![]() Gabriela B. N. de Paula1 ,
Gabriela B. N. de Paula1 , ![]() Mariane Gavazzoni1,
Mariane Gavazzoni1, ![]() Cláudio H. Zawadzki1,2,
Cláudio H. Zawadzki1,2, ![]() Carlos A. Fernandes1,2,3,
Carlos A. Fernandes1,2,3, ![]() Ana L. B. Portela-Castro2,4,
Ana L. B. Portela-Castro2,4, ![]() Roberto L. Lui5 and
Roberto L. Lui5 and ![]() Vladimir P. Margarido1,5
Vladimir P. Margarido1,5 ![]()
PDF: EN XML: EN | Cite this article
Abstract
Hypostomus is the most specious genus of Hypostominae, composed of several species with high intraspecific morphological and color pattern variation, making their identification a complex issue. One of the species with problematic identification is Hypostomus tietensis that was described from a single specimen, resulting in uncertainties about its color pattern and correct identification. To assist in this context, cytogenetic analyzes were carried out in three putative populations of H. tietensis from the Upper Paraná River basin, one of them from the type locality. The three populations showed considerable cytogenetic differences, with 2n = 72 chromosomes for the population from the type locality and 2n = 76 chromosomes for the others. Terminal NORs were detected (Ag- and 18S rDNA-FISH), being simple for the type locality population (acrocentric pair23, long arm) and the Pirapó River (subtelocentric pair11, short arm), and multiple for Do Campo River (subtelocentric pairs11 and 12, short and long arm,respectively). C-banding was efficient in differentiating the type locality population from the others. Cytogenetic data revealed that populations from Pirapó and Do Campo rivers, although treated until now as Hypostomus aff. tietensis, represent a cryptic species, and those morphological analyses are necessary to differentiate and for describing this new species.
Keywords: Catfish, Chromosomal variation, Cytogenetics, Neotropical fishes, Upper Paraná River basin.
Hypostomus é o gênero mais especioso de Hypostominae, composto por várias espéciescom uma alta variação tanto morfológica, como no padrão de coloração intraespecífica, tornando sua identificação uma questão complexa. Uma das espécies com identificação complexa é Hypostomus tietensis, a qual foi descrita a partir de um único espécime, resultando em incertezas sobre o seu padrão de cor e identificação. Para auxiliar nesse contexto, análises citogenéticas foram realizadas em três populações putativas de H. tietensis da bacia do Alto rio Paraná, sendo uma delas da localidade tipo. As três populações apresentaram diferenças citogenéticas consideráveis, com 2n = 72 cromossomos para a população da localidade tipo e as demais com 2n = 76. RONs terminais foram detectadas (Ag- e FISH-DNAr 18S), sendo simples para a população da localidade tipo (par acrocêntrico 23, braço longo) e do rio Pirapó (par subtelocêntrico 11, braço curto) e múltiplas para rio Do Campo (pares subtelocêntricos 11 e 12, braço curto e longo, respectivamente), confirmado pela FISH-DNAr 18S. O bandamento C foi eficiente em diferenciar a população da localidade tipo das demais. Os dados citogenéticos revelaram que as populações dos rios Pirapó e do rio Do Campo, embora tratadas até agora como Hypostomus aff. tietensis, representam uma espécie críptica, e que análises morfológicas são necessárias para diferenciar e descrever esta nova espécie.
Palavras-chave: Cascudos, Variação cromossômica, Citogenética, Peixes Neotropicais, Bacia do Alto Rio Paraná.
Introduction
Loricariidae is the largest family in number of species within the order Siluriformes, and one of the largest in relation to all fish families in the world, with 163 genera and more than 1,020 species (Fricke et al., 2022). The phylogeny of the family is not well established and, despite its monophyletism, many conflicts still permeate the taxonomy and phylogeny of its subfamilies, which have been constantly revised (Armbruster, 2004; Reis et al., 2006; Chiachio et al., 2008; Cramer et al., 2011; Lujan et al., 2015; Pereira, Reis, 2017; Roxo et al., 2019).
Hypostomus Lacepède, 1803, belongs to the Hypostominae subfamily and is one of the most diverse and complex groups of fish, whose distribution extends from Central America to southern South America. The more than 150 species that make up the genus have various shades, which vary from light to dark tones, and may also have spots of different shapes spread over the body (Azevedo et al., 2021). The great intraspecific variability, in relation to color and morphology patterns, has made the taxonomic identification of species extremely difficult, contributing to several uncertainties within the genus (Weber, 2003; Armbruster, 2004).
