari, 1992).
Few contributions have focused on the taxonomy of species of Cyphocharax from Brazilian eastern coastal drainages. The most recent studies were made by Vari et al. (2010) and Dutra et al. (2016), which described C. pinnilepis Vari, Zanata & Camelier, 2010 from the de Contas River, C. jagunco Dutra, Penido, Guimarães de Mello & Pessali, 2016 from the Jequitinhonha River, and C. lundi Dutra, Penido, Guimarães de Mello & Pessali, 2016 from the das Velhas River in the Upper São Francisco River basin. Later, Dutra et al. (2017) recognized C. albula (Lütken, 1874) as a senior synonym of C. lundi, resurrected the species, and removed it from the synonymy of C. gilbert (Quoy & Gaimard, 1824) (versus Vari, 1992).
In his revisionary study of the genus, Vari (1992) considered C. gilbert to be a widely distributed species occurring across eastern Brazilian coastal drainages from Bahia to Rio de Janeiro and eastern São Paulo. More recently, Melo et al. (2018) expanded the C. gilbert clade to include all species from coastal drainages from eastern Brazil and the Paraná River and La Plata system, as well as Cyphocharax corumbae (Pavanelli & Britski, 1999). Those authors considered C. gilbert to represent a single species. However, analyses of specimens from the Upper Paraíba do Sul River basin (eastern São Paulo), previously attributed to ‘Cyphocharax gilbert’ indicate that they belong to a distinct and undescribed species, which is described herein.
Material and methods
Morphological analyses. Measurements and counts follow Vari (1992) and Melo, Vari (2014), with the addition of the number of circumpeduncular scales, scale rows around the least depth of the caudal peduncle (Fink, Weitzman, 1974). All measurements were taken point-to-point to the nearest 0.1 mm with digital calipersunder a stereomicroscope with incident light, preferably on the left side. In the description, frequencies are given in parentheses after each count, and an asterisk indicates counts for the holotype. Two specimens were cleared and stained according to Taylor, Van Dyke (1985). Vertebral counts included the four vertebrae of the Weberian apparatus as individual precaudal centra, whereas the fused PU1+U1 was considered a single centrum.
Molecular analyses. Total genomic DNA of seven specimens corresponding to the new species were extracted from muscle tissue according to the Ivanova et al. (2006) protocol (Tab. 1). We optimized PCR conditions to amplify a 657 base pair (bp) fragment of the mitochondrial Cytochrome c oxidase subunit I protein-coding gene (COI) using the primers FishF6 and FishR7 (Jennings et al., 2019). PCR reactions were run in a volume of 12.5 ul, with 1.25 μl of 10X buffer (10 mM Tris-HCl+15 mM MgCl2), 0.5 μl dNTPs (200 nM of each), 0.5 μl each 5 mM primer, 0.05 μl Platinum® Taq Polymerase (Invitrogen), 1 μl genomic DNA (10-50 ng), and 8.7 μl ddH2O. The thermo-cycler profile consisted of an initial denaturation (2 min at 95ºC) followed by 30 cycles of chain denaturation (30 sec at 94ºC), primer hybridization (30 s at 52ºC), nucleotide extension (30–60 sec at 72ºC), and a final extension (8 min at 72ºC). All PCR products were first visually identified on 1% agarose gel and purified using ExoSap-IT® (USB Corporation) following manufacturer instructions. Sequencing was performed at the sequencing facility Centro de Pesquisa sobre o Genoma Humano e Células Tronco at Universidade de São Paulo, São Paulo, Brazil.
TABLE 1 | Tissue samples of Cyphocharax tamuya used in the present study.
Voucher | Tissue
number | GenBank Accession | Locality |
MZUSP 126668 | Mzict006210 | ON997071 | Paraibuna dam |
MZUSP 126668 | Mzict006211 | ON997072 | Paraibuna dam |
MZUSP 126668 | Mzict006212 | ON997073 | Paraibuna dam |
MZUSP 126668 | Mzict006213 | ON997074 | Paraibuna dam |
MZUSP 126668 | Mzict006214 | ON997075 | Paraibuna dam |
MZUSP 126669 | Mzict006215 | OP056028 | Paraibuna dam |
MZUSP 126669 | Mzict006216 | ON997076 | Paraibuna dam |
Alignment and phylogenetic analyses. Consensus sequences for the new species were assembled and edited in Geneious Prime 2022.0.1 and aligned with 203 sequences of cytochrome oxidase (COI) from Melo et al. (2018) (Tab. S1) using the MUSCLE algorithm (Edgar, 2004). A total matrix with 210 samples (see supplementary material) and 657 bp was used to estimate a Maximum Likelihood (ML) tree in RAxML v. 8.0.19 (Stamatakis, 2014) on the 40 CPU 128GB Brycon server at LBP/UNESP. We conducted 20 Maximum Likelihood searches for the phylogenetic tree that best fit the data under the GTRGAMMA substitution model and used non-parametric bootstrapping to assess nodal support by allowing the program to automatically determine the number of replicates using the autoMRE criterion. To test the validity of the new species, we implemented three species delimitation approaches. First, we employed the Bayesian implementation of the Poisson Tree Process model (bPTP, Zhang et al., 2013), using the best ML tree as the input file in the PTP web server (http://species.h-its.org) with default settings. The second method was the Generalized Mixed Yule-Coalescent model (GMYC) (Pons et al., 2006; Fujisawa, Barraclough, 2013). This model aims to discern stochastic birth-death processes separating species from neutral coalescent processes occurring within species, by analyzing time intervals between branching events in a single time-calibrated gene tree (Pons et al., 2006). For this analysis, we estimated a calibrated tree using the lognormal relaxed molecular clock model, which assumes that the rates of molecular evolution are uncorrelated, but log-normally distributed among lineages as implemented in BEAST v1.8.2 (Drummond, Rambaut, 2007). We obtained an ultrametric gene tree with two independent runs of 50 million generations with one tree sampled every 1000th generation. BEAST log files were examined in Tracer v1.6 (Rambaut et al., 2018) to assess stationarity and parameter convergence. A maximum clade credibility tree was estimated in TreeAnnotator v1.8.1, based on the sampled trees after discarding the first 25% generations as burn-in. The ultrametric gene tree was used in the GMYC analysis using a single threshold via GMYC web server. Third, we employed the program Assemble Species by Automatic Partitioning (ASAP) which uses pairwise genetic distances to provide species partitions ranked by a scoring system that uses no biological prior insight about intraspecific diversity (Puillandre et al., 2021). Finally, to contextualize the results of the species delimitation analyses, we estimated pairwise genetic distances for a sub-matrix containing other Cyphocharax species in Mega version 11.0.10 using the Kimura 2-parameter model.
