Tiago C. Faria[1] ![]()
![]() , Karen L. A. Guimarães[2]
, Karen L. A. Guimarães[2] ![]() , Luís R. R. Rodrigues[2]
, Luís R. R. Rodrigues[2]![]() ,Claudio Oliveira[3]
,Claudio Oliveira[3] ![]() andFlávio C.T. Lima[1]
andFlávio C.T. Lima[1] ![]()
PDF: EN XML: EN | Supplementary: S1, S2, S3 | Cite this article
Abstract
A new species of Hyphessobrycon belonging to the Hyphessobrycon heterorhabdus species-group from the lower rio Tapajós, state of Pará, Brazil, is described. The new species is allocated into the Hyphessobrycon heterorhabdus species-group due to its color pattern, composed by an anteriorly well-defined, horizontally elongated humeral blotch that becomes diffuse and blurred posteriorly, where it overlaps with a conspicuous midlateral dark stripe that becomes blurred towards the caudal peduncle and the presence, in living specimens, of a tricolored longitudinal pattern composed by a dorsal red or reddish longitudinal stripe, a middle iridescent, golden or silvery longitudinal stripe, and a more ventrally-lying longitudinal dark pattern composed by the humeral blotch and dark midlateral stripe. It can be distinguished from all other species of the group by possessing humeral blotch with a straight or slightly rounded ventral profile, lacking a ventral expansion present in all other species of the group. The new species is also distinguished from Hyphessobrycon heterorhabdus by a 9.6% genetic distance in the cytochrome c oxidase I gene. The little morphological distinction of the new species when compared with its most similar congener, H. heterorhabdus, indicates that the new species is one of the first truly cryptic fish species described from the Amazon basin.
Keywords: Biodiversity, Cryptic Species, DNA Barcoding, Rio Amazonas, Rio Tapajós.
Uma nova espécie de Hyphessobrycon pertencente ao grupo Hyphessobrycon heterorhabdus é descrita da região do baixo rio Tapajós, estado do Pará, Brasil. A nova espécie é incluída no grupo Hyphessobrycon heterorhabdus devido ao seu padrão de coloração, composto por uma mancha umeral alongada, anteriormente bem definida, que se torna difusa e borrada posteriormente, onde se sobrepõe a uma conspícua faixa escura médio-lateral que se torna borrada próxima ao pedúnculo caudal, e pela presença, em exemplares vivos, de um padrão longitudinal tricolor, composto por uma faixa longitudinal vermelha ou avermelhada dorsal, uma faixa média iridescente dourada ou prateada e, mais ventralmente, o padrão longitudinal escuro composto pela faixa escura médio-lateral e mancha umeral. A espécie pode ser distinguida das outras espécies pertencentes ao grupo por possuir uma mancha umeral com região ventral retilínea ou levemente arredondada, sem uma expansão ventral presente nas demais espécies do grupo. A espécie também se diferencia de Hyphessobrycon heterorhabdus por uma distância genética de 9,6% no gene citocromo c oxidase I. A sutil diferença morfológica da nova espécie quando comparada ao seu congênere mais similar, H. heterorhabdus, indica que a nova espécie é uma das primeiras espécies de peixes verdadeiramente crípticas descritas da Bacia Amazônica.
Palavras-chave: Biodiversidade, DNABarcoding, Espécie Críptica, Rio Amazonas, Rio Tapajós.
Introduction
The genus Hyphessobrycon Durbin, 1908, currently with 160 species (Ota et al., 2020; Faria et al., 2020; Terán et al., 2020), is the most species rich genus of Characidae, now that Astyanax Baird & Girard, 1854, the formerly more species rich genus, was split into several genera (Terán et al., 2020). Hyphessobrycon is diagnosed by a combination of non-apomorphic morphological characters, which includes lateral line incompletely perforated, and caudal fin lacking scales at its basis (Eigenmann, 1918). As predicted by Weitzman, Fink (1983), the genus has been repeatedly recovered as non-monophyletic, both by molecular and total-evidence phylogenies (Oliveira et al., 2011; Mirande, 2019; Ohara et al., 2019).
Regardless of the non-monophyly of the genus, some authors have proposed putative monophyletic species-groups within Hyphessobrycon using either color pattern or anal-fin hooks morphology characters (e.g., Weitzman, Palmer, 1997; Ingenito et al., 2013; Ota et al., 2020). Among those groups, the Hyphessobrycon heterorhabdus species-group was recently redefined, with all its component species [H. amapaensis Zarske & Géry, 1998, H. ericae Moreira & Lima, 2017, H. heterorhabdus (Ulrey, 1894), H. montagi Lima, Coutinho & Wosiacki, 2014, H. sateremawe Faria, Bastos, Zuanon & Lima, 2020, and H. wosiacki Moreira & Lima, 2017] possessing a well-defined, elongated humeral blotch, continuous with a midlateral, well-defined dark stripe that becomes blurred towards the caudal peduncle (Lima et al., 2014; Moreira, Lima, 2017) and a tricolored longitudinal pattern, composed dorsally by a red or reddish longitudinal stripe, a middle iridescent, golden or silvery longitudinal stripe, together with the longitudinal dark pattern composed by the humeral blotch and dark midlateral stripe (Faria et al., 2020).
Although the morphological characters remain as the main approach in ichthyological taxonomic literature, molecular data have established itself as an important tool to help understand taxonomic boundaries among species, as well as populational, conservation, and even some ecological aspects of the biology of fishes (e.g., Schaefer et al., 2011; Weigt et al., 2012; Pereira et al., 2013; Landi et al., 2014). Among the molecular tools used in taxonomy, the DNA barcoding surely stands out. This methodology is based on the analysis of the mitochondrial gene cytochrome c oxidase I, having revealed itself as an effective way to identify potential cryptic species in taxonomic fish studies (e.g., Tornabene et al., 2010; Baldwin, Weigt, 2012; Costa et al., 2012; Melo et al., 2016; Allen et al., 2016). In the present paper we describe a new cryptic species belonging to the Hyphessobrycon heterorhabdus species-group from the lower rio Tapajós, Amazon basin in Brazil. This species was previously identified as a putative undescribed species using the DNA barcoding, being distinguished from the nearest population of H. heterorhabdus by a distance of 9.15% (Guimarães et al., 2018). An ongoing revisionary morphological study of the H. heterorhabdus species-group being conducted by two of the authors (TCF and FCTL) revealed a slight but consistently distinct color pattern between this taxon and H. heterorhabdus.