One of the species of the genus with problematic identification is Hypostomus tietensis (Ihering, 1905) (Fig. 1), which was described by Ihering (1905) from a single specimen from the Upper Tietê River. However, there is a problem regarding the definition of its color pattern, since Ihering (1905) defined H. tietensis as having the upper part of the head with dark stripes or vermiculations, and the body with dark spots, and in a study carried out with species from the Upper Paraná River, Jerep et al. (2007) divided the main bulk of Hypostomus species from the Upper Paraná River into two groups based on color pattern, and included H. tietensis in the group of species with light rounded spots. Jerep et al. (2007) analyzed the holotype and several specimens from the Upper Tietê River basin, which showed irregularly spaced light stripes on a dark background. They further suggest that Ihering (1905) may have made a mistake, as defining the color pattern is complex and difficult task to do with just one specimen. As a result, several populations have been designated by taxonomists as Hypostomus aff. tietensis, or Hypostomus cf. tietensis in other regions of the Upper Paraná, such as in the flood area and basins of the Paranapanema and Piquiri rivers (Reis et al., 2020), indicating the existence of variations in relation to the described species from the type locality.
FIGURE 1| Holotype of Hypostomus tietensis, 160 mm SL, Tiête River, São Paulo, Brazil (https://data.nhm.ac.uk/object/a9a46e32-84d2-4996-9e69-71e8a2a8aca2/1633478400000, catalog number NHMUK 1905.6.9.1).
In view of the difficulties in identifying several species of Hypostomus and its unclear phylogeny, cytogenetics has proved itself as an important tool in understanding the systematics of the genus (Artoni, Bertollo, 2001; Kavalco et al., 2005; Alves et al., 2006; Bueno et al., 2012, among). Therefore, the objective of the present study was to cytogenetically analyze allopatric populations of H. tietensis, including that from the type locality and from two other localities in the Upper Paraná River basin where the occurrence of H. tietensis were previously recorded (Coleção Ictiológica do Núcleo de Pesquisas em Limnologia, Ictiologia e Aquicultura – NUPELIA), in order to test the hypothesis that there is cryptic diversity and to collaborate in the identification and characterization of the species.
Material and methods
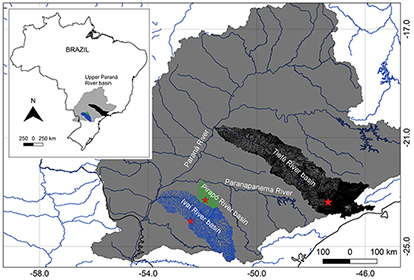
Three populations samples of Hypostomus tietensis were collected in the Upper Paraná River basin: 1- Piraí River a putative “type locality” (NUP 21903, a tributary of the upper stretches of the Tietê River basin, 23º22’22”S 47º22’13”W) with eight males and three females, 2- Pirapó River (NUP 2830, a tributary of Paranapanema River basin, 23º18’15”S 51º53’40”W) with 10 males and seven females, 3- Do Campo River (NUP 3025, a tributary of Ivaí River basin 24º04’41”S 52º26’12”W) with 13 males and five females (Fig. 2). The specimens were deposited in the Coleção Ictiológica do Núcleo de Pesquisas em Limnologia, Ictiologia e Aquicultura (NUP), Universidade Estadual de Maringá, Maringá.
All specimens were euthanized by an overdose of clove oil (Griffiths, 2000), and chromosome preparations were obtained from cells of anterior region of kidney (Bertollo et al., 2015). Metaphase chromosomes were obtained by the air-drying method and classified and organized into metacentric, submetacentric, subtelocentric, and acrocentric (Levan et al., 1964). The fundamental number (FN) was established considering the metacentric, submetacentric and subtelocentric chromosomes with two arms and the acrocentric chromosome with only one arm. Nucleolar organizing regions (AgNORs) were evidenced by silver nitrate impregnation (Howell, Black, 1980).Heterochromatin were determined following the C-banding techniqueand stained with propidium iodide (Sumner, 1972; Lui et al., 2012). The physical mapping of the 18S sequences was carried out by Fluorescence in situ Hybridization (FISH) according to Pinkel et al. (1986), using DNA probes obtained from Prochilodus argenteus Spix & Agassiz, 1829(Hatanaka, Galetti, 2004). Probes were labeled bynick translation method with biotin-16-dUTP (Roche®). Detection of signals was performed with avidin-FITC with biotinylated anti-avidin (Sigma-Aldrich), with the chromosomes counterstained with 4’,6-diamidino-2-phenylindole (DAPI, 50 μg/mL). Metaphases were photographed using a BX 51 epifluorescence microscope, coupled with Olympus DP 71 digital camera (Olympus America, Inc.) with the Olympus DP Controller software 3.2.1.276.
FIGURE 2| Map of Brazil showing the Upper Paraná River basin. Red stars represent Hypostomus tietensis populations, from Tietê River basin “type locality” (in black), from Pirapó River (in green), and from Do Campo River, tributary of Ivaí River basin (in blue).
Results
Hypostomus tietensis – Piraí River “type locality”
The diploid number was 72 chromosomes, with a karyotype formula of 8m+8sm+20st+36a and a fundamental number (FN) equal to 108 for the male and the female (Fig. 3A). A single pair of NORs was revealed with silver nitrate staining, located in the terminal regions of the long arms of the acrocentric pair 23 (Fig. 3A, highlighted above) confirmed by 18S rDNA-FISH (Fig. 3A, highlighted below). The heterochromatin detected by the C-banding showed large conspicuous terminals blocks in the long arm of chromosomes a (19, 20, 21 and 22) and pairs a 24, 32, 33, 34 (Fig. 3B).
Hypostomus aff. tietensis – Pirapó River
The diploid number found was 76 chromosomes, with a karyotype formula of 8m+6sm+24st+38a and a fundamental number (FN) equal to 114 for males and females (Fig. 3C). A single pair of NORs was observed, located in terminal regions of the short arms of subtelocentric pair 11 (Fig. 3C, highlighted above) confirmed by 18S rDNA-FISH (Fig. 3C, highlighted below). Heterochromatin were coincident with NORs and more evident in the subtelocentric pairs 13 (Fig. 3D).
Hypostomus aff. tietensis – Do Campo River
The diploid number found was 76 chromosomes, with a karyotype formula of 8m+6sm+24st+38a and a fundamental number (FN) equal to 114 for males and females (Fig. 3E). Multiple NORs were found in terminal regions of the short arms of subtelocentric pair 11 and long arm of only one subtelocentric chromosome of pair 12 (Fig. 3E, highlighted above) confirmed by 18S rDNA-FISH (Fig. 3E, highlighted below). Heterochromatin were coincident with NORs and more evident in the subtelocentric pair 13 (Fig. 3F).
FIGURE 3| Karyotypes of Hypostomus tietensis (A, B) Piraí River “type locality”, (C, D) Pirapó River, and Do Campo River (E, F) with conventional Giemsa-staining (A, C, E), AgNORs (highlighted above) and 18S rDNA-FISH (highlighted below); C-banding (B, D, F). m = metacentrics; sm = submetacentrics; st = subtelocentrics; a = acrocentrics. Scale bars = 10 µm.
Discussion
This is the first paper about cytogenetic characterization of Hypostomus tietensis, once available cytogenetic data for H. tietensis included data from abstracts presented on the Brazilian Symposiums(for review, Bueno et al., 2012). Even so, recently, considering these cytogenetic data, a study carried out on Hypostomus species from southeastern Brazil, using molecular and associated to cytonegenetics data, subdivided the genus into subgroups according to their diploid number, including H. tietensis within the subgroup of species that they have 2n = 66–68 chromosomes (Rocha-Reis et al., 2021).
Based on the results obtained in this study, the populations from Pirapo and Do Campo rivers showed considerable cytogenetic differences in relation to the population of the type locality (such as diploid number, karyotypic formula, AgNORs and heterochromatin distribution pattern) (Tab. 1), which can significantly contribute to the diagnosis of the species. The population of H. tietensis from the type locality had 2n = 72 chromosomes with FN = 108, unlike the other two H. aff. tietensis populations that present 2n = 76 chromosomes with FN = 114. Karyotypic differences were also observed in other populations of H. aff. tietensis from the Paranapanema River and Araquá River, where both had 2n = 68 chromosomes, with different karyotype formulas, 14m + 12sm + 42st/a (Oliveira, Foresti, 1993) and 18m + 10sm + 12st + 28a for H. aff. tietensis (Ishida et al., 2002). In a cytogenetic study carried out in seven species of the tribe Hypostominae, Alves et al. (2006) obtained the same result of 2n = 68, with a karyotype formula 18m+ 10sm+ 12st+ 28a for Hypostomus ancistroides (Ihering, 1911) from the Araquá River, which may be the same species, with possible errors in the identification of H. aff. tietensis, once they belong to same morphological group and may be misidentified. It is worthy to note that the range of 2n = 64 to 2n = 68 chromosomes is the diploid values to species pertaining to the super-group Hypostomus plecostomus (Linnaeus, 1758) sensu Queiroz et al. (2020): H. ancistroides, H. boulengeri (Eigenmann & Kennedy, 1903), H. commersoni Valenciennes, 1836, among others. In the Upper Paraná River basin, species lacking keels along lateral series of flanks, presenting no evident crests on compound-pterotic and having abdominal region somewhat naked, usually belongs to the super-group Hypostomus auroguttatus Kner, 1854. Species of the H. auroguttatus super-group present 2n = 72, 2n = 74 or 2n = 76 chromosomes (Bueno et al., 2012). Therefore, although H. tientensis were not in the bulk of species from Queiroz et al. (2020) it is very probable the species to belong to this super-group. In addition, up to now, the only species to the super-group H. plecostomus naturally inhabiting upper stretches of the upper Paraná River basin is the H. ancistroides complex. Then, it is very probable that the previous H. tietenis identifications by cytogenetic works in fact refers to some population of the H. ancistroides species complex.
The karyotypic numbers found are within the expected range for Hypostomus species from the southern basins of South America proposed by Bueno et al. (2013), where the high diploid number with several acrocentric chromosomes is considered a derived characteristic within the group, since 2n = 54 chromosomes is the putative plesiomorphic condition for Loricariidae, being the centric fissions responsible for playing an important role in the karyotypic evolution of the genus (Artoni, Bertollo, 2001). In Hypostomus, chromosome numbers range from 2n = 64 in H. faveolus Zawadzki, Birindelli & Lima, 2008, H. cochliodon Kner, 1854, and H. soniae Hollanda Carvalho & Weber, 2005to 2n = 84 in H. perdido Zawadzki, Tencatt & Froehlich, 2014 (Artoni, 1996; Cereali et al., 2008; Bueno et al., 2013, 2014; Becker et al., 2014; Oliveira et al., 2019), with 2n = 72 and 2n = 76 chromosomes being one of the most frequent conditions, observed in more than 30% of the species analyzed cytogenetically (Bueno et al., 2012). Although Mezzomo et al. (2020) consider the chromosome number range from 52 to 84, the 2n = 52 belongs to H. chrysostiktos, which the current status is Pterygoplichthys chrysostiktos (Birindelli, Zanata & Lima, 2007) (Fricke et al., 2022).
AgNORs and 18S rDNA-FISH were also efficient in differentiation of the type locality population (terminal, long arm, acrocentric pair 23) from the other two populations (terminal, short arm, subtelocentric pair 11). Although in the Do Campo River population some individuals presented an extra chromosome bearing NOR (long arm of chromosome st 12), variation in the number of NORs is common in Hypostomus, both among populations of the same species and within populations (Bueno et al., 2012). The presence of multiple NORs located in terminal regions is frequent in Hypostomus species (Artoni, Bertollo, 1996; Alves et al., 2006; Rubert et al., 2008; Martinez et al., 2011), being derived in the Hypostominae group, where the presence of a single site is considered basal in Loricariidae and in fish in general (Artoni, Bertollo, 1996, 2001; Alves et al, 2005). The most frequent locations of AgNORs are on the st/a chromosomes, corresponding to regions of heterochromatin in many populations, as has been reported in other species (Kavalco et al., 2005; Rubert et al., 2008; Traldi et al., 2013). This association may contribute to the evolution of the group, as it allows the dispersion of NORs sites throughout the genome through transposable elements, which may carry complete or partial NORs (Moreira-Filho et al., 1984; Vicari et al., 2008; Schmid et al., 2017).
TABLE 1 | Cytogenetic data in Hypostomus tietensis. 2n = diploid number, KF = karyotypic formula, FN = fundamental number, m = metacentrics, sm = submetacentrics, st = subtelocentrics, a = acrocentrics.
Populations | 2n | KF | FN | AgNORs | 18S rDNA-FISH |
Piraí River | 72 | 8m+8sm+20st+36a | 108 | Simple | Simple |
Pirapó River | 76 | 8m+6sm+24st+38a | 114 | Simple | Simple |
Do Campo River | 76 | 8m+6sm+24st+38a | 114 | Multiple | Multiple |
The heterochromatin analysis was also efficient to differentiate the chromosomal structure among the three populations. Individuals form the type locality population showed large conspicuous blocks on acrocentric chromosomes pairs19, 20, 21 and 22. In the populations of the Pirapó River and Do Campo River, heterochromatins were more evident in interstitial position on chromosome st par 13, and terminal blocks coincident with NORs in each population. In H. aff. tietensis studied by Ishida et al. (2002), the distribution was different, with the presence of heterochromatin in centromeric and pericentromeric positions, which suggests that this population should not represent H. tietensis.
Genetic variation does not immediately generate a phenotypic differentiation in the external morphology of an organism, revealing the presence of cryptic species (Bickford et al., 2007;Endo et al., 2012). The morphological similarity and diversity of karyomorphs in allopatric populations of Hypostomus represent cryptic species and species complexes (Lorscheider et al., 2015). Hypostomus paulinus (Ihering, 1905), for example, has populations with 2n = 72 to 2n = 76 chromosomes, being considered a species complex, as well as H. iheringii (Regan, 1908) (2n = 76 and 80), H. topavae (Godoy, 1969), H. nigromaculatus (Schubart, 1964), H. ancistroides, and H. regani (Ihering, 1905) that have karyomorphs with distinct karyotype formulas (Endo et al., 2012; Lorscheider et al., 2015; Rubert et al., 2016).
There are no strong morphological differences among the three populations analyzed in this work. The three populations present pale marks (blotches, spots or vermiculations) on a darker background, short to moderate jaw rami, flattened body, and comparatively to congeners, small eyes. However, the cytogenetic characters allowed to evidence a notable difference among the population of the type locality and the other populations, such as diploid number, karyotypic formula, NORs and heterochromatin distribution pattern, showing that the populations of the Pirapó River and Do Campo River evolved in a way independent in each basin, in relation to the population of the type locality, the valid H. tietensis. The data suggest the possible presence of cryptic species within this taxon, considering the populations of the Pirapó River and Do Campo River as Hypostomus aff. tietensis. Furthermore, given the range of karyomorphs already found for the species (2n = 68 to 76), H. tietensis can eventually be considered a species complex.
Acknowledgments
This study was supported by CAPES (Coordenadoria de Aperfeiçoamento de Ensino Superior), the Araucária Foundation (Fundação Araucária de Apoio e Desenvolvimento Científico e Tecnológico do Estado do Paraná) and CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico). The authors are grateful to the Instituto Chico Mendes de Conservação da Biodiversidade (MMA/ICMBio) for authorizing the capture of the fish specimens (license number: SISBIO 69480–2). We also thank the Western Paraná State University (UNIOESTE), NUPELIA, and the students Iago Penido (UEM), Tiago Pessali (PUC-MG), Hélen Proença (UEM), Gabriela Nardi (UEM) for helping with field work.
References
Alves AL, Oliveira C, Foresti F. Comparative cytogenetic analysis of eleven subfamilies Neoplecostominae and Hypostominae (Siluriformes: Loricariidae). Genetica. 2005; 124:127−36. https://doi.org/10.1007/s10709-004-7561-4
Alves AL, Oliveira C, Nirchio M, Granado A, Foresti F. Karyotypic relationships among the tribes of Hypostominae (Siluriformes: Loricariidae) with description of XO sex chromosome system in a Neotropical fish species. Genetica. 2006; 128(1):1–09. https://doi.org/10.1007/s10709-005-0715-1
Armbruster JW. Phylogenetic relationships of the suckermouth armored catfishes (Loricariidae) with particular emphasis on the Hypostominae and the Ancistrinae. Zool J Linn Soc. 2004; 141(1):1−80. https://doi.org/10.1111/j.1096-3642.2004.00109.x
Artoni RF, Bertollo LAC. Cytogenetic studies on Hypostominae (Pisces, Siluriformes, Loricariidae). Considerations on karyotype evolution in the genus Hypostomus. Caryologia. 1996; 49(1):81−90. https://doi.org/10.1080/00087114.1996.10797353
Artoni RF, Bertollo LAC. Trends in the karyotype evolution of Loricariidae fish (Siluriformes). Hereditas. 2001; 134(3):201−10. https://doi.org/10.1111/j.1601-5223.2001.00201.x
Azevedo FM, Zawadzki CH, Soria TV, Fabrin TMC, Oliveira AV, Prioli SMAP, Prioli AJ. Integrative taxonomy reveals the historically poorly defined armoured catfish Hypostomus variipictus (Ihering 1911), from the upper rio Paraná basin, Brazil (Siluriformes, Loricariidae). J Fish Biol. 2021; 99(1):143−52. https://doi.org/10.1111/jfb.14706
Becker QMC, Castro RJ, Silva AM, Vizzotto PC. Cytogenetic characterization of two species of Hypostomus (Siluriformes, Loricariidae) from tributaries of the Vermelho River, upper Paraguay River basin. Biodiversidade. 2014; 13(1):1–10.
Bertollo LAC, Cioffi MB, Moreira-Filho O. Direct chromosome preparation from freshwater teleost fishes, fish cytogenetic techniques. In: Ozouf-Costaz C, Pisano E, Foresti F, Toledo LFA, editors. Fish Citogenetic Techniques: Ray-fin fishes and Chondrichthyans. 2015. Available from: https://doi.org/10.1201/b18534-4
Bickford D, Lohman DJ, Sodhi NS, Ng PKL, Meier R, Winker K, Ingram KK, Das I. Cryptic species as a window on diversity and conservation. Trends Ecol Evol. 2007; 22(3):148−55. https://doi.org/10.1016/j.tree.2006.11.004
Bueno V, Zawadzki CH, Margarido VP. Trends in chromosome evolution in the genus Hypostomus Lacepede, 1803 (Osteichthyes, Loricariidae): a new perspective about the correlation between diploid number and chromosomes types. Rev Fish Biol Fish. 2012; 22:241−50. https://doi.org/10.1007/s11160-011-9215-9
Bueno V, Venere PC, Zawadzki CH, Margarido VP. Karyotypic diversification in Hypostomus Lacépède, 1803 (Siluriformes, Loricariidae): biogeographical and phylogenetic perspectives. Rev Fish Biol Fish. 2013; 23:103−12. https://doi.org/10.1007/s11160-012-9280-8
Bueno V, Venere PC, Konerat JT, Zawadzki CH, Vicari MR, Margarido VP. Physical Mapping of the 5S and 18S rDNA in ten species of Hypostomus Lacépède 1803 (Siluriformes: Loricariidae): Evolutionary tendencies in the genus. Sci World J. 2014; 2014:943825. http://dx.doi.org/10.1155/2014/943825
Cereali SS, Pomini E, Rosa R, Zawadzki CH, Froehlich O, Giuliano-Caetano L. Karyotype description of two species of Hypostomus (Siluriformes, Loricariidae) of the Planalto da Bodoquena, Brazil. Genet Mol Res. 2008; 7(3):583–91. http://dx.doi.org/10.4238/vol7-3gmr404
Chiachio MC, Oliveira C, Montoya-Burgos JI. Molecular systematic and historical biogeography of the armored Neotropical catfishes Hypoptomatinae and Neoplecostominae (Siluriformes: Loricariidae). Mol Phylogenet Evol. 2008;49(2):606−17. https://doi.org/10.1016/j.ympev.2008.08.013
Cramer CA, Bonatto SL, Reis RE. Molecular phylogeny of the Neoplecostominae and Hypoptopomatinae (Siluriformes: Loricariidae) using multiple genes. Mol Phylogenet Evol.2011; 59(1):43−52. https://doi.org/10.1016/j.ympev.2011.01.002
Endo KS, Martinez ERM, Zawadzki CH, Paiva LRS, Julio Júnior HF. Karyotype description of possible new species of the Hypostomus ancistroides complex (Teleostei: Loricariidae) and other Hypostominae. Acta Sci Biol Sci. 2012; 34(2):181–89. https://doi.org/10.4025/actascibiolsci.v34i2.9318
Fricke R, Eschmeyer WN, Fong JD. Eschmeyerʼs catalog of fishes: genera/species by family/subfamily [internet]. San Francisco: California Academy of Sience; 2022. Available from: https://researcharchive.calacademy.org/research/ichthyology/catalog/fishcatmain.asp
Griffiths SP. The use of clove oil as an anaesthetic and method for sampling intertidal rockpool fishes. J Fish Bio. 2000; 57(6):1453–64. https://doi.org/10.1111/j.1095-8649.2000.tb02224.x
Hatanaka T, Galetti PM. Mapping 18S and 5S ribosomal RNA genes in the fish Prochilodus argenteus Agassiz, 1929 (Characiformes, Prochilodontidae). Genetica. 2004; 122:239–44. https://doi.org/10.1007/s10709-004-2039-y
Howell WM, Black DA. Controlled silver-staining of nucleolus organizer regions with a protective colloidal developer: a 1 -step method. Experientia. 1980; 36:1014–15. https://doi.org/10.1007/BF01953855
Ihering RV. Description of four new loricariid fishes of the genus Plecostomus from Brazil. Ann Mag Nat Hist. 1905; 15(90):558–61. https://doi.org/10.1080/03745480509442852
Jerep FC, Shibatta OA, Zawadzki CH. A new species of Hypostomus Lacépède, 1803 (Siluriformes: Loricariidae) from the upper rio Paraná basin, Southern Brazil. Neotropical Ichthyology. 2007; 5(4):435–42. https://doi.org/10.1590/S1679-62252007000400002
Kavalco KF, Pazza R, Bertollo LAC, Moreira-Filho O. Heterochromatin characterization of four fish species of the family Loricariidae (Siluriformes). Hereditas. 2005; 141(3):237–42. https://doi.org/10.1111/j.1601-5223.2004.01850.x
Levan A, Fredga K, Sandberg AA. Nomenclature for centromeric position on chromosomes. Hereditas. 1964; 52(2):201–20. https://doi.org/10.1111/j.1601-5223.1964.tb01953.x
Lorscheider CA, Zawadzki CH, Vicari MR, Martins-Santos IC, Artoni RF. Karyotypic diversity of the armored catfish genus Hypostomus (Siluriformes: Loricariidae) in the context of its occurrence and distribution. J Fish Biol. 2015; 87(4):1099–110. https://doi.org/10.1111/jfb.12762
Lui RL, Blanco DR, Moreira-Filho O, Margarido VP. Propidium iodide for making heterochromatin more evident in the C-banding technique. Biotech Histochem. 2012; 87(7):433–38. https://doi.org/10.3109/10520295.2012.696700
Lujan NK, Armbruster JW, Lovejoy NR, López-Fernández H. Multilocus molecular phylogeny of the suckermouth armored catfishes (Siluriformes: Loricariidae) with a focus on subfamily Hypostominae. Mol Phylogenet Evol. 2015; 82:269–88. https://doi.org/10.1016/j.ympev.2014.08.020
Martinez ERM, Zawadzki CH, Foresti F, Oliveira C. Cytogenetic analysis of five Hypostomus species (Siluriformes, Loricariidae). Genet Mol Biol. 2011; 34(4):562–68. https://doi.org/10.1590/S1415-47572011005000038
Mezzomo P, Mielniczki‑Pereira AA, Sausen TL, Marinho JR, Cansian RL. Molecular inferences about the genus Hypostomus Lacépède, 1803 (Siluriformes: Loricariidae): a review. Mol Biol Rep. 2020; 47:6179–92. https://doi.org/10.1007/s11033-020-05542-z
Moreira-Filho O, Bertollo LAC, Galetti Jr., PM. Structure and variability of nucleolar organizer regions in Parodontidae fish. Can J Genet Cytol. 1984; 26(5):564–68. https://doi.org/10.1139/g84-089
Oliveira LC, Ribeiro MO, Costa GM, Zawadzki CH, Prizon-Nakajima AC, Borin-Carvalho LA, Martins-Santos I, Portela-Castro ALB. Cytogenetic characterization of Hypostomus soniae Hollanda-Carvalho & Weber, 2004 from the Teles Pires River, southern Amazon basin: evidence of an early stage of an XX/XY sex chromosome system. Comp Cytogenet. 2019; 13(4):411–22. https://doi.org/10.3897/CompCytogen.v13i4.36205
Pereira EHL, Reis RE. Morphology-based phylogeny of the suckermouth armored catfishes, with emphasis on the Neoplecostominae (Teleostei: Siluriformes: Loricariidae). Zootaxa. 2017; 4264(1):1–104. https://doi.org/10.11646/zootaxa.4264.1.1
Pinkel D, Straume T, Gray JW. Cytogenetic analysis using quantitative, high sensitivity, fluorescence hybridization. Proc Natl Acad Sci USA. 1986; 83(9):2934–38. https://doi.org/10.1073/pnas.83.9.2934
Queiroz LJ, Cardoso Y, Jacot-des-Combes C, Bahechar IA, Lucena CA, Rapp Py-Daniel L, Soares LMS, Nylinder S, Oliveira C, Parente TE, Torrente-Vilara G, Covain R, Buckup P, Montoya-Burgos JI. Evolutionary units delimitation and continental multilocus phylogeny of the hyperdiverse catfish genus Hypostomus. Mol Phylogenet Evol. 2020; 145:e106711. https://doi.org/10.1016/j.ympev.2019.106711
Reis RE, Pereira EHL, Armbruster JW. Delturinae, a new loricariid catfish subfamily (Teleostei, Siluriformes), with revisions of Delturus and Hemipsilichthys. Zool J Linn Soc. 2006; 147(2):277–99. https://doi.org/10.1111/j.1096-3642.2006.00229.x
Reis RB, Frota A, Deprá, GC, Ota RR. Freshwater fishes from Paraná State, Brazil: an annotated list, with comments on biogeographic patterns, threats, and future perspectives. Zootaxa. 2020; 4868(4):451–94. https://doi.org/10.11646/zootaxa.4868.4.1
Rocha-Reis DA, Pazza R, Kavalco KF. High congruence of karyotypic and molecular data on Hypostomus species from Brazilian southeast. Org Divers Evol. 2021; 21:135–43. https://doi.org/10.1007/s13127-021-00478-z
Roxo FF, Ochoa LE, Sabaj MH, Lujan NK, Covain R, Silva GSC, Melo BF, Albert JS, Chang J, Foresti F, Alfaro ME, Oliveira C. Phylogenomic reappraisal of the Neotropical catfish family Loricariidae (Teleostei: Siluriformes) using ultraconserved elements. Mol Phylogenet Evol. 2019; 135:148–65. https://doi.org/10.1016/j.ympev.2019.02.017
Rubert M, Zawadzki CH, Giuliano-Caetano L. Cytogenetic characterization of Hypostomus nigromaculatus (Siluriformes: Loricariidae). Neotrop Ichthyol. 2008; 6(1):93–100. https://doi.org/10.1590/S1679-62252008000100011
Rubert M, Rosa R, Zawadzki CH, Mariotto S, Moreira-Filho O, Giuliano-Caetano L. Chromosome mapping of 18S ribosomal RNA genes in eleven Hypostomus species (Siluriformes, Loricariidae): diversity analysis of the sites. Zebrafish. 2016; 13(4):360–68. https://doi.org/10.1089/zeb.2016.1279
Schmid M, Steinlein C, Feichtinger W, Nanda I. Chromosome banding in Amphibia. XXXV. Highly mobile Nucleolus Organizing Regions in Craugastor fitzingeri (Anura, Craugastoridae). Cytogenet Genome Res. 2017; 152(4):180–93. https://doi.org/10.1159/000481554
Sumner AT. A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res. 1972; 75(1):304–06. https://doi.org/10.1016/0014-4827(72)90558-7
Traldi JB, Blanco DR, Vicari MR, Martinez JF, Lui RL, Barros AV, Artoni RF, Moreira-Filho O. Chromosomal diversity in Hypostomus (Siluriformes, Loricariidae) with emphasis on physical mapping of 18S and 5S rDNA sites. Genet Mol Res. 2013; 12(1):463−71. http://dx.doi.org/10.4238/2013.February.8.11
Vicari MR, Noleto RB, Artoni RF, Moreira-Filho O, Bertollo LAC. Comparative cytogenetics among species of the Astyanax scabripinnis complex. Evolutionary and biogeographical inferences. Genet Mol Biol. 2008; 31(1 suppl):173–79. https://doi.org/10.1590/S1415-47572008000200002
Weber C. Subfamily Hypostominae (Armored catfishes). In: Reis RE, Kullander SO, Ferraris Jr., CJ., editors. Check list of the freshwater fishes of South and Central America. Porto Alegre, Edipucrs. 2003.
Authors
![]() Gabriela B. N. de Paula1 ,
Gabriela B. N. de Paula1 , ![]() Mariane Gavazzoni1,
Mariane Gavazzoni1, ![]() Cláudio H. Zawadzki1,2,
Cláudio H. Zawadzki1,2, ![]() Carlos A. Fernandes1,2,3,
Carlos A. Fernandes1,2,3, ![]() Ana L. B. Portela-Castro2,4,
Ana L. B. Portela-Castro2,4, ![]() Roberto L. Lui5 and
Roberto L. Lui5 and ![]() Vladimir P. Margarido1,5
Vladimir P. Margarido1,5 ![]()
[1] Universidade Estadual de Maringá, Pós-Graduação em Biologia Comparada, Av. Colombo, 5790, 87020-900 Maringá, PR, Brazil. (GBMP) gabipaula3@gmail.com, (MG) marianegavazzoni@gmail.com, (CHZ) chzawadzki@nupelia.uem.br, (CAF) cafernandes@uem.br, (VPM) vladimir.margarido@unioeste.br (corresponding author).
[2] Universidade Estadual de Maringá, Núcleo de Pesquisas em Limnologia, Ictiologia e Aquicultura, Av. Colombo, 5790, 87020-900 Maringá, PR, Brazil. (ALBPC) albpcastro@uem.br.
[3] Universidade Estadual de Maringá, Pós-Graduação em Biotecnologia Ambiental, Av. Colombo, 5790, 87020-900 Maringá, PR, Brazil.
[4] Universidade Estadual de Maringá, Departamento de Biotecnologia, Genética e Biologia Celular, Av. Colombo, 5790, 87020-900 Maringá, PR, Brazil.
[5] Universidade Estadual do Oeste do Paraná, Centro de Ciências Biológicas e da Saúde, R. Universitária, 1619, Jardim Universitário, 85819-170 Cascavel, PR, Brazil. roberto.lui@unioeste.br.
Authors’ Contribution 

Gabriela B. N. de Paula: Conceptualization, Formal analysis, Investigation, Methodology, Writing-original draft, Writing-review and editing.
Mariane Gavazzoni: Methodology, Supervision, Validation, Writing-original draft, Writing-review and editing.
Cláudio H. Zawadzki: Conceptualization, Formal analysis, Investigation, Methodology, Supervision, Writing-original draft, Writing-review and editing.
Carlos A. Fernandes: Conceptualization, Writing-original draft, Writing-review and editing.
Ana L. B. Portela-Castro: Conceptualization, Writing-original draft, Writing-review and editing.
Roberto L. Lui: Conceptualization, Writing-original draft, Writing-review and editing.
Vladimir P. Margarido: Conceptualization, Formal analysis, Funding acquisition, Investigation, Methodology, Supervision, Validation, Writing-original draft, Writing-review and editing.
Ethical Statement
This study was carried out in strict accordance with the recommendations of the Guide for the Care and Use of Laboratory Animals, approved by the Committee on the Ethics of Animal Experiments of the Universidade Estadual de Maringá (License number: Protocol 1968010719 – CEUA/UEM; License number: SISBIO 69480–2).
Competing Interests
The authors declare no competing interests.
How to cite this article
Paula GBN, Gavazzoni M, Zawadzki CH, Fernandes CA, Portela-Castro ALB, Lui RL, Margarido VP. Identification of cryptic species in allopatric populations of Hypostomus tietensis (Siluriformes: Loricariidae) through cytogenetics analyses. Neotrop Ichthyol. 2022; 20(2):e210158. https://doi.org/10.1590/1982-0224-2021-0158
Copyright
This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Distributed under
Creative Commons CC-BY 4.0

© 2022 The Authors.
Diversity and Distributions Published by SBI
![]() Accepted May 2, 2022 by Claudio Oliveira
Accepted May 2, 2022 by Claudio Oliveira
![]() Submitted November 10, 2021
Submitted November 10, 2021
![]() Epub June 03, 2022
Epub June 03, 2022