Abbreviations. Institutional abbreviations follow Sabaj (2020). Abbreviations used in the text are: CS = cleared and counterstained specimens, SL = standard length, and HL = head length.
Results
Cyphocharax tamuya, new species
urn:lsid:zoobank.org:act:DF3AFC11-26F6-4F99-8F99-D32390195637
(Figs. 1, 2A,B; Tab. 2)
Cyphocharax gilbert (non Quoy, Gaimard). —Vari, 1992:93 (analyzed specimens from Paraibuna River).
Holotype. MZUSP 125833, 106.8 mm SL, Paraibuna River, Paraibuna, São Paulo, Brazil, 23°22’59”S 45°40’00”W, 19 May 1977, I. Sazima, U. Caramaschi & D. S. Rocha.
Paratypes. All from Brazil, São Paulo, Paraíba do Sul River basin: MZUSP 21668 (3, 87.9–106.8 mm SL), collected with holotype. MZUSP 20686 (5, 83.6–97.9 mm SL), Córrego do Rogero, Santa Branca, ca. 23°22’S 45°44’W, 22 Jan 1961, H. A. Britski. MZUSP 20687 (1, 87.4 mm SL), Córrego do Rogero, Santa Branca, ca. 23°22’S 45°44’W, 14 Jan 1961, H. A. Britski. MZUSP 20729 (3, 88.2–89.9 mm SL), Córrego do Rogero, Santa Branca, ca. 23°22’S 45°44’W, 16 Sep 1962, H. A. Britski & J. Rossi. MZUSP 43822 (5+1CS, 74.1–113.6 mm SL), Paraibuna River, Paraibuna, 23°25’59”S 45°17’59”W, 22 Apr 1992, R. S. Lima. MZUSP 108111 (4+1CS, 64.0–164.5 mm SL), Paraitinga River, São Luiz do Paraitinga, 23°13’34”S 45°20’26”W, 15 Jun 2010, CESP-Paraibuna & ICMBio staffs. MZUSP 108117 (2, 68.5–79.1 mm SL), Paraitinga River, São Luiz do Paraitinga, 23°14’57”S 45°18’49”W, 16 Jun 2010, MZUSP team, CESP-Paraibuna & ICMBio staffs. MZUSP 126668 (5, 155.5–196.2 mm SL), MZUSP 126669 (4, 176.5–178.5 mm SL), Paraibuna dam, below Capim de Angola, Paraibuna, 23°33’50”S 45°35’00”W, 22 Aug 2021, G. Dutra & G. Vita.
FIGURE 1| Lateral view of Cyphocharax tamuya. A. MZUSP 125833, holotype, 106.8 mm SL, Paraibuna River, Paraíba do Sul River basin, São Paulo, Brazil; B. MZUSP 108117, paratype, 69.7 mm SL, Paraitinga River, Paraíba do Sul River basin, São Paulo, Brazil.
Diagnosis. Cyphocharax tamuya differs from most congeners, except C. aninha Wosiacki & Miranda, 2014, C. gilbert, C. gillii (Eigenmann & Kennedy, 1903), C. gouldingi Vari, 1992, C. meniscaprorus Vari, 1992, C. mestomyllon Vari, 1992, C. modestus (Fernández-Yépez, 1948), C. naegelii (Steindachner, 1881), C. oenas Vari, 1992, C. platanus (Günther, 1880), C. santacatarinae (Fernández-Yépez, 1948), C. spilotus (Vari, 1987), C. spiluropsis (Eigenmann & Eigenmann, 1889), and C. spilurus (Günther, 1864) by the absence of distinctive pigmentation of the body and fins combined with a dark blotch on the midlength of the caudal peduncle [vs. possession of a reticulate color pattern formed by pigmentation on the scale borders plus a round dark blotch on the caudal peduncle in C. gangamon Vari, 1992; a longitudinal series of dark blotches on the scales of the flanks plus a caudal peduncle blotch extending from the base of the median caudal-fin rays to the vertical through the posterior limit of the adipose-fin base in C. sanctigabrielis; multiple series of dark stripes or spots running between most scale rows in C. cramptoni Bortolo & Lima, 2020, C. helleri (Steindachner, 1910), C. multilineatus (Myers, 1927), and C. pantostictos Vari & Barriga Salazar, 1990; a patch of dark pigmentation on the dorsal finin C. notatus (Steindachner, 1908)and C. vexillapinnus Vari, 1992; a single dark midlateral stripe in C. boiadeiro Melo, 2017, C. corumbae, C. laticlavius Vari & Blackledge, 1996, C. saladensis (Meinken, 1933), and C. signatus Vari, 1992; a midlateral series of irregular patches of dark pigmentation along the lateral line in C. albula, C. jagunco, C. punctatus (Vari & Nijssen, 1986), and C. vanderi (Britski, 1980); a random pattern of small dark spots on the lateral and dorsolateral surfaces of the body in C. voga (Hensel, 1870); a midlateral spot of dark pigmentation ventral to the dorsal-fin base in C. biocellatus Vari, Sidlauskas & Le Bail, 2012; or the lack of a pronounced caudal-peduncle blotch in C. abramoides (Kner, 1858), C. aspilos Vari, 1992, C. derhami Vari & Chang, 2006, C. festivus Vari, 1992, C. leucostictus (Eigenmann & Eigenmann, 1889), C. magdalenae, C. microcephalus (Eigenmann & Eigenmann, 1889), C. muyrakytan Bortolo, Lima & Melo, 2018, C. nigripinnis Vari, 1992, C. pinnilepis, C. plumbeus (Eigenmann & Eigenmann, 1889), and C. stilbolepis Vari, 1992]. Cyphocharax tamuya can be distinguished from most congeners with a dark blotch on the caudal peduncle, except C. naegelii, by the presence of 19–20 circumpeduncular scales [vs. 16–18 in C. gilbert (n = 43), and 16 in C. modestus (n = 3), C. santacatarinae (n = 10), C. spilotus (n = 8), C. voga (n = 11), and 23–25 in C. platanus (n = 9)]. It differs from C. naegelii by possessing a round caudal peduncle blotch that does not reach the proximal portions of the median caudal-fin rays [vs. a horizontally elongate blotch that extends onto the median caudal-fin rays (Fig. 2)], the greatest body depth corresponding to 34.7–39.9% of SL (vs. 29.0–33.0% of SL), the caudal peduncle depth corresponding to 13.3–15.2% of SL (vs. 12.0–13.0% of SL), and the presence of 33 vertebrae (vs. 34). The new species differs from C. platanus by the presence of 34–41 pored lateral-line scales from the supracleithrum to the hypural joint (vs. 48–54), and the presence of 6–7 scale rows between the lateral-line scale series and the anal-fin origin (vs. 8–10). Furthermore, the new species differs from C. voga by the presence of 33 vertebrae (vs. 34–37), from C. santacatarinae by the presence of 6–7 scale rows between the lateral-line scale series and anal-fin origin (vs. 4–5), and from C. spilotus by the presence of 34–41 pored lateral-line scales from the supracleithrum to the hypural joint (vs. 29–33).
FIGURE 2| Caudal peduncle and fin of Cyphocharax species: A. C. tamuya, MZUSP 108117, paratype; B. C. voga, MZUSP 39782; C. C. santacatarinae, MZUSP 58659; D. C. gilbert, MZUSP26861; E. C. naegelii, MZUSP 207752; F. C. platanus, MZUSP 45858.
Description. Morphometric data for holotype and 24 paratypes presented in Tab. 2. Body somewhat elongate. Greatest body depth at vertical through dorsal-fin origin. Dorsal profile of head strongly convex from margin of upper lip to vertical through anterior nostril, slightly concave to nearly straight from that point to supraoccipital spine. Dorsal profile of body slightly convex from that point to dorsal-fin terminus; nearly straight from that point to adipose-fin origin and then slightly concave to origin of anteriormost dorsal caudal-fin procurrent ray. Ventral profile of body convex from lower lip to anal-fin terminus; concave from that point to origin of anteriormost ventral caudal-fin procurrent ray. Prepelvic region smoothly flattened transversely. Postpelvic region of body transversely round.
Head compressed, pointed overall in lateral view. Mouth subterminal, located at horizontal through ventral margin of pupil. Upper jaw slightly longer than lower jaw. Nostrils close together and separated by thin flap of skin. Anterior nostril circular, near midpoint between snout tip and anterior margin of eye. Posterior nostril crescent shaped. Adipose eyelid more developed anteriorly, with vertically ovoid opening near center of eye. Eye on anterior one-half of head length. Branchial membranes joined at isthmus. Branchiostegal rays 4(2).
Scales cycloid. Lateral-line series with 34(1), 36(5), 37(9), 38*(7), 39(2) or 41(1) perforated scales from supracleithrum to hypural joint; perforated scales posterior to hypural joint 1(2), 2*(13), 3(8) or 4(1) [zero in one specimen]. Scale rows between dorsal-fin origin and lateral line 6*(11) or 7(14). Scale rows between anal-fin origin and lateral line 6*(12) or 7(13). Scales between anus and anal-fin origin 2*(20) or 3(5). Predorsal scales 11(2), 12(1), 13*(14), 14(7) or 16(1). Circumpeduncular scales 19(3) or 20*(22). Scales covering basal portion of caudal-fin rays distinctly smaller than those on caudal peduncle.
Pectoral fin pointed, with i,12(5), 13*(12), 14(7) or 15(1) rays; tip of adpressed pectoral fin reaching vertical between eighth to tenth lateral line scale. Supraneurals 5(2), first and second inserted immediately anterior to neural spine of 5th vertebra. Dorsal fin pointed, with ii,9*(24) or 10(1) rays; first unbranched ray about one-half length of second unbranched ray, second unbranched and first branched rays longest; branched rays gradually decreasing in length posteriorly. Dorsal-fin pterygiophores 11(2); first inserted immediately posterior to neural spine of 9th(2) vertebra. Pelvic fin emarginate, with i,8*(25) rays; fin origin at vertical through third or fourth branched dorsal-fin ray. Tip of adpressed pelvic fin falling short of anus by 1*(6), 2(12) or 3(5) scales. Anal fin emarginate, with ii,7*(25) rays; first unbranched ray about one-third length of second unbranched ray; second unbranched ray and first branched rays longest, subsequent branched rays gradually decreasing in length. Anal-fin pterygiophores 8(1) or 9(1); first inserted immediately posterior to haemal spine of 23rd(2) vertebra. Adipose fin present. Caudal fin with i,9/8,i*(25) rays; forked with lobes somewhat pointed. Dorsal procurrent rays of caudal-fin 8(2); ventral procurrent rays of caudal-fin 5(2). Total vertebrae 33(2). Precaudal vertebrae 20(1) or 21(1), caudal vertebrae 12(1) or 13(1).
TABLE 2 | Morphometric data for Cyphocharax tamuya (n = total number of specimens examined; SD = Standard deviation); ranges include values for the holotype.
Holotype | Range | n | Mean | SD | |
Standard length (mm) | 106.8 | 64.0–164.5 | 20 | – | – |
Percent of standard length | |||||
Greatest body depth | 36.7 | 34.7–41.1 | 25 | 37.8 | 1.7 |
Snout to dorsal-fin origin | 50.4 | 47.4–53.7 | 25 | 50.9 | 1.5 |
Snout to pectoral-fin origin | 29.9 | 27.0–32.5 | 25 | 29.6 | 1.3 |
Snout to pelvic-fin origin | 56.4 | 54.0–58.1 | 25 | 56.0 | 1.1 |
Snout to anal-fin origin | 79.5 | 79.5–85.1 | 25 | 82.0 | 1.4 |
Snout to anus | 76.1 | 69.9–81.5 | 25 | 77.8 | 2.4 |
Dorsal-fin origin to hypural
joint | 56.3 | 54.8–59.2 | 25 | 57.2 | 1.3 |
Dorsal-fin origin to anal-fin origin | 44.0 | 44.0–50.7 | 25 | 47.4 | 1.6 |
Dorsal-fin origin to pelvic-fin origin | 35.8 | 33.5–40.7 | 25 | 37.2 | 2.2 |
Dorsal-fin origin
to pectoral-fin origin | 36.2 | 31.9–38.2 | 25 | 35.7 | 1.7 |
Caudal-peduncle depth | 13.7 | 13.3–15.2 | 25 | 14.2 | 0.5 |
Pectoral-fin length | 18.6 | 16.1–19.6 | 24 | 18.2 | 0.8 |
Pelvic-fin length | 19.1 | 17.3–22.0 | 25 | 19.6 | 1.0 |
Dorsal-fin length | 26.3 | 24.5–28.8 | 25 | 26.3 | 1.1 |
Head length | 29.9 | 26.2–32.3 | 25 | 29.5 | 1.2 |
Percent of head length | |||||
Snout length | 28.2 | 22.8–33.8 | 25 | 28.8 | 2.3 |
Orbital diameter | 31.6 | 21.7–33.5 | 25 | 28.7 | 2.8 |
Postorbital length | 46.2 | 42.2–48.5 | 25 | 45.7 | 1.8 |
Interorbital distance | 41.1 | 38.1–47.4 | 25 | 41.6 | 2.5 |
Coloration in alcohol. Specimens retain silvery guanine deposits on infraorbitals and opercular series. Background coloration of body tan. Dark coloration on dorsal portion of head, maxilla, upper lip and infraorbital 1, lighter ventrally. Dark chromatophores on postorbital region of head slightly larger than those on snout. Overall pigmentation of latter portion of postorbital region consequently lighter than adjoining surrounding areas. Scale rows from predorsal to lateral-line series with dark chromatophores concentrated at focus of each scale. Scales below lateral line with few, sparse chromatophores, lacking distinctive pigmentation pattern. Lateral surface of body of small specimens (up to 69.7 mm SL; Fig. 2B) with faint dark midlateral stripe up to 2 scales deep running from vertical through dorsal-fin insertion to anterior margin of caudal peduncle blotch. Dark midlateral stripe inconspicuous or absent in specimens larger than 70 mm SL. Midlateral surface of caudal peduncle with dark round blotch, not extending onto proximal portions of median caudal-fin rays. Caudal peduncle blotch more conspicuous in juveniles (Fig. 2B). Distal tip of dorsal fin with discrete, dark chromatophores on rays and associated membranes. Anal-fin rays with dark chromatophores on distal tip, more concentrated on anteriormost rays. Adipose, caudal, pectoral and pelvic fins overall hyaline. Caudal fin with discrete dark chromatophores more concentrated on lower caudal-fin lobe in some specimens.
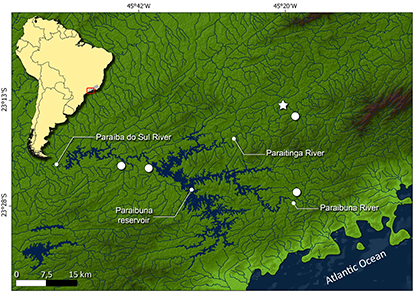
Geographical distribution. Cyphocharax tamuya is known from the Paraíbuna and Paraitinga rivers, two main headwater tributaries of the Paraíba do Sul River, São Paulo, Brazil (Fig. 3).
FIGURE 3| Map of eastern portion of South America showing the type locality (star) and distribution of Cyphocharax tamuya. Symbols may represent more than one collection event.
Etymology. The epithet “tamuya” honours the indigenous Tamuya people. The word Tamuya, from Tupinambá language, means “grandfather or the oldest”. Linguistics variations to Tamuya include “Tamúya”, “Tamujas” or “Tamuía”, all keeping the same origin and meaning. The Tamuyas were driven extinct during the colonization of the Brazilian coast in the seventeenth century. Before their extinction, they lived in the area of occurrence of Cyphocharax tamuya. A noun in apposition.
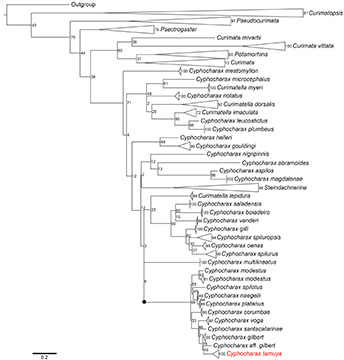
FIGURE 4| Maximum likelihood tree of Curimatidae using COI (210 sequences with 657 bp) depicting the Cyphocharax gilbert clade (black dot). Numbers at right of nodes represent bootstrap support.
Conservation status. The Upper Paraíba do Sul River basin area was impacted in the 1970’s by the construction of the Paraibuna Dam, which changed the water dynamics in the area. However, considering that Cyphocharax species inhabit lentic waters, the dam may not impact the population of C. tamuya in that area. In fact, part of the specimens examined herein originate from inundation zone above the dam and were collected after the dam’s construction (e.g., MZUSP 21668). Furthermore, the species also occurs in at least three localities upstream of the dam. Thus, we recommend that C. tamuya be classified as Least Concern (LC), according to the International Union for Conservation of Nature (IUCN) categories and criteria (IUCN Standards and Petitions Subcommittee, 2019).
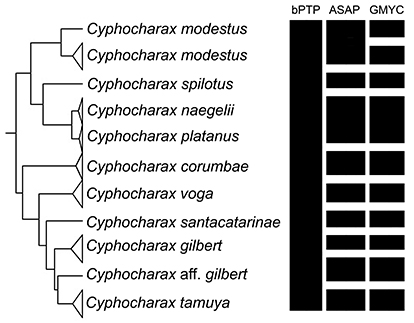
FIGURE 5| Results of the molecular delimitations methods bPTP, ASAP and GMYC for the species included in the Cyphocharax gilbert clade. MOTUs are represented by black rectangles.
TABLE 3 | Average barcoding divergence (%) among species of the Cyphocharax gilbert clade based on the Kimura two-parameters model using COI data matrix. Values in bold refer to intraspecific distance within each species. Values above the diagonal refer to standard deviation.
1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | ||
1 | C. corumbae | 0.001 | 0.001 | 0.002 | 0.002 | 0.001 | 0.004 | 0.003 | 0.000 | 0.002 | 0.004 | 0.002 |
2 | C. gilbert | 0.036 | 0.002 | 0.001 | 0.001 | 0.001 | 0.001 | 0.001 | 0.001 | 0.001 | 0.004 | 0.002 |
3 | C. aff. gilbert | 0.048 | 0.019 | – | 0.002 | – | 0.001 | 0.001 | – | – | 0.005 | 0.002 |
4 | C. modestus | 0.033 | 0.035 | 0.048 | 0.005 | 0.001 | 0.001 | 0.002 | 0.001 | 0.002 | 0.005 | 0.001 |
5 | C. cf. modestus | 0.033 | 0.038 | 0.048 | 0.013 | – | 0.001 | 0.000 | – | – | 0.005 | 0.001 |
6 | C. naegelii | 0.020 | 0.036 | 0.047 | 0.028 | 0.027 | 0.002 | 0.001 | 0.001 | 0.001 | 0.005 | 0.002 |
7 | C. platanus | 0.024 | 0.039 | 0.047 | 0.030 | 0.031 | 0.005 | 0.001 | 0.000 | 0.000 | 0.005 | 0.002 |
8 | C. santacatarinae | 0.039 | 0.021 | 0.031 | 0.042 | 0.041 | 0.036 | 0.036 | – | – | 0.005 | 0.002 |
9 | C. spilotus | 0.030 | 0.040 | 0.049 | 0.033 | 0.031 | 0.026 | 0.029 | 0.042 | – | 0.006 | 0.002 |
10 | C. tamuya | 0.053 | 0.025 | 0.032 | 0.051 | 0.050 | 0.049 | 0.053 | 0.030 | 0.053 | 0.004 | 0.005 |
11 | C. voga | 0.036 | 0.025 | 0.030 | 0.045 | 0.041 | 0.035 | 0.035 | 0.026 | 0.039 | 0.039 | 0.003 |
Molecular species delimitation. The molecular dataset included 210 sequences of 657 base pairs in length. Stop codons, deletions or insertions were not observed in any sequences. In the matrix 386 positions were conserved and 271 were variable, with base compositions of 23% adenine, 27.9% cytosine, 30.1% thymine and 18.4% of guanine. The data were not saturated as evaluated via the Iss.c value. The maximum likelihood tree from RAxML shows the same topology obtained in the ultrametric tree from BEAST and the three delimitation analyses (Fig. 4). The bPTP approach delimited 95 lineages, and Cyphocharax tamuya was grouped with the remaining species composing the C. gilbert clade of Melo et al. (2018) with a posterior probability of 0.926 (Fig. 5). In contrast, the results from the ASAP analysis showed a low asap score of 4.00 with a p-value 0.0063, and delimited 101 species from the total dataset (six more than did bPTP). The ASAP analysis delimited all species belonging to the C. gilbert clade as unique lineages except for C. naegelii and C. platanus which were grouped in a single lineage. The best fitting model in the GMYC analysis received a likelihood score of 1471.86 in comparison with the null model of 1435.10. The GMYC analysis identified a total of 106 entities with a confidence interval of 103-111 and a threshold time of -0.007. This analysis recognized the new species as a differentiated entity, again grouped C. naegelii and C. platanus in the same lineage, and recovered two distinct lineages within C. modesus. Regarding the genetic distances from a sub-matrix containing the COI sequences from the all known species in C. gilbert clade (Tab. 3), the overall mean distance among species was 3%. The lowest genetic distance between C. tamuya and any other known species was 2.5% (from C. gilbert), followed by 3% from C. santacatarinae, and 3.2% from C. aff. gilbert.
Discussion
Vari (1989) hypothesized the Cyphocharax gilbert group as including C. gilbert, C. modestus, C. santacatarinae, and C. voga, based on the shared presence of a round caudal peduncle blotch. Vari also suggested C. naegelii and C. platanus as close relatives of the C. gilbert group based on overall similarity. Recently, Melo et al. (2018) corroborated the C. gilbert group as a well-supported monophyletic clade within Curimatidae, also including C. corumbae, C. naegelii, C. platanus,and C. spilotus, in agreement with the morphological hypothesis of Vari (1989). The maximum likelihood analysis presented herein recovered a similar topology to that proposed by Melo et al. (2018). Despite the methodological limitations of single locus analyses in phylogenetic inferences (e.g., Hajibabaei et al., 2006, 2007), it is important to note that the present analysis supports the assignment of C. tamuya to the C. gilbert clade and adds molecular evidence to the morphological similarity to support the hypothesis of a close relationship between these species.
In his revisionary study of Cyphocharax, Vari (1992) interpreted the conspicuous variation in the shape of the dark blotch on the caudal peduncle among the specimens of ‘C. gilbert’ as reflecting intra-specific plasticity. Upon the revalidation of C. albula, Dutra et al. (2017) presented evidence partially refuting Vari’s delimitation of C. gilbert, which the description of C. tamuya herein further corroborates. In short, Vari’s concept of C. gilbert subsumed several entities now recognized as distinct species.
The examination of specimens corresponding to C. gilbert across its distribution revealed that specimens from coastal rivers between the Paraíba do Sul and Doce rivers drainages present a dark round blotch on the caudal peduncle that never reaches the proximal portion of the median caudal-fin rays. The holotype of C. gilbert demonstrates a similar pattern. In contrast, at least part of the specimens distributed in coastal rivers of northeastern Brazil between the Mucuri River (Gomes et al., 2015), and the de Contas River (Vari, 1992), possess an elongate caudal peduncle blotch extending onto the median caudal-fin rays. This pattern differs from the holotype of C. gilbert and suggests that these northeastern specimens may not be conspecific with C. gilbert. The single locus (COI) evidence presented herein also supports a subdivision of Vari’s concept of C. gilbert, by recovering a genetic difference (1.9%) between C. aff. gilbert from the Mucuri River and specimens of C. gilbert from closer to the type locality. That result supports Gomes et al. (2015), suggestion that C. aff gilbert represents another distinct species. Our single-locus phylogeny places this species as the sister group of C. tamuya, with this clade being the sister of C. gilbert from the Paraíba do Sul River basin and coastal rivers eastern Brazil between the Paraíba do Sul and the Baía de Guanabara, though this placement requires testing with a multilocus dataset. Further examination of some of the specimens that Vari included in his study (MZUSP 20686, 20687, 20729, 21668), revealed that a higher number of circumpeduncular scaleseasily distinguishes C. tamuya from C. gilbert and C. aff. gilbert. Vari (1992) did not examine this character.
Overall, the present results highlight the need for a comprehensive taxonomic review of the C. gilbert group using integrative taxonomic approaches. Such review would not only clarify the currently neglected diversity under this name, but also allow proper conservation strategies for the species occurring in deeply populated areas in Brazil, where rising urbanization, agriculture and river damming pressure the known and unknown components of ichthyological diversity.
Comparative material examined. In addition to the comparative material examined listed in Dutra et al. (2016, 2017), the following species were examined, all from Brazil: Cyphocharax gilbert: MNHN 5430, holotype of Curimatus gilbert, 83.2 mm SL, Macacu River. MZUSP 26861 (1, 105.4 mm SL), Macacu River. MZUSP 114685 (1, 127.1 mm SL), MZUSP 114744 (4, 152.5–165.5 mm SL), Paraíba do Sul River. MCNIP 207 (1, 143.3 mm LEA), MCNIP 208 (2, 81.2–88.5 mm LEA), MCNIP 209 (1, 87.6 mm LEA), MCNIP 455 (10, 96.3–116.5 mm LEA), MCNIP 456 (1, 117.5 mm LEA), MCNIP 457 (2, 107.0–107.5 mm LEA), MCNIP 458 (1, 116.2 mm LEA), MZUSP 51524 (2, 88.6–108.0 mm LEA), MZUSP 66167 (1, 170.1 mm LEA), MZUSP 66169 (1, 121.2 mm LEA), MZUSP 90769 (11+2CS, 39.5–99.9 mm LEA), MZUSP 101026 (2, 73.3–77.2 mm LEA), Doce River. Cyphocharax aff. gilbert:MCNIP 1488 (1, 105.1 mm SL), MCNIP 1489 (2, 97.9– 110.2 mm SL), Mucuri River. MZUSP 58873 (20, 56.9–80.8 mm SL), MZUSP 58881 (2, 82.6–83.8 mm SL), Pratinha River. MZUSP 86101 (6, 35.1–47.5 mm SL), Paraguaçu River. MZUSP 87891 (1, 84.9 mm SL), MZUSP 102558 (1, 82.0 mm SL), MZUSP 102599 (1, 94.6 mm SL), de Contas River. MZUSP 102644 (1, 79.4 mm SL), Rio Almada. Cyphocharax modestus: MZUSP 1350 (1 paratype, 94.9 mm SL), MZUSP 5250 (1 paratype, 90.5 mm SL), MZUSP 5251 (1 paratype, 82.6 mm SL), Batalha River. Cyphocharax naegelii: NWM 68808 (holotype of Curimatus naegelii, 110.0 mm SL), “Vicinity of [de] Rio Janeiro”. MZUSP 20692 (8, 92.4–137.1 mm SL), Cachoeira das Emas. MZUSP 20703 (1, 134.6 mm SL), MZUSP 20752 (38+2CS, 92.5–101.6 mm SL), Mogi-Guaçu River. Cyphocharax platanus: MZUSP 1735 (3, 136.5–149.0 mm SL), MZUSP 1431 (2, 130.7–145.9 mm SL), Uruguai River. MZUSP 45858 (84.2–104.6 mm SL), Uruguai River. Cyphocharax santacatarinae: MZUSP 58659 (1, 94.3 mm SL), Iporanga River. MZUSP 70427 (10, 129.6–159.5 mm SL), Ribeira de Iguape River. Cyphocharax spilotus: MZUSP 37133 (holotype of Curimata spilota, 58.9 mm SL), MZUSP 37134 (7 paratypes of Curimata spilota, 36.0–46.5 mm SL), Santa Maria River. Cyphocharax voga: MZUSP 39782 (10, 87.2–126.4 mm SL), arroio do Salso.
Acknowledgments
The authors are thankful to the following individuals and institutions for access to material under their care: G. Santos (MCNIP), W. Wosiacki and I. Maschio (MPEG), A. Datovo, M. Gianetti, and O. Oyakawa (MZUSP), and Anja Palandacic (NMW). We also thank L. Peixoto (MZUSP) and Anja Palandacic (NMW) for kindly providing information on the types of Curimatus gilbert and C. naegelii, respectively. We finally thank D. Canepelle and O. Oyakawa for assisted with the acquisition of fresh specimens of C. tamuya. The authors were financially supported by the Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP grant # 2018/09445–9 to GMD, and #2019/23167–4 to PVG), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq process 313834/2021-0 to ALNF), and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES process 88882.377148/2019–01 to GV). This contribution was also supported by the Diversity and Evolution of Gymnotiformes Project (FAPESP/Smithsonian process 2016/19075–9).
References
Bortolo GC, Lima FCT. A new species of Cyphocharax (Characiformes: Curimatidae) with a horizontal color pattern from the rio Tapajós drainage, Amazon basin, Brazil. Neotrop Ichthyol. 2020; 18(2):e190135. http://doi.org/10.1590/1982-0224-2019-0135
Bortolo GC, Lima FCT, Melo BF. A new Cyphocharax from the Lower rio Tapajós, Amazon basin, Brazil (Characiformes: Curimatidae). Copeia. 2018; 106(2):346–52. http://doi.org/10.1643/CI-17-656
Drummond AJ, Rambaut A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol. 2007; 7:214. https://doi.org/10.1186/1471-2148-7-214
Dutra GM, Penido IS, Mello GCG, Pessali TC. Two new species of Cyphocharax (Teleostei: Characiformes: Curimatidae) from headwaters of the Jequitinhonha and São Francisco River basins, Minas Gerais, Brazil. Zootaxa. 2016; 4103(2):154–64. http://doi.org/10.11646/zootaxa.4103.2.5
Dutra GM, Penido IS, Pessali TC, Netto-Ferreira. Ressurrection of Curimatus albula Lütken (Characiformes: Curimatidae), a senior synonym of Cyphocharax lundi Dutra, Penido, Mello & Pessali. Zootaxa. 2017; 4344(3):597–600. https://doi.org/10.11646/zootaxa.4344.3.10
Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004; 32(5):1792–97. http://doi.org/10.1093/nar/gkh340
Fink WL, Weitzman SH. The so-called cheirodontin fishes of Central America with descriptions of two new species (Pisces: Characidae). Smithson Contrib Zool. 1974; 172:1–46. http://doi.org/10.5479/si.00810282.172
Fricke R, Eschmeyer WN, Van der Laan R. Eschmeyer’s catalog of fishes: species by family/subfamily [Internet]. San Francisco: California Academy of Sciences; 2020a. Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/SpeciesByFamily.asp
Fricke R, Eschmeyer WN, Van der Laan R. Eschmeyer’s catalog of fishes: genera, species, references [Internet]. San Francisco: California Academy of Sciences; 2020b. Available from: http://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp
Fujisawa T, Barraclough TG. Delimiting species using single-locus data and the generalized mixed yule coalescent approach: A revised method and evaluation on simulated data sets. Syst Biol. 2013; 62(5):707–24. http://doi.org/10.1093/sysbio/syt033
Gomes LC, Pessali TC, Sales NG, Pompeu PS, Carvalho DC. Integrative taxonomy detects cryptic and overlooked fish species in a neotropical river basin. Genetica. 2015; 143:581–88. http://doi.org/10.1007/s10709-015-9856-z
Hajibabaei M, Singer GAC, Hickey DA. Benchmarking DNA barcodes: An assessment using available primate sequences. Genome. 2006; 49(7):851–54. http://doi.org/10.1139/g06-025
Hajibabaei M, Singer GAC, Herbert PND, Hickey DA. DNA barcoding: How it complements taxonomy, molecular phylogenetics and population genetics.Trends in Genetics. 2007; 23(4):167–72. https://doi.org/10.1016/j.tig.2007.02.001
International Union for Conservation of Nature (IUCN). Standards and petitions subcommittee. Guidelines for using the IUCN Red List categories and criteria. Version 14 [Internet]. Gland; 2019. Available from: https://www.iucnredlist.org/resources/redlistguidelines
Ivanova NV, Dewaard J, Hebert PDN. An inexpensive, automation-friendly protocol for recovering high-quality DNA. Mol Ecol Notes. 2006; 6(4):998–1002.http://doi.org/10.1111/j.1471-8286.2006.01428.x
Jennings WB, Ruschi PA, Ferraro G, Quijada CC, Silva-Malanski ACG, Prosdocimi F, Buckup PA. Barcoding the Neotropical freshwater fish fauna using a new pair of universal COI primers with a discussion of primer dimers and M13 primer tails. Genome. 2019; 62(2):77–83. http://doi.org/10.1139/gen-2018-0145
Melo BF. Cyphocharax boiadeiro, a new species from the Upper rio Araguaia, Amazon basin, Brazil (Characiformes: Curimatidae). Zootaxa. 2017; 4247(2):114–20. http://doi.org/10.11646/zootaxa.4247.2.2
Melo BF, Vari RP. New species of Cyphocharax (Characiformes: Curimatidae) from the Upper rio Negro, Amazon basin. Neotrop Ichthyol. 2014; 12(2):327–32. http://doi.org/10.1590/1982-0224-20130153
Melo BF, Sidlauskas BL, Hoekzema K, Vari RP, Dillman CB, Oliveira C. Molecular phylogenetics of Neotropical detritivorous fishes of the family Curimatidae (Teleostei: Characiformes). Mol Phylogenet Evol. 2018; 127:800–12. https://doi.org/10.1016/j.ympev.2018.06.027
Pons J, Barraclough TG, Gomez-Zurita J, Cardoso A, Duran DP, Hazell S, Kamoun S, Sumlin WD, Vogler AP. Sequence-based species delimitation for the DNA taxonomy of undescribed insect. Syst Biol. 2006; 55(4):595–609. https://doi.org/10.1080/10635150600852011
Puillandre N, Brouillet S, Achaz G. ASAP: assemble species by automatic partitioning. Mol Ecol Resour. 2021; 21(2):609–20. https://doi.org/10.1111/1755-0998.13281
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA. Posterior summarisation in Bayesian phylogenetics using Tracer 1.7. Syst Biol. 2018; 67(5):901–04. https://doi.org/10.1093/sysbio/syy032
Sabaj MH. Codes for Natural History Collections in Ichthyology and Herpetology. Copeia. 2020; 108(3):593–669. https://doi.org/10.1643/ASIHCODONS2020
Stamatakis A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinform. 2014; 30(9):1312–13. https://doi.org/10.1093/bioinformatics/btu033
Taylor WR, Van Dyke GC. Revised procedures for staining and clearing small fishes and other vertebrates for bone and cartilage study. Cybium. 1985; 9(2):107–19.
Vari RP. Asystematics of the Neotropical characiform genus Cyphocharax Fowler (Pisces: Ostariophysi). Smithson Contrib Zool. 1989; 471:1–71. http://doi.org/10.5479/si.00810282.471
Vari RP. Systematics of the Neotropical characiform genus Cyphocharax Fowler (Pisces, Ostariophysi). Smithson Contrib Zool. 1992; 529:1–137. http://doi.org/10.5479/si.00810282.529
Vari RP, Sidlauskas BL, Le Bail PY. New species of Cyphocharax (Ostariophysi: Characiformes: Curimatidae) from Suriname and French Guiana and a discussion of curimatid diversity on the Guiana Shield. Cybium. 2012; 36(1):63–69. http://doi.org/10.26028/cybium/2012-361-007
Vari RP, Zanata AM, Camelier P. New species of Cyphocharax (Ostariophysi: Characiformes: Curimatidae) from the rio de Contas drainage, Bahia, Brazil. Copeia. 2010; 2010(3):382–87. http://doi.org/10.1643/CI-09-140
Wosiacki WB, Miranda DPS. Description of a new small species of the genus Cyphocharax (Characiformes: Curimatidae) from the lower Amazon basin. Copeia. 2013; 2013(4):627–33. http://doi.org/10.1643/CI-12-127
Zhang J, Kapli P, Pavlidis P, Stamatakis A. A general species delimitation method with applications to phylogenetic placements. Bioinform. 2013; 29(22):2869–76. http://doi.org/10.1093/bioinformatics/btt499
Authors
![]() Guilherme Moreira Dutra1
Guilherme Moreira Dutra1 ![]() ,
, ![]() George Vita1,
George Vita1, ![]() Péricles Vinícius Gentile1,
Péricles Vinícius Gentile1, ![]() Luz Eneida Ochoa2 and
Luz Eneida Ochoa2 and ![]() Andre Luiz Netto-Ferreira3
Andre Luiz Netto-Ferreira3
[1] Museu de Zoologia da Universidade de São Paulo, Av. Nazaré, 481, Ipiranga, 04263-000 São Paulo, SP, Brazil. (GMD) guilhermedutr@yahoo.com.br (corresponding author), (GV) george.vitao@gmail.com, (PVG) pericles.gentile@gmail.com.
[2] Universidad Nacional de Colombia, Sede de la Paz, Cesar, Colombia. leochoao@unal.edu.co.
[3] Laboratório de Ictiologia, Departamento de Zoologia, Instituto de Biociências, Universidade Federal do Rio Grande do Sul, Avenida Bento Gonçalves, 9500, 91501-970 Porto Alegre, RS, Brazil. alnferreira@gmail.com.
Authors’ Contribution 

Guilherme Moreira Dutra: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Project administration, Writing-original draft, Writing-review and editing.
George Vita: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Project administration, Writing-original draft, Writing-review and editing.
Péricles Vinícius Gentile: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Project administration, Writing-original draft, Writing-review and editing.
Luz Eneida Ochoa: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Writing-original draft, Writing-review and editing. Andre Luiz Netto-Ferreira: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Project administration, Writing-original draft, Writing-review and editing.
Ethical Statement
Not applicable.
Competing Interests
The authors declare no competing interests.
How to cite this article
Dutra GM, Vita G, Gentile PV, Ochoa LE, Netto-Ferreira AL. Integrative taxonomy reveals a new species of Cyphocharax (Characiformes: Curimatidae) from the Upper Paraíba do Sul River basin, Brazil. Neotrop Ichthyol. 2022; 20(3):e220017. https://doi.org/10.1590/1982-0224-2022-0017
Copyright
This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Distributed under
Creative Commons CC-BY 4.0

© 2022 The Authors.
Diversity and Distributions Published by SBI
![]() Accepted July 4, 2022 by Brian Sidlauskas
Accepted July 4, 2022 by Brian Sidlauskas
![]() Submitted June 9, 2021
Submitted June 9, 2021
![]() Epub Aug 8, 2022
Epub Aug 8, 2022