Material and methods
Morphological analysis. Counts and measurements follow Fink, Weitzman (1974), except for the number of horizontal scale rows below lateral line, which are counted to the pelvic-fin insertion (excluding the axillary scale) rather than to the anal-fin origin, and the addition of three measurements: distance from pelvic-fin origin to anal-fin origin, dorsal-fin base length, and anal-fin base length. Standard length (SL) is expressed in millimeters (mm) and all other measurements are expressed as percentages of SL, except subunits of the head, which are expressed as percentages of head length (HL). In the description, counts are followed by their absolute frequency in parentheses. Asterisks indicate the counts of the holotype. Circulii and radii were counted on scales from the row immediately dorsal to the lateral line at the vertical through the dorsal-fin origin. Counts of supraneurals, branchiostegal rays, gill-rakers of the first branchial arch, teeth cusps, minute dentary teeth, unbranched anal-fin rays, procurrent caudal-fin rays, uroneurals, and position of pterygiophores were taken from cleared and stained (C&S) specimens prepared according to Taylor, Van Dyke (1985). Vertebrae of the Weberian apparatus were counted as four elements and the compound caudal centrum (PU1+U1) as a single element. Catalog numbers are followed by the total number of specimens and their SL range. The number of cleared and stained specimens, if any, is given after the SL range of the total number of specimen, followed by their respective SL range. Institutional abbreviations follow Sabaj (2019).
Molecular analysis. In order to investigate the genetic divergence among the new species and its putatively closest taxa, we analyzed DNA barcoding sequences from 10 specimens collected within the known range of the species. DNA extraction followed an adapted salting-out protocol (Aljanabi, Martinez, 1993; Vitorino et al., 2015). For amplification of Cytochrome c oxidase subunit I (COI) we used the standard primers FishF1 and Fish R1 (Ward et al., 2005), in PCR reactions assembled and processed as detailed in Guimarães et al. (2018). The DNA barcoding sequences were obtained by capillary sequencing with an ABI3500 genetic analyzer, using the ABI PRISM Big Dye Terminator kit (Applied Biosystems) following the manufacturer instructions.
The sequences were visually inspected and trimmed with BioEdit software (Hall, 1999). The dataset was complemented with DNA barcodes from H. ericae (n=10), H. heterorhabdus (n=17), H. agulha Fowler, 1913 (n=2) and Pristella maxillaris (Ulrey, 1894) (BOLD:AAC9546), that were downloaded from internet (www.boldsystems.org). The latter species was adopted as the outgroup. The sequences were aligned using the ClustalW Algorithm (Thompson et al., 1994) implemented in the software BioEdit (Hall, 1999). Detailed information associated to the specimens and sequences are given in S1 and S2. For initial data exploration, we constructed a pairwise distance matrix and a neighbor-joining (NJ) tree topology using the K2P distance model (Kimura, 1980), with the software MEGA X (Kumar et al., 2018). We used FigTree v.1.2.2 (http://tree.bio.ed.ac.uk/software/figtree) for tree design edition.
Molecular species delimitation. To identify putative new species by molecular data we applied three species delimitation approaches: 1) Generalized Mixed Yule Coalescent – GMYC (Pons et al., 2006; Fujisawa, Barraclough, 2013); 2) Barcode Index Number (BIN) (Ratnasingham, Hebert, 2013) and 3) Automatic Barcode Gap Discovery – ABGD (Puillandre et al., 2012). First, for the GMYC method we removed the repeated haplotypes and constructed an ultrametric tree based on GTR evolution model, Gamma shape distribution, molecular clock lognormal relaxed and Yule process speciation. The Bayesian inference with 100 million MCMC iterations and sampled at each 1000 iterations with 10% burn-in was processed in BEAST v1.8.0 (Drummond, Rambaut, 2007). The convergence and stability were checked with the software Tracer v.1.7.1 and retained for Effective Sample Sizes (ESS) > 200 (Drummond, Rambaut, 2007). The resulting trees were combined with TreeAnotator v1.8.0 and saved as a Newick file (Drummond, Rambaut, 2007). To test the branching events for speciation (Yule process) and coalescence null hypothesis we used the packages Splits (Species Limits by Threshold Statistics) and Ape (Analyses of Phylogenetics and Evolution), with the single threshold model implemented in R 3.4.0 statistical software (R Core Team, 2014).
The Barcode Index Number (BIN) is an online tool implemented in the BOLD System workbench (www.boldsystems.org) that uses the algorithm RESL (Refined Single Linkage Analysis) to find clusters of DNA barcodes from entry data and the BOLD archived library (Ratnasingham, Hebert, 2013). This procedure assigns a tag (BIN) for the clusters recovered and assumes it as an Operational Taxonomic Unit (OTU) and can reveal putative new species based on a maximum intra-cluster distance threshold (Ratnasingham, Hebert, 2013).
The barcoding gap is largely assumed as a genetic signature of species divergence. We used the method Automatic Barcode Gap Discovery (ABGD) to explore the existence of barcoding gap among the Hyphessobrycon taxa analyzed. This approach estimates the species boundary searching for discontinuities between distributions of pairwise genetic distances (Puillandre et al., 2012). We processed this analysis in the online platform (bioinfo.mnhn.fr/abi/public/abgd/abgdweb.html) with an input file (distance matrix) software MEGA X (Kumar et al., 2018) and setting for K80 model, intraspecific divergence (min. 0.001 and max. 0.1) and barcoding gap as default (X=1.5).
Results
Hyphessobrycon cantoi, new species
urn:lsid:zoobank.org:act:82D27CE6-8349-4F17-A58D-745B3B1B76D2
(Figs. 1–5; Tab. 1)
Hyphessobrycon gr. heterorhabdus. —Guimarães et al., 2018:7–8, 10 (Brazil, Pará, Santarém, streams União do Vegetal, Sonrisal, and Irurá, lower rio Tapajós basin; genetic distance from Hyphessobrycon heterorhabdus; as a putative undescribed species).
Holotype. ZUEC 17228, 30.2 mm SL, female, Brazil, Pará State, Santarém, stream tributary of Lago Verde, lower rio Tapajós basin, 02°31’19”S 54°54’58”W, J. D. Bogotá-Gregory, 17 May 2015.
FIGURE 1 | Hyphessobrycon cantoi. A. ZUEC 17228, holotype, 30.2 mm SL, female, Brazil, Pará, Santarém, Alter do Chão, stream tributary of Lago Verde, rio Tapajós basin. B. ZUEC 14597, paratype, 32.1 mm SL, female, Brazil, Pará, Santarém, Mararu, igarapé do Diamantino, rio Amazonas basin.
Paratypes. All from Brazil, Pará State, Santarém. Rio Tapajós basin: ZUEC 12437, 37, 20.8–30.3 mm SL, same locality and collector as holotype, 29 Jan 2015. ANSP 207974, 5, 24.3–29.5 mm SL; FMNH 144981, 5, 24.3–26.4 mm SL; INPA 59495, 5, 24.8–30.2 mm SL; UF 245654, 5, 23.7–27.3 mm SL; ZUEC 12440, 136, 20.9–33.3 mm SL, 3 C&S, 22.2–27.0 mm SL, same data as holotype. ZUEC 12439, 146, 13.1–30.6 mm SL, same locality and collector as holotype, 8 Aug 2015. LBP 30214, 10, 12.5–29.5 mm SL, União do Vegetal, rio Tapajós basin, 02°28’50.10”S 54°47’19.10”W, L. R. R. Rodrigues, 27 Sep 2020. MPEG 35994, 4, 15.9–27.4 mm SL, comunidade Cucurunã, igarapé Cucurunã, 02°28’54”S 54°46’13”W, M. Sudário, 12 May 2017. MPEG 35996, 14, 18.7–28.8 mm SL, same locality as previous, M. Sudário, 21 Apr 2017. UFOPA 703, 25, 10.4–28.6 mm SL, igarapé Irurá, 02°27’50”S 54°44’6”W, D. Franco & F. Ribeiro, 1 May 2012. UFOPA 723, 21, 13.5–31.9 mm SL, igarapé Irurá, 02°29’23”S 54°44’16”W, D. Franco & F. Ribeiro, 11 May 2012. UFOPA 776, 51, 12.3–25.1 mm SL, igarapé Irurá, 02°27’32”S 54°44’6”W, D. Franco & F. Ribeiro, 1 Sep 2012. UFOPA 885, 17, 20.3–32.4 mm SL, igarapé Sonrisal, 02°32’S 54°55”W, T. Torres, C. Silva & J. Souza, 13 Jan 2014. UFOPA 901, 89, 14.8–32.0 mm SL, igarapé Irurama, 02°29’3”S 54°50’12”W, T. Torres, C. Silva & J. Souza, 15 Jan 2014. UFOPA 932, 8, 24.7–32.7 mm SL, igarapé Santa Luzia, 02°32’37”S 54°52’8”W, T. Torres, C. Silva, J. Souza, A. Canto & F. Ribeiro, 18 Jan 2014. Rio Amazonas basin: ZUEC 14597, 46, 20.2–32.1 mm SL, Mararu, igarapé do Diamantino, 02°30’16”S 54°39’33”W, T. C. Faria & K. L. A. Guimarães, 9 Aug 2018.
Diagnosis. Hyphessobrycon cantoi can be distinguished from all congeners, except H. amapaensis, H. ericae, H. heterorhabdus, H. sateremawe and H. wosiackii, by the presence of an elongated, anteriorly well-defined humeral blotch that becomes progressively diffuse and blurred posteriorly, overlapping with a midlateral dark stripe. Hyphessobrycon cantoi can be distinguished from H. ericae and H. wosiackii by lacking a caudal peduncle blotch (vs. presence of a caudal peduncle blotch). Hyphessobrycon cantoi can be distinguished from H. amapaensis, H. heterorhabdus and H. sateremawe by lacking a ventral extension of the humeral blotch (vs. ventral extension of the humeral blotch present, althoughabsence of ventral extension may occurs in specimens of H. amapaensis). Hyphessobrycon cantoi can be further distinguished from H. amapaensis by presenting a conspicuous midlateral dark stripe (vs. inconspicuous midlateral dark stripe) and by possessing a relatively thin red longitudinal stripe (vs. midlateral red stripe very thick and conspicuous). Hyphessobrycon cantoi can be also further distinguished from H. sateremawe by presenting a humeral blotch narrower, occupying vertical height equivalent to less than one scale row to middle of body (vs. humeral blotch and continuous midlateral stripe broad, occupying vertical height equivalent of two scale rows to middle of body). Hyphessobrycon cantoi is also distinguished from H. heterorhabdus by >9% of genetic distance in the cytochrome c oxidase I (COI) gene. Hyphessobrycon cantoi can be distinguished from H. heterorhabdus by 53–79 mutations and from H. ericae by 87 mutations in the COI gene (S3).
FIGURE 2 | Hyphessobrycon cantoi. Living specimen, Brazil, Pará, Santarém, stream tributary of lago Maicá (not preserved).
Description. Morphometric data of holotype and paratypes in Tab. 1. Body compressed. Greatest body depth at vertical through dorsal-fin origin. Dorsal profile of head slightly convex from upper lip to vertical through posterior nostril, straight from that point to tip of supraoccipital spine. Dorsal profile of body slightly convex from latter point to dorsal-fin origin. Dorsal-fin base straight, posteroventrally slanted, slightly convex from end of dorsal fin base to adipose-fin insertion and slightly concave between adipose-fin insertion and origin of anteriormost dorsal procurrent caudal-fin ray. Ventral profile of head and body convex from anterior tip of lower jaw to anal-fin origin. Anal-fin base straight, posterodorsally slanted. Ventral profile of caudal peduncle slightly concave.
Jaws equal, mouth terminal. Posterior terminus of maxilla reaching vertical through anterior margin of iris. Maxilla approximately at 45 degrees angle relative to longitudinal axis of body. Nostrils close to each other, anterior opening oval, posterior opening crescent-shaped. Premaxillary teeth in two rows. Outer teeth row with 2(20), 3(20), or 4(2) conic teeth. Inner row with 5(41) bi- to tetracuspid teeth, symphyseal tooth narrower than remaining teeth. Maxilla with 2(1), 3(1), or 4(1) conical to tricuspid teeth. Dentary with 9(1), 10(1) or 11(1) teeth, anteriormost 3–4 teeth larger, tricuspid, 6 remaining teeth considerably smaller and conical. Central cusp of all teeth more developed than remaining lateral cusps.
Scales cycloid. Two to seven parallel radii strongly marked, circulii well marked anteriorly, on covered portion, weakly marked posteriorly. Lateral line slightly deflected downward and incompletely pored, with 7(4), 8(26), 9(16), 10*(4), 11(3) or 13(1) perforated scales. Longitudinal scales series including lateral-line scales 31(2), 32(6), 33*(22), 34(13), 35(7) or 36(2). Longitudinal scale rows between dorsal-fin origin and lateral line 5*(58) or 6(2). Longitudinal scale rows between lateral line and pelvic-fin origin 3*(49) or 4(1). Predorsal scales 9(13), 10*(37), 11(9) or 12(1). Circumpeduncular scales 11(1) or 12*(58). Caudal fin with few small scales basally.
Dorsal-fin rays ii,8(2), 9*(57) or 10(1). Dorsal-fin origin slightly anterior of middle of standard length. First dorsal-fin pterygiophore inserting behind neural spine of 9th(3) vertebrae. Adipose fin present. Anteriormost anal-fin pterygiophore inserting posterior to haemal spine of 14th(1) or 15th(2) vertebrae. Anal-fin rays iv,17*(13), 18(30), 19(15) or 20(2). Last unbranched and first to third anteriormost branched rays distinctly longer than remaining rays, subsequent rays gradually decreasing in size. Pectoral-fin rays i,9(6), 10*(38), 11(15) or 12(1). Pelvic-fin rays i,6(9) or 7*(51). Tip of pelvic fin reaching anteriormost anal-fin rays. Caudal fin forked, lobes roughly rounded and of similar size. Nine (1) or 11(2) dorsal procurrent caudal-fin rays, and 7(1) or 8(2) ventral procurrent caudal-fin rays. Vertebrae 32(1) or 33(2). Supraneurals 4(3), upper portion wider. Branchiostegal rays 4. First gill arch with 2(2) or 3(1) hypobranchial, 6(1) or 8(2) ceratobranchial, 1(3) on cartilage between ceratobranchial and epibranchial, and 6(3) epibranchial gill-rakers.
TABLE 1 | Morphometric data for Hyphessobrycon cantoi. N = number of specimens measured. S.D = Standard deviation. Holotype measurements included in the range.
| Holotype | Range | Mean ± S.D. | N |
Standard length (mm) | 30.2 | 24.5–33.3 | – | 60 |
Percents of standard length |
|
|
|
|
Depth at dorsal-fin origin | 33.1 | 28.7–34.3 | 31.9±1.4 | 60 |
Snout to dorsal-fin origin | 50.1 | 48.6–52.1 | 50.2±0.8 | 60 |
Snout to pelvic-fin origin | 48.7 | 46.1–50.7 | 48.2±1.0 | 60 |
Snout to anal-fin origin | 64.9 | 61.7–67.5 | 64.3±1.1 | 60 |
Caudal peduncle depth | 8.9 | 7.3–9.7 | 8.5±0.5 | 60 |
Caudal peduncle length | 13.2 | 12.2–15.9 | 13.9±0.8 | 60 |
Pectoral-fin length | 19.5 | 17.8–22.7 | 20.5 ±0.9 | 60 |
Pelvic-fin length | 16.9 | 14.8–18.8 | 16.9±0.8 | 60 |
Dorsal-fin base | 13.6 | 11.5–15.2 | 13.8±0.8 | 60 |
Dorsal-fin length | 29.5 | 25.2–32.8 | 29.1±1.4 | 60 |
Anal-fin base | 25.2 | 24.4–28.7 | 26.1 ±1.0 | 60 |
Head length | 27.5 | 25.4–28.6 | 27.1±0.6 | 60 |
Percents of head length |
|
|
|
|
Horizontal orbital diameter | 38.5 | 33.8–42.0 | 37.3±1.5 | 60 |
Snout length | 26.5 | 20.8–29.6 | 26.8±1.8 | 60 |
Least interorbital width | 28.9 | 26.4–34.6 | 31.2±1.7 | 60 |
Upper jaw length | 45.8 | 41.7–47.5 | 44.4±1.4 | 60 |
Color in alcohol. Overall body color beige (Fig. 1). Dorsal portion of head and body darker. Ventral portion of head and body with few scattered dark chromatophores. Snout and lower lip dark. Dark chromatophores scattered across infraorbitals and opercle, concentrated dorsally. Predorsal and preadipose scales with conspicuous central dark blotches. Three dorsalmost scale rows with conspicuous reticulated pattern formed by dark chromatophores concentrated at scales margins. Area immediately above humeral blotch and midlateral dark stripe clear, with evenly-scattered, light-grey chromatophores. Humeral blotch narrow and conspicuous. Anterior region well defined, horizontally elongated, with diffuse dorsal expansion and ventral margin approximately straight to slightly convex, becoming gradually diffuse posteriorly, thicker in some specimens. Posterior region of humeral blotch gradually becoming diffuse and coalescing with midlateral dark stripe. Midlateral dark stripe thicker anteriorly, narrower and more diffuse posteriorly, reaching caudal peduncle. Ventral half of abdominal region clear, with few, evenly-scattered chromatophores. Region above anal fin darker, with concentration of dark chromatophores increasing towards caudal peduncle. Dark chromatophores aligned along myocommata of hypaxial muscles above anal fin or above posterior half of anal fin. Caudal fin mostly hyaline, with high concentration of dark chromatophores on middle caudal rays. Anal fin mostly hyaline, with dark chromatophores scattered along interradial membranes, more concentrated along distal region of anal-fin lobe. Dark chromatophores line parallel to anal-fin base. Dorsal-fin rays mostly hyaline, with dark chromatophores concentrated on anteriormost rays. Adipose fin with few scattered dark chromatophores mainly concentrated on proximal region. Pectoral and pelvic fins with dark chromatophores scattered.
Color in life. Based on a picture provided by A. L. C. Canto (Fig. 2), and pictures taken by one of the authors (LRRR). Overall body color clear, with olivaceous hue. Lower half of head and abdominal region silvery. Dorsal portion of eye red. Tricolor longitudinal pattern starting immediately posterior to opercle and ending immediately before base of caudal fin, composed of narrow dorsal red stripe, narrow middle iridescent stripe and narrow ventral dark longitudinal pattern composed of humeral blotch followed posteriorly by midlateral stripe. Longitudinal red stripe anteriorly continuous, becoming row of red spots at midbody. Middle stripe golden, relatively narrow and well defined, thicker anterior to humeral blotch, becoming row of golden spots at midbody. Scattered, iridescent greenish chromatophores immediately ventral to humeral blotch and anterior region of midlateral stripe.
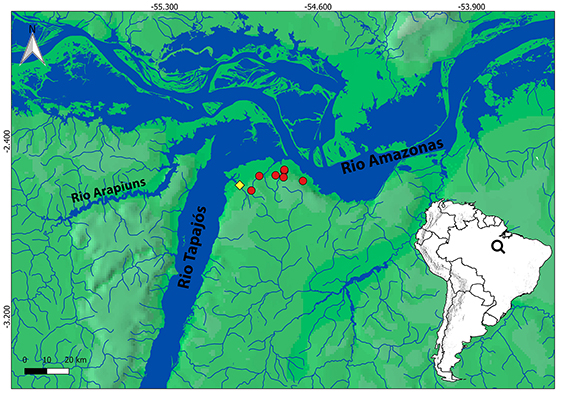
FIGURE 3 | Map of the lower rio Tapajós and its confluence with the rio Amazonas, showing the known distribution of Hyphessobrycon cantoi (red circles indicate paratypes localities; yellow diamond indicates type locality).
Sexual dimorphism. Last unbranched and two anteriormost branched anal fin rays larger in females, resulting in a slightly more pointed and developed anal-fin lobe (ZUEC 12440; ZUEC 14597). Males present tiny bony hooks on distal half of last unbranched anal-fin ray and all branched anal-fin rays (5 to 44), on caudal-fin rays (0 to 8), on distal half of dorsal-fin rays (0 to 30), on all extension of pelvic-fin rays (7 to 40) and on distalmost region of pectoral fin (0 to 12) (all bony hook counts made in a single C&S specimen, ZUEC 12440, 25.2 mm SL). Bony hooks larger on anteriormost rays of anal fin and on pelvic fin (ZUEC 12440, 65, 21.0–26.7 mm SL; ZUEC 14597, 5, 20.3–24.8 mm SL). Bony hooks of dorsal, caudal and pectoral fins only discernible in C&S individuals. Smaller male presenting bony hooks 20.3 mm SL (ZUEC 12440). Females reach larger sizes than males (largest female examined with 33.3 mm SL and largest male with 27.8 mm SL).
Geographical distribution. Hyphessobrycon cantoi is known from streams draining into the rio Tapajós mouth, and in streams immediately eastward draining directly into the rio Amazonas, state of Pará, Brazil (Fig. 3).
Ecological notes. Hyphessobrycon cantoi is known from slow-flowing, clear water streams. Specimens collected from Igarapé do Diamantino, Santarém, state of Pará, were observed in schools with up to 25 individuals, typically swimming among floating leaves of aquatic plants (Nymphaea sp. or Nymphoides sp.). Syntopic species were Copella callolepis (Regan, 1912), Crenuchus spilurus Gunther, 1863, and Nannostomus marginatus Eigenmann, 1909. Another locality from where the species is known (a stream tributary of Lago Verde) presented the following range of physico-chemical parameters across a period of 10 months (November-August): temperature 26.4–27.6ºC, pH 4.3–5.2, dissolved oxygen 3.6–7.9, conductivity 8.52–13.12, and turbidity 4.22–13.3 (Juan D. Bogotá-Gregory, 2020, pers. comm.). Gut content of 3 C&S individuals (ZUEC 12440) revealed the presence of ants, beetles, Diptera larvae, a Trichoptera larvae, unidentified arthropods remains and unidentified organic matter.
Etymology. The specific name is a homage to André Luiz C. Canto, curator of the fish collection of the Universidade Federal do Oeste do Pará (UFOPA), in recognition of his contribution to the knowledge of the fishes from the rio Tapajós basin. A genitive noun.
Conservation status. Hyphessobrycon cantoi is only known from streams in the lower Tapajós and nearby streams emptying directly into the rio Amazonas, with an Extent of Occurrence (EOO) of 134.9 km² in Santarém. Some of the streams where the species was collected lie at periurban areas, and consequently are under some anthropogenic disturbance, i.e., deforestation, and domestic sewage disposal. The type locality and some of the collection sites of the species lie within the Área de Proteção Ambiental Alter do Chão, which gives them some level of protection against environmental degradation, but even these areas are currently under real state pressure and human induced wildfires. Considering the small EOO and expected increase in degradation of many of the streams from where the species is currently known in the coming years due to the expansion of the urban area of Santarém, we recommend Hyphessobrycon cantoi to be considered Near Threatened (NT) due to its closeness to fulfilling the criteria as Vulnerable (VU) B2 biii following the International Union for Conservation of Nature (IUCN) rules (2019).
Remarks. Variation on the humeral blotch thickness was observed in populations of Hyphessobrycon cantoi, with thicker humeral blotches occurring in populations of the westernmost portion of the distribution of the species (Lago Verde drainage, rio Tapajós basin) in comparison with population of the easternmost portion of the distribution (igarapé do Diamantino, rio Amazonas basin). Specimens from the Lago Verde drainage also present fewer chromatophores on the abdominal region when compared to the easternmost populations, where, in some individuals, a subtle reticulated pattern can be discerned (compare Figs. 1A-B).
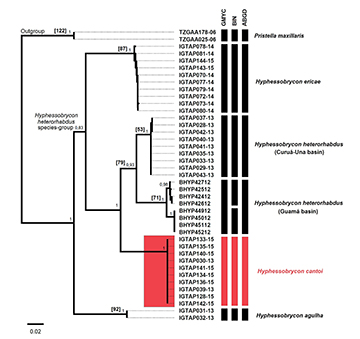
FIGURE 4 | Neighbor-joining tree of Hyphessobrycon cantoi and others lineages of H. heterorhabdus species-group. Red color indicating the new species. Species were delimited through GMYC, BIN and ABGD analysis. Values in brackets expressed the number variable sites that separate H. cantoi and values in branches indicate bootstrap values.
Molecular species delimitation. We analyzed 37 COI sequences (DNA barcoding) belonging to the Hyphessobrycon heterorhabdus species-group from the Tapajós, Curuá-Una and Guamá basins. The sequences were 602bp long and revealed 121 polymorphic sites, base composition of T (31.9%), C (26.5%), A (23.0%), G (18.5%). We did not observe stop codons and indels.The Neighbor-joining reconstruction clearly delimited the H. heterorhabdus species-group (Bootstrap = 100%) as sister group to H. agulha and have H. cantoi aligned with H. heterorhabdus and H. ericae (Fig. 4). The three methods used to delimit species based on COI sequences were congruent to illuminate the molecular identity and the specific status of H. cantoi, which diverged from its congeners by 9.6 to 17.8% (Tab. 2). Additionally, we observed two clusters within H. heterorhabdus that segregate individuals from rio Curuá-Una and rio Guamá basins by 6.4% COI divergence, and H. agulha diverged from the H. heterorhabdus species-group by 14.8 to 18.6%.
TABLE 2 | Pairwise genetic distances (K80 model) between Hyphessobrycon species and Pristella maxillaris (outgroup).
Species | 1 | 2 | 3 | 4 | 5 | 6 | |
1 | H. cantoi |
|
|
|
|
|
|
2 | H. heterorhabdus (Curuá-Una basin) | 0.096 |
|
|
|
|
|
3 | H. heterorhabdus (Guamá basin) | 0.124 | 0.064 |
|
|
|
|
4 | H. ericae | 0.163 | 0.152 | 0.164 |
|
|
|
5 | H. agulha | 0.178 | 0.170 | 0.186 | 0.148 |
|
|
6 | Pristella maxillaris | 0.246 | 0.223 | 0.234 | 0.200 | 0.189 |
|
Discussion
Hyphessobrycon cantoi present the typical color pattern of the Hyphessobrycon heterorhabdus species-group (Lima et al., 2014; Moreira, Lima, 2017; Faria et al., 2020). viz.: an anteriorly well defined humeral blotch continuous with a midlateral dark stripe that becomes blurred posteriorly and a tricolored longitudinal pattern composed dorsally by a red or reddish longitudinal stripe, a middle iridescent, golden or silvery longitudinal stripe, and ventrally by the longitudinal dark pattern composed by the humeral blotch and dark midlateral stripe. The midlateral stripe overlaps with the humeral blotch posteriorly and typically is not possible to clearly identify the limits between both color features components. Also, some species the midlateral dark stripe is poorly developed or even presumably absent (e.g., Hyphessobrycon amapaensis, most populations of H. ericae, H. montagi, and H. wosiackii). A full analysis of the color patterns present in the species currently included in the group will be presented elsewhere (TCF, FCTL, work in progress).
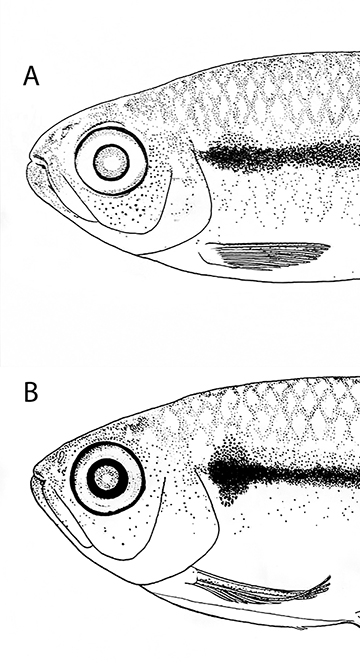
Among the species belonging to the Hyphessobrycon heterorhabdus species-group, H. cantoi closely resembles H. heterorhabdus, currently reported from eastern Pará (Eigenmann, 1921; Géry, 1966) to the rio Curuá-Una basin (Zarske, Géry, 1997; Silva-Oliveira et al., 2016) and the rio Cupari basin (a tributary of the rio Tapajós) (Silva-Oliveira et al., 2016). The type locality of Hyphessobrycon heterorhabdus was vaguely stated as being “Para; Brazil” (Ulrey, 1894, 1895). A full discussion on the question of type-locality of H. heterorhabdus will be presented elsewhere (TCF, FCTL, work in progress), but we can advance herein that it very likely corresponds to the outskirts of Belém. All examined populations of H. heterorhabdus distributed along coastal basins of Pará westward to the rio Curuá-Una and rio Cupari possess a ventral extension of the humeral blotch, which is absent in H. cantoi (Fig. 5). This ventral extension is particularly developed in the H. heterorhabdus populations occurring in the rio Curuá-Una basin, which are the geographically closest to H. cantoi. A full discussion on the geographical variation of Hyphessobrycon heterorhabdus will be presented elsewhere (TCF, FCTL, work in progress). No other color pattern or morphological character was identified as distinguishing both species.
FIGURE 5 | Schematic representation of humeral blotch and anterior portion of midlateral stripe between Hyphessobrycon cantoi (A, ZUEC 14597, paratype 32.1 mm SL, rio Amazonas basin) and H. heterorhabdus (B,INPA 41126, 27.8 mm SL, rio Curuá-Una basin).
The molecular data (DNA barcoding) demonstrated the genetic singularity of H. cantoi that was previously reported as a putative undescribed species (Guimarães et al., 2018). Herein, we found robust evidence from phylogenetic and statistical species delimitation (GMYC, BIN, ABGD) supporting the specific status for this taxon. The speciation of H. cantoi is followed by deep divergence (9.6–12.4%) from its nearest neighbor species, H. heterorhabdus. A conservatism of morphological features accompanied of high genetic differentiation seems to be a trend for some Hyphessobrycon lineages (e.g., Guimarães et al., 2019).
The integrative taxonomy has emerged as a powerful approach to investigate cryptic diversity and complex taxonomic groups (Dayrat, 2005; Padial et al., 2010). Herein, we demonstrate a successful combination of molecular and morphological data to diagnose and describe a new characid fish species from the Amazon basin. The genus Hyphessobrycon is a species-rich, taxonomically problematic group widespread throughout the Amazon, Orinoco and guyanese basins, and some additional cases of cryptic diversity within the genus have been recently published (e.g., Guimarães et al., 2019, 2020). On the other hand, populations from distinct tributaries may exhibit markedly genetic divergence, as observed herein in H. heterorhabdus from the rio Curuá-Una and rio Guamá basins, without any significative morphological evidence pointing them as distinct taxa. Future works using integrative taxonomy will be useful to help unravel the diversity and evolutionary history of Hyphessobrycon and other speciose characid genera.
Comparative material examined. In addition to Faria et al. (2020). Hyphessobrycon heterorhabdus: INPA 41126, 2, 23.3–27.7 mm SL, Brazil, Pará, rio Curuá-Una basin.
Acknowledgments
The present paper is derived from the first author M.Sc. dissertation entitled “Sistemática do grupo Hyphessobrycon heterorhabdus (Ulrey) (Characiformes, Characidae)”, presented in January 2020 at the Instituto de Biologia, Universidade Estadual de Campinas, and we thank the members of the several committees that assessed the dissertation, namely Flávio D. Passos, André R. Garrafoni, André Victor L. Freitas, Rafaela P. Ota, Fabrizio M. Machado, Ricardo C. Benine, and Túlio F. Teixeira for several suggestions for its improvement. André Luiz C. Canto, Frank Raynner V. Ribeiro, Jordson S. Souza, Marcos C. Lima, Poliane B. Silva, Claudia S. Chaves and Luan Maciel provided invaluable logistical support to the first author during a trip to Santarém. André Luiz C. Canto also provided Fig. 2. Juan D. Bogotá-Gregory collected part of the material herein studied and provided the physico-chemical parameters of one of the collecting sites. Tatiane C. Mariguela sequenced the samples of H. heterorhabdus from rio Guamá as part of a DNA barcoding study in Hyphessobrycon. Part of the material herein studied was collected with funds provided by the projects “Aquatic Faunal Survey of the Lower Amazon” (NSF grant DEB–1146734 to William G. R. Crampton) and “Systematics of the tetras (genera Hemigrammus, Hyphessobrycon, Thayeria, Parapristella and Bryconella”) (FAPESP grant # 2011/51532–7 to Flávio C.T. Lima). Figs. 1A–B were prepared by Eduardo G. Baena. Figs. 5A-B were prepared by Zaqueu dos Santos (UFOPA), and Fig. 3 by Mendelsohn Fujiie (UFOPA). We are grateful to André Luiz C. Canto (UFOPA) and Wolmar B. Wosiacki (MPEG) for access to material used in this study. The authors were funded by CAPES (TCF; LRRR, grant #AUXPE 3318/2013; KLAG, grant #88882.457158/2019–01), CNPq/FAPEAM (INCT-ADAPTA II, LRRR), and FAPESP (FCTL, grant #2013/20936–0). Specimens used in this study were collected under the licences ICMBio/Sisbio #55903 (FCTL) and SISBIO #32653–3 (LRRR).
References
Aljanabi SM, Martinez I. Universal and rapid salt-extraction of high quality genomic DNA for PCR-based techniques. Nucleic Acids Res. 1993; 25(22):4692–93. https://doi.org/10.1093/nar/25.22.4692
Allen GR, Unmack PJ, Hadiaty RK. The Goeldiei group of rainbowfishes (Melanotaeniidae) from the Birds Neck region of New Guinea (Papua and West Papua provinces, Indonesia) with descriptions of five new species and recognition of Melanotaenia dumasi Weber. Aqua. 2016; 22(1):1–32.
Baldwin CC, Weigt LA. A new species of soapfish (Teleostei: Serranidae: Rypticus), with redescription of R. subbifrenatus and comments on the use of DNA barcoding in systematic studies. Copeia. 2012; 2012(1):23–36. https://doi.org/10.1643/CG-11-035
Costa WJEM, Amorim PF, Mattos JLO. Species delimitation in annual killifishes from the Brazilian caatinga, the Hypsolebias flavicaudatus complex (Cyprinodontiformes: Rivulidae): implications for taxonomy and conservation. Syst Biodivers. 2012; 10:71–91. https://doi.org/10.1080/14772000.2012.664177
Dayrat B. Towards integrative taxonomy. Biol J Linn Soc. 2005; 85(3):407–15. https://doi.org/10.1111/j.1095-8312.2005.00503.x
Drummond AJ, Rambaut A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol. 2007; 7(1):1-08. https://doi.org/10.1186/1471-2148- 7-214
Eigenmann CH. The American Characidae [Part 2]. Mem Mus Comp Zool. 1918; 43(2):103–208, pls. 9–11, 13, 17–29, 33, 78–80, 93.
Eigenmann CH. The American Characidae. Mem Mus Comp Zool. 1921; 43(3):208–310, pls. 30-32. 40–55, 61–62, 64, 66, 69, 85, 87, 89, 92.
Faria TC, Bastos DA, Zuanon J, Lima FCT. A new Hyphessobrycon (Characiformes: Characidae) of the Hyphessobrycon heterorhabdus species-group from the Central Amazon basin, Brazil. Zootaxa. 2020;4859(2):275–84. https://doi.org/10.11646/zootaxa.4859.2.6
Fink WL, Weitzman SH. The so-called Cheirodontin fishes of Central America with descriptions of two new species (Pisces: Characidae). Smithson Contr Zool. 1974; 172:1–46. https://doi.org/10.5479/si.00810282.172
Fujisawa T, Barraclough TG. Delimiting species using single-locus data and the generalized mixed Yule coalescent approach: a revised method and evaluation on simulated data sets. Syst Biol. 2013; 62(5):707–24. https://doi.org/10.1093/sysbio/syt033
Géry J. Hyphessobrycon vilmae sp. nov., a new tetra from the upper Juruena, Brazil, with keys to the heterorhabdus-like species. Ichthyologica, the Aquarium Journal. 1966; 37:63–70.
Guimarães EC, Brito PS, Feitosa LM, Carvalho Costa LF, Ottoni FP. A new cryptic species of Hyphessobrycon Durbin, 1908 (Characiformes, Characidae) from the Eastern Amazon, revealed by integrative taxonomy. Zoosyst Evol. 2019; 95(2):345–60. https://doi.org/10.3897/zse.95.34069
Guimarães, EC, Brito PS, Bragança, PHN, Santos JP, Katz AM, Carvalho Costa, LF, Ottoni, FP. Integrative taxonomy reveals two new cryptic species of Hyphessobrycon Durbin, 1908 (Teleostei: Characidae) from the Maracaçumé and middle Tocantins River basins, eastern Amazon region. Eur J Taxon 2020; 723:77–107. https://doi.org/10.5852/ejt.2020.723.1145
Guimarães KLA, Sousa MPA, Ribeiro FRV, Porto JIR, Rodrigues LRR. DNA barcoding of fish fauna from low order streams of Tapajós River basin. PLoS One. 2018; 13(12):e0209430. http://dx.doi.org/10.1371/journal.pone.0209430
Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acid S. 1999; 41:95–98.
Ingenito LFS, Lima FCT, Buckup PA. A new species of Hyphessobrycon Durbin (Characiformes: Characidae) from the rio Juruena basin, Central Brazil, with notes on H. loweae Costa & Géry. Neotrop Ichthyol. 2013; 11(1):33–44. http://dx.doi.org/10.1590/S1679-62252013000100004
International Union for Conservation of Nature (IUCN). Standards and Petitions Subcommittee. Guidelines for using the IUCN Red List categories and criteria. Version 14. Gland; 2019. Available from: http://www.iucnredlist.org/documents/RedListGuidelines.pdf
Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980; 16:111–20.
Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 2018; 35(6):1547–49. https://doi.org/10.1093/molbev/msy096
Landi M, Dimech M, Arculeo M, Biondo G, Martins R, Carneiro M, Carvalho GR, Lo Brutto S, Costa FO. DNA barcoding for species assignment: the case of Mediterranean marine fishes. PLoS One. 2014; 9(9):e106135. http://dx.doi.org/10.1371/journal.pone.0106135
Lima FCT, Coutinho DP, Wosiacki WB. A new Hyphessobrycon (Ostariophysi: Characiformes: Characidae) from the middle Amazon basin, Brazil. Zootaxa. 2014;3872(2):167–79. http://dx.doi.org/10.11646/zootaxa.3872.2.3
Melo BF, Ochoa LE, Vari RP, Oliveira C. Cryptic species in the Neotropical fish genus Curimatopsis (Teleostei, Characiformes). Zool Scr. 2016; 45(6):650–58. http://dx.doi.org/10.1111/zsc.12178
Mirande JM. Morphology, molecules and the phylogeny of Characidae (Teleostei, Characiformes).Cladistics. 2019; 35(3):282–300. https://doi.org/10.1111/cla.12345
Moreira CR, Lima FCT. Two new Hyphessobrycon (Characiformes: Characidae) species from Central Amazon basin, Brazil. Zootaxa. 2017;4318(1):123–34. http://dx.doi.org/10.11646/zootaxa.4318.1.5
Ohara WM, Teixeira TF, Albornoz-Garzón JG, Mirande JM, Lima FCT. Hyphessobrycon rheophilus, a new species from rapids of the Amazon and Orinoco river basins (Characiformes: Characidae: Stethaprioninae). Zootaxa. 2019;4712(4):561–75. http://dx.doi.org/10.11646/zootaxa.4712.4.5
Oliveira C, Avelino GS, Abe KT, Mariguela TC, Benine RC, Orti G, Vari RP, Castro RMC. Phylogenetic relationships within the speciose family Characidae (Teleostei: Ostariophysi: Characiformes) based on multilocus analysis and extensive ingroup sampling.BMC EvolBiol. 2011; 11(275):1–25. https://doi.org/10.1186/1471-2148-11-275
Ota RR, Carvalho FR, Pavanelli CS. Taxonomic review of the Hyphessobrycon panamensis species-group (Characiformes: Characidae). Zootaxa. 2020; 4751(3):401–36. http://doi.org/10.11646/zootaxa.4751.3.1
Padial JM, Miralles A, De la Riva I, Vences M. The integrative future of taxonomy. Front Zool. 2010; 7(16):1–14. https://doi.org/10.1186/1742-9994-7-16
Pereira LHG, Hanner R, Foresti F, Oliveira C. Can DNA barcoding accurately discriminate megadiverse Neotropical freshwater fish fauna? BMC Genet. 2013; 14(20):1–14. https://doi.org/10.1186/1471-2156-14-20
Pons J, Barraclough TG, Gomez-Zurita J, Cardoso A, Duran DP, Hazell S, Kamoun S, Sumlin WD, Vogler AP. Sequence-based species delimitation for the DNA taxonomy of undescribed insects. Syst Biol. 2006; 55(4):595–609. https://doi.org/10.1080/10635150600852011
Puillandre N, Lambert A, Brouillet S, Achaz G. ABGD, Automatic Barcode Gap Discovery for primary species delimitation. Mol Ecol. 2012; 21(8):1864–77. https://doi.org/10.1111/j.1365-294X.2011.05239.x
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. Vienna: Austria; 2014. Available from: http://www.R-project.org
Ratnasingham S, Hebert PDN. A DNA-based registry for all animal species: The Barcode Index Number (BIN) System. PLoS One. 2013; 8(7):e66213. https://doi.org/10.1371/journal.pone.0066213
Sabaj MH. Standard symbolic codes for institutional resource collections in herpetology and ichthyology: an online reference [Internet]. Washington: American Society of Ichthyologists and Herpetologists; 2019. Available from: https://asih.org/sites/default/files/2019-04/Sabaj_2019_ASIH_Symbolic_Codes_v7.1.pdf
Schaefer SA, Chakrabarty P, Geneva AJ, Sabaj Pérez MH. Nucleotide sequence data confirm diagnosis and local endemism of variable morphospecies of Andean astroblepid catfishes (Siluriformes: Astroblepidae). Zool J Linn Soc-Lond. 2011; 162(1):90–102. http://dx.doi.org/10.1111/j.1096-3642.2010.00673.x
Silva-Oliveira C, Canto ALC, Ribeiro FRV. Stream ichthyofauna of the Tapajós National Forest, Pará, Brazil. Zookeys. 2016;580:125–44. http://doi.org/10.3897/zookeys.580.6659
Taylor WR, Van Dyke GC. Revised procedures for staining and clearing small fishes and other vertebrates for bone and cartilage study. Cybium. 1985(2);9:107–19.
Terán GE, Benitez MF, Mirande JM. Opening the Trojan horse: phylogeny of Astyanax, two new genera and resurrection of Psalidodon (Teleostei: Characidae). Zool J Linnean Soc. 2020; 1217–34. https://doi.org/10.1093/zoolinnean/zlaa019
Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994; 22(22):4673–80.
Tornabene L, Baldwin C, Weigt LA, Pezold F. Exploring the diversity of western Atlantic Bathygobius (Teleostei: Gobiidae) with cytochrome c oxidase-I, with descriptions of two new species. Aqua. 2010; 16(4):141–70.
Ulrey AB. Preliminary descriptions of some new South American Characinidae. Am Nat. 1894; 28:610–11.
Ulrey AB. The South American Characinidae collected by Charles Frederick Hartt. Ann N Y Acad Sci. 1895; 8:257–300.
Vitorino CA, Oliveira RCC, Margarido VP, Venere PC. Genetic diversity of Arapaima gigas (Schinz, 1822) (Osteoglossiformes: Arapaimidae) in the Araguaia-Tocantins basin estimated by ISSR marker. Neotrop Ichthyol. 2015; 13(3):557–68. https://dx.doi.org/10.1590/1982-0224-20150037
Ward RD, Zemlak TS, Innes BH, Last PR, Hebert PDN. DNA barcoding Australia’s fish species. Philos Trans R Soc B Biol Sci. 2005; 360(1462):1847–57. https://doi.org/10.1098/rstb.2005.1716
Weigt LA, Baldwin CA, Driskell A, Smith DG, Ormos A, Reyier EA. Using DNA barcoding to assess Caribbean reef fish biodiversity: expanding taxonomic and geographic coverage. PLoS One. 2012; 7(7):e41059. http://dx.doi.org/10.1371/journal.pone.0041059
Weitzman SH, Fink WL. Relationships of the neon tetras, a group of South American freshwater fishes (Teleostei, Characidae), with comments on the phylogeny of New World characiforms. Bull Mus Comp Zool. 1983; 150(6):339–95.
Weitzman SH, Palmer L. A new species of Hyphessobrycon (Teleostei: Characidae) from the Neblina region of Venezuela and Brazil, with comments on the putative ‘rosy tetra clade. Ichthyol Explor Fres. 1997; 7(3):209–42.Zarske A, Géry J. Hyphessobrycon amapaensis spec. nov., ein neue and mutmaßliche Stellvertreterart von Hyphessobrycon heterorhabdus (Ulrey, 1894) aus dem Bundesstaat Amapa in Brasilien (Teleostei: Characiformes: Characidae). Zool Abh. 1998; 50(1):19–26.
Zarske A, Géry J. Hyphessobrycon amapaensis spec. nov., ein neue and mutmaßliche Stellvertreterart von Hyphessobrycon heterorhabdus (Ulrey, 1894) aus dem Bundesstaat Amapa in Brasilien (Teleostei: Characiformes: Characidae). Zool Abh. 1998; 50(1):19–26.
Authors
Tiago C. Faria[1] ![]()
![]() , Karen L. A. Guimarães[2]
, Karen L. A. Guimarães[2] ![]() , Luís R. R. Rodrigues[2]
, Luís R. R. Rodrigues[2]![]() ,Claudio Oliveira[3]
,Claudio Oliveira[3] ![]() andFlávio C.T. Lima[1]
andFlávio C.T. Lima[1] ![]()
[1] Museu de Zoologia da Universidade Estadual de Campinas “Adão José Cardoso”, Caixa Postal 6109, 13083-863 Campinas, SP, Brazil. (TCF) tiago_faria5@hotmail.com; (FCTL) fctlima@gmail.com (corresponding author).
[2] Laboratório de Genética & Biodiversidade, Instituto de Ciências da Educação, Programa de Pós-Graduação em Recursos Naturais da Amazônia, Universidade Federal do Oeste do Pará, Rua Vera Paz, s/n (Unidade Tapajós) Bairro Salé, 68040-255 Santarém, PA, Brazil. (KLAG) karen.guimaraes.bio@gmail.com; (LRRR) luisreginaldo.ufpa@hotmail.com.
[3] Laboratório de Biologia e Genética de Peixes (LBP), Instituto de Biociências de Botucatu (IBB), Universidade Estadual Paulista “Júlio de Mesquita Filho” (UNESP), Botucatu, SP, Brazil. claudio.oliveira@unesp.br.
Authors Contribution 
Tiago C. Faria: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Project administration, Validation, Visualization, Writing-original draft, Writing-review and editing.
Karen L. A. Guimarães: Data curation, Formal analysis, Investigation, Methodology, Software, Validation, Writing-original draft, Writing-review and editing.
Luís R. R. Rodrigues: Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Resources, Software, Supervision, Validation, Writing-original draft, Writing-review and editing.
Claudio Oliveira: Data curation, Software, Writing-review and editing.
Flávio C.T. Lima: Conceptualization, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Supervision, Validation, Visualization, Writing-original draft, Writing-review and editing.
Ethical Statement
Specimens used in this study were collected under the licences ICMBio/Sisbio #55903 (FCTL) and SISBIO #32653–3 (LRRR).
Competing Interests
The authors declare no competing interests.
How to cite this article
Faria TC, Guimarães KLA, Rodrigues LRR, Oliveira C, Lima FCT. A new Hyphessobrycon (Characiformes: Characidae) of the Hyphessobrycon heterorhabdus species-group from the lower Amazon basin, Brazil. Neotrop Ichthyol. 2021; 19(1):e200102. https://doi.org/10.1590/1982-0224-2020-0102
Copyright
This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Distributed under
Creative Commons CC-BY 4.0

© 2021 The Authors.
Diversity and Distributions Published by SBI
![]() Submitted September 29, 2020
Submitted September 29, 2020
![]() Accepted December 28, 2020
Accepted December 28, 2020
by George Mattox
![]() Epub 08 March, 2021
Epub 08 March, 